Research Article Open Access
Killer Typhoons Began to Impact the Japanese Islands from ca.1.55 Ma, Based on Phylogeography of Chlorogomphus (Gliding Dragonfly)
| Osozawa S1* and Wakabayashi J2 | |
| 1Department of Earth Sciences, Graduate School of Science, Tohoku University, Sendai 980-8578, Japan | |
| 2Department of Earth and Environmental Sciences, California State University, Fresno, CA 93740, USA | |
| Corresponding Author : | Osozawa S Department of Earth Sciences Graduate School of Science Tohoku University Sendai 980-8578, Japan Tel: +8-22-795- 6619 E-mail: osozawa@m.tohoku.ac.jp |
| Received April 12, 2015; Accepted May 08, 2015; Published May 18, 2015 | |
| Citation: Osozawa S, Wakabayashi J (2015) Killer Typhoons Began to Impact the Japanese Islands from ca.1.55 Ma, Based on Phylogeography of Chlorogomphus (Gliding Dragonfly). J Earth Sci Climat Change S3:003. doi: 10.4172/2157-7617.S3-003 | |
| Copyright: ©2015 Osozawa S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Visit for more related articles at Journal of Earth Science & Climatic Change
Abstract
Chlorogomphus is a golden ring dragonfly, and characterized by swarming behavior, and strong gliding and flying. Therefore this dragonfly is expected to be easily carried by typhoon winds. This dragonfly immigrated and colonized the Pacific coast of southern Japan main islands from the Ryukyu islands. In this study, we demonstrate that the Japan mainland population is phylogenetically common to the northern Ryukyu population, but distinct from the southern Ryukyu and Taiwan populations. The East China Sea was formed between the Ryukyu island arc and Asian mainland by the rifting of the Okinawa trough that started at 1.55 Ma. Prior to this time typhoons lost strength when heading overland over the Ryukyu continental arc and continental China, but since then have maintained their strength northward because they were over water (the newly opened Okinawa trough). Chlorogomphus is interpreted to have speciated on each island, but it also migrated northward as a result of typhoons that carried it further than it could normally fly. The present paper shows that such typhoons that now ravage Japan may have been generated since 1.55 Ma and will continue to be dangerous, because the opening of the Okinawa trough is an ongoing process. This phenomenon is unrelated to global warming.
| Keywords |
| Ryukyu; East China sea; COI; 28SrRNA; Immigration |
| Introduction |
| As global warming progresses, typhoons may tend to become more powerful, and such typhoons are called super typhoons. This paper suggests that historical configuration of land and sea may also affect the strength of typhoons, and such damaging typhoons that have impacted Japan were generated after 1.55 Ma. |
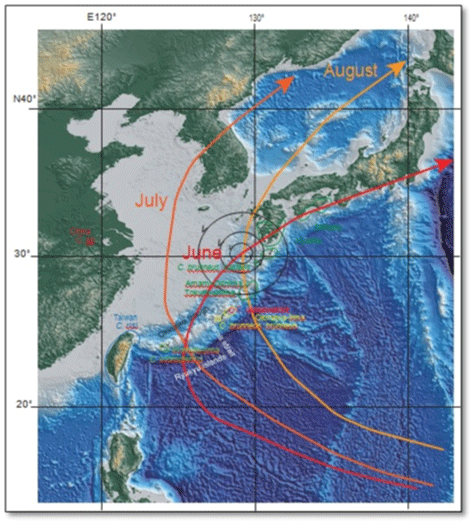
| The East China Sea is a submerged continental shelf (Figure 1). This subsidence was a consequence of rifting and subsequent opening of the Okinawa trough that has continued to the present [1]. The Kuroshio warm current began to inflow to the East China Sea since 1.55 Ma [1]. Before 1.55 Ma typhoons made landfall on the sub-aerially exposed Chinese continent, that including the present Ryukyu arc (that was a continental margin arc before the rifting), and soon attenuated in energy. Since 1.55 Ma the typhoon path in this region has remained over water and the storms retain their power and intensity on their northward path because they do not make landfall on the continent. |
| ¨The rifting and subsidence separated the Ryukyu Islands, that comprise the Ryukyu arc, from the Chinese continent, and such isolation triggered allopatric speciation of terrestrial insects [2]. We have been conducting molecular phylogenetic studies of the Ryukyu endemic species, including analysis of related species in Japan, Taiwan, Korea, and China [3,4]. |
| Chlorogomphus is a type of dragonfly, and the Ryukyu islands, southern Japanese islands, and Taiwan yield endemic species (Figure 1). From the target specimens, we obtained mitochondrial COI and nuclear 28SrRNA sequences, and the sequences showed allopatrical speciation in Okinawa-jima, Iriomote-jima, and Taiwan (Figure 1). However, sequences of populations from Amami-Oshima, Tokuno-shima, and Shikoku, Japan, were similar, so speciation was not recognized there (Figure 1). This research aims to explain why speciation did not take place on Amami-Oshima, Tokuno-shima, and Shikoku. |
| Materials and Methods |
| Taxon sampling |
| We collected Chlorogomphus specimens from Shikoku-Japan, Amami-Oshima, Tokuno-shima, Okinawa-jima, Iriomote-jima, Taiwan, and Zhejiang-China (Table 1), but could not collect samples from Kyushu-Japan, Nakano-shima-Tokara islands, and Tokashikijima- Kerama islands (Figure 1). |
| Female Chlorogomphus are characterized by dark wings, contrasting with the transparent wings of male. However, females of C. okinawensis in Okinawa-jima and C. suzukii in Taiwan have transparent wings, and they are sympatrically distributed with other Chlorogomphus whose females have dark wings. We collected the former from Okinawa-jima (Table 1), but were unsuccessful in collecting specimens from Taiwan. |
| DNA extraction, polymerase chain reaction amplification, and sequencing |
| Collected leg muscle samples used for analyses were stored in 1.5 mL tubes filled by 99% ethanol, with the remaining material stored at -30ºC in a freezer. DNA extracting was done using GenElutemTM Mammalian Genomic DNA Miniprep Kit by Sigma-Aldrich. |
| Primers used for amplification for mitocondrial COI-tRNA-COII gene were COS2265 (5-GCA CAA GAA AGA GGG AAA AAA GA- 3), and COA3625 (5-GCC CCA CAA ATT TCG GAAC ATT G-3), following Kiyoshi et al. [5]. When these primers were not effective, internal primers were newly designed as cgOsoF (5-TGA TTA GCA ACT CTT CAC GG-3) and cgOsoR (5-TGT AAG GGT AAA ACT GCT CG-3). |
| Primers used for amplification for nuclear 28SrRNA are 28sf (5- AAG GTA GCC AAA TGC CTC ATC-3) and 28sr (5-AGT AGG GTA AAA CTA ACC T-3), following Hasegawa and Kasuya [6]. |
| GoTaq Green Master Mix G2, Promega, for COI amplication, and Tks GflexTM DNA Polymerase, Takara, for 28SrRNA amplication were used for PCR, following that protocol. Annealing temperature for COI and 28SrRNA primers, including internal primers, was 53ºC. PCR product was purified using Wizard SV Gel and PCR Clean-Up System, Promega. Sequencing was done by Macrogen Japan, and National Museum of Nature and Science, Tsukuba. 1422 bp and 816 bp sequences were readable both for COI-tRNA-COII and 28SrRNA genes. |
| Sequence alignment and phylogenetic analysis |
| The non-protein cording tRNA-Leu region was excluded from the alignment, and a 1381 bp COI-COII sequence was used. |
| Sequence alignment was done using ClustalW accompanied in MEGA5 [7]. No gap was observed in the aligned sequences including 28SrRNA. Codon translation was checked by ExPASy-Translate tool, Bioinformatics Resource Tool, and such COI sequence data are reflected in our registered data in DDBJ/GenBank. The sequences were also checked for similarity by BLAST (Basic Local Alignment Search Tool) offered by DDBJ/GenBank. |
| Our evolutionary analyses were conducted using MEGA5 [6], as noted in Osozawa et al. [2]. For constructing the ML (Maximum Likelihood) trees, we chose the evolutionary model of Tamura and Nei [8], and uniform rates are presumed. The number of bootstrap replications was 500. Codon positions included were 1st+2nd+3rd+Noncoding. |
| Our geological model predicts the simultaneous but multiple branching of nodes in the ML tree. Because of the geological isolation of islands at 1.55 Ma, we assigned this age to the simultaneously branching nodes. The branching was not actually simultaneous, but we selected a calibration point at the basal node (starting point of the vicariance). |
| A phylogenetic tree of the COI gene was displayed using raxmlGUI v1.3 [9]. The topology for COI-COII was displayed using other PC software packages. A topology was constructed using a series of software of BEAST (Bayesian Evolutionary Analysis Sampling Trees), running BEAUti v1.8.0, BEAST v1.8.0, Tracer v1.6, TreeAnnotator v1.8.0, and Figure Tree v1.4.2, in ascending order [10]. Substitution rate (2.81%/m.y.) for COI-COII calculated by MEGA5 was entered in BEAUti v1.8.0, although the molecular clock test was not passed. |
| Combined analyses with the COI-COII and 28SrRNA genes were performed using BEAST. The substitution rate (0.119%/m.y.) for the 28SrRNA gene calculated by MEGA5 was incorporated into BEAUti v1.8.0, and two files for each gene were combined into one file using LogCombiner v1.8.0. The combined data were input into TreeAnnotator v1.8.0, and the consequent tree was drawn using Figure Tree v1.4.2. |
| The relationships among nuclear 28SrRNA as well as COI-COII haplotypes were visually assessed by constructing statistical parsimony networks with a 95% connection limit using TCS v1.21 [11,12]. |
| Results |
| ML tree by MEGA5, raxmlGUI, combined tree by BEAST |
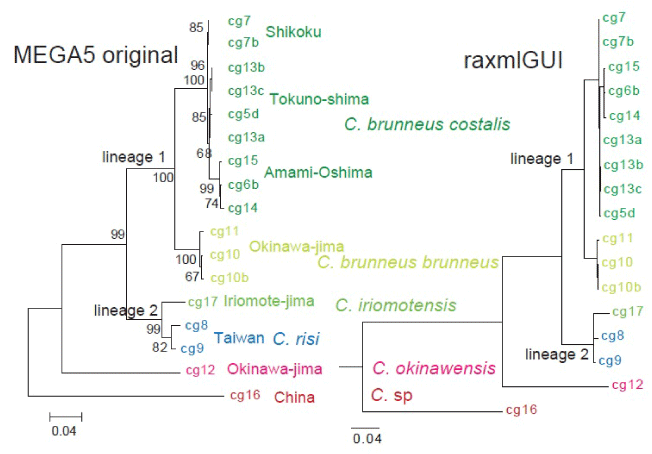
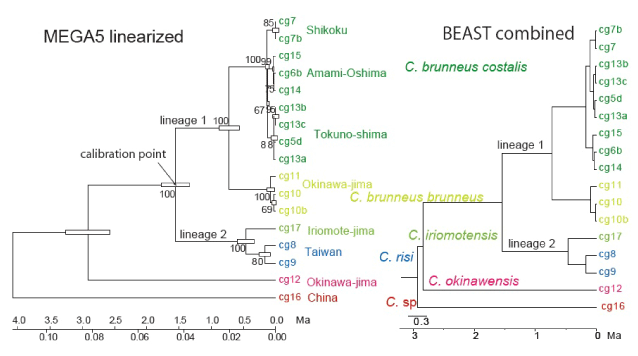
| The topology, branching order, clade, subspecies, and geographical components in the ML tree constructed by MEGA5 are concordant to the raxmlGUI tree (Figure 2). The topology is also completely concordant between the linearized and calibrated ML tree made by MEGA5 and the calibrated BEAST tree made by combined analyses of COI-COII and 28SrRNA (Figure 3). |
| Apart from the outgroup species from China and C. okinawensis (female with transparent wings) from Okinawa-jima, two lineages of Shikoku to Okinawa-jima and Iriomote-jima-Taiwan are recognized (Figure 2). The tree was calibrated with the assumption that these lineages had differentiated at 1.55 Ma (Figure 3). Base substitution rate of COI-COII was estimated 2.81%/m.y. (28SrRNA: 0.119%/m.y.). |
| Lineage 1 consists of two clades of C. brunneus costalis (Shikoku, Amami-Oshima, Tokuno-shima), and C. brunneus brunneus (Okinawa-jima) (Figure 2), which had differentiated at 0.7 Ma (Figure 3). C. brunneus costalis was not differentiated in Shikoku, Amami- Oshima, and Tokuno-shima (Figures 2 and 3). |
| Lineage 2 consists of C. iriomotensis (Iriomote-jima) and C. risi (Taiwan (Figure 2), which had differentiated at 0.5 Ma (Figure 3). C. risi from northern and southern Taiwan had differentiated at 0.2 Ma (Figure 3). |
| Haplotype network by TCS |
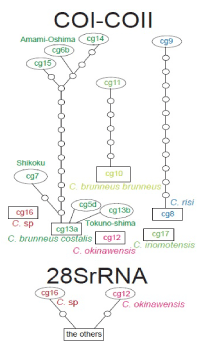
| In the COI-COII haplotype network, each Chlorogomphus species and subspecies constitute distinct haplogroups (Figure 4). The haplotype network for C. brunneus costalis among the Shikoku, Amami-Oshima, and Tokuno-shima populations was especially well constructed (Figure 4). |
| The 28SrRNA sequence is almost identical among the analyzed samples. The haplotype network shows that C. sp. from China and C. okinawensis from Okinawa-jima constitute distinct haplotypes compared to the others (Figure 4). |
| Discussion |
| Speciation but partial gene flow of Chlorogomphus |
| We selected a calibration point at a basal node, corresponding to the node between lineage 1 and 2 (Figure 3), and obtained a reasonable base substitution rate compared to other insect species [2~4]. We conclude that this differentiation is due to allopatric speciation also recognized for the above insect species [2~4]. |
| For speciation, the wide seaway between Okinawa-jima and Iriomote-jima, including the Kerama strait (Figure 1) might have acted as strong barrier, whereas extent of differentiation of each subspecies or geographic clade is relatively minor (speciated at a younger age), especially for C. brunneus costalis among Shikoku, Amami-Oshima, and Tokuno-shima (Figures 3 and 4). The other physical barriers [2~4] were apparently not as effective for Chlorogomphus, and mild to very strong gene flow within these areas probably took place. |
| Northward immigration of Chlorogomphus carried by winds of killer typhoons |
| We propose that typhoons may have have promoted gene flow by carrying dragonflies further than they would normally fly. In the Shikoku (and Kyushu), Amami-Oshima, and Tokuno-shima (Figure 1) area, marginal counterclockwise winds associated with a typical June typhoon with an eye to the north of Amami-Oshima (Figure 1) can carry gliding and swarming dragonflys during this emergence time. Chlorogomphus characteristically swarms in large groups in June, and disperses later. Another possible typhoon transportation mechanism is that large masses of dragonflies were concentrated and carried in the typhoon eye, and then dispersed from the typhoon when it made landfall and diminished in power to an extratropical cyclone. Such a transport mechanism was observed for Anax parthenope in September, 2013. |
| We propose transport by typhoon from Okinawa-jima to Tokunoshima (and Amami-Oshima) and from Taiwan (or within Taiwan) to Iriomote-jima as a mechanism that produced the relatively shallow differentiation of Chlorogomphus in this region (Figure 3). |
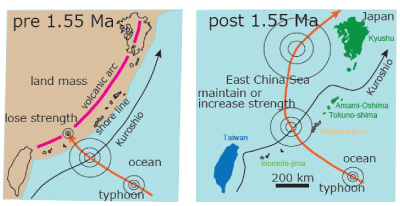
| Prior to the 1.55 Ma rifting of the Okinawa trough, Ryukyu arc (that became the Ryukyu islands) was situated at the eastern margin of the Chinese continent [1]. If strong typhoon landed on this continent, it is expected to have rapidly lost its power (Figure 5). In contrast, after 1.55 Ma, the continental shelf subsided, the East China Sea was formed [1], and a strong typhoon tracking northward would not make land fall and lose power (Figure 5). The Kuroshio warm current (Figure 5) began to inflow and run along the Okinawa trough at 1.55 Ma [1]. Accordingly since 1.55 Ma a typhoon following a northward path may actually increase in power in this region (Figure 5). |
| From 1.55 Ma onward, powerful typhoons, that we call killer typhoons, have ravaged Japan, and because the sea floor spreading in the Okinawa trough continues, this dangerous situation for Japan will persist for the foreseeable future. |
| It may be expected that precipitation associated typhoons reaching Japan drastically changed at ca.1.55 Ma, but this has not been documented in the geological and paleo-meteorological records. Rifting and opening of the Okinawa trough may have significantly affected regional climate as well as insect phylogeny in East Asia. We suggest that the current typhoon behavior in this region has been influenced by the geologic evolution and consequent distribution of land and sea since 1.55 Ma, rather than more recent global warming. |
| Acknowledgement |
| We thank Asahiko Taira and Koji Tojo for constructive reviews. Takehiko Yamanaka, Junko Kobayashi, and Akira Mishima offered Iriomote-jima, Okinawajima, and China sample, respectively. This project was financed through the Osozawa Fund, Tohoku University. Suichi Ogino, Hachijojima Tourist Association, CTI Engineering Co., Ltd., and NEWJEC, Inc. contributed to this fund. |
| References |
|
Tables and Figures at a glance
| Table 1 |
Figures at a glance
 |
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 |
Relevant Topics
- Atmosphere
- Atmospheric Chemistry
- Atmospheric inversions
- Biosphere
- Chemical Oceanography
- Climate Modeling
- Crystallography
- Disaster Science
- Earth Science
- Ecology
- Environmental Degradation
- Gemology
- Geochemistry
- Geochronology
- Geomicrobiology
- Geomorphology
- Geosciences
- Geostatistics
- Glaciology
- Microplastic Pollution
- Mineralogy
- Soil Erosion and Land Degradation
Recommended Journals
Article Tools
Article Usage
- Total views: 14237
- [From(publication date):
specialissue-2015 - Aug 16, 2025] - Breakdown by view type
- HTML page views : 9633
- PDF downloads : 4604
