Research Article Open Access
Potential Role of Fresh Water Apple Snails on H5N1 Influenza Virus Persistence and Concentration in Nature
Marc Souris1,2,3, Daniel Gonzalez4, Witthawat Wiriyarat5, Kamlang Chumpolbanchorn5, Supaluk Khaklang1, Suwannapa Ninphanomchai1, Weena Paungpin5, Somjit Chaiwattanarungruengpaisan5, Ladawan Sariya5, Dubravka Selenic1, Meriadeg AR Gouilh6, Pattamaporn Kittayapong1 and Jean-Paul Gonzalez7*
1Center of Excellence for Vectors and Vector Borne Diseases, Faculty of Science, Mahidol University at Salaya, 999 Phutthamonthon 4, Nakhon Pathom 73170, Thailand
2Institut de Recherche pour le Développement (IRD), UMR 190, 44 Bd de Dunkerque 13572 Marseille Cedex 02, France
3Asian Institute of Technology, P.O. Box 4, Klong Luang, Pathumthani 12120, Thailand
4Montpellier II University, Faculté des Sciences, Place E. Bataillon, 34095 Montpellier, France
5The Monitoring Surveillance Center for Zoonotic Diseases in Wildlife and Exotic Animals, Faculty of Veterinary Science, Mahidol University at Salaya, 999 Phutthamonthon 4, Nakhon Pathom 73170, Thailand
6Institut Pasteur, Laboratory for Urgent Response to Biological Threats, 25 rue du docteur Roux, 75015 Paris, France
7Metabiota Inc., 8757 Georgia Avenue, site 420, Silver Spring 20910, MD, USA
- *Corresponding Author:
- Jean-Paul Gonzalez
Senior Scientist for Emerging Diseases & Biosecurity
Metabiota Inc., 8757 Georgia Avenue
site 420, Silver Spring 20910, MD, USA
Tel: +1 415-398-4712
E-mail: jpgonzalez@metabiota.com
Received Date: December 11, 2014; Accepted Date: January 30, 2015; Published Date: February 04, 2015
Citation: Souris M, Gonzalez D, Wiriyarat W, Chumpolbanchorn K, Gonzalez JP, et al. (2015) Potential Role of Fresh Water Apple Snails on H5N1 Influenza Virus Persistence and Concentration in Nature. Air Water Borne Diseases 4:119. doi:10.4172/2167-7719.1000119
Copyright: © 2015 Souris M et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Air & Water Borne Diseases
Abstract
Background
Influenza A viruses have the remarkable characteristic of sustainability in the environment. Mucus from snails contains sialic acids as the one that may allow avian influenza virions to bind to the vertebrate host cell membrane. Subsequently snails could potentially promote persistence and/or concentration of influenza Avirions through mucus, in wetland environment when virus is released from bird infected feces.
Methods
This article describes experimental research on the potential outcome of apple snails regarding the persistence and concentration of the H5N1 influenza virions in fresh water. The presence of virus was detected from water and snails by hemagglutination test, and H5N1 viral genetic material determined by quantitative RT-PCR
Results
Active virus in the water was demonstrated up to twelve days after water infestation without snails, and up to fourteen days with snails. Also, up to eleven days, the virus and genetic material were detected and tittered from snails. Although the presence of snails did not significantly change the persistence of H5N1 virus in the water, number of positive snail sampled and quantitative RT-PCR data suggest that snails may have the ability to concentrate and carry viral particles.
Conclusions
Ultimately snails could play a role in the virus ecology by concentrating viral particles from water and facilitating virus contact with the bird hosts that feed on them.
Keywords
H5N1; Fresh Water Snail
Introduction
A virus (Orthomyxoviridae family, Influenza virus A genus) had spread to Asia in 2003, it then, expend further widely into Africa and Europe. Since the major 2004 and 2005 epidemics the virus reappeared regularly in many countries without clear understanding of the fundamental of such re-emergences. Nowadays, endemic in several areas (e.g.: Egypt, South East Asia), AI remains a major threat to animal and human health. Virus emergence, spread and re-emergence among bird livestock remain unanswered, while conditions of persistence in wild are poorly understood. Some wild birds are a natural reservoir of AI viruses (AIV), but other wildlife may also participate for the maintenance and/or dissemination the AI H5N1 virus in its natural freshwater habitats including bivalves, zebra mussels, or water fleas among others [1-3] representing an abundant food source, while no studies have been conducted on snails, for wild birds.
The receptors of AIVconsist of sialic acid (SA) derived from a monosaccharide linked to the galactose of a glycoprotein or a glycolipid embedded in the host cell’s membrane and involved in the processes of cell recognition and virus adhesion. N-acetyl-neuraminic [NeuAc], and N-glycolyl-neuraminic [NeuGc] are the SAs detected by AIV including two types of osidic linkage between SA and galactose (α2,3linkage or α2,6 linkage) [4]. The structure of AIV hemagglutinin determines which SA virion can bind to. Species’ specificity for AIV is directly related to the presence on the surface of the host cells of SAs, which possess these characteristics. AIV-A hemagglutinin bind preferentially to SA type NeuGc with the α2, 3 linkage. These SAs are present in the epithelial cells of the avian digestive tract, and determines the digestive tropism of AIV in birds [5]. Snails, of freshwater or land, have mucus that contains SAs similar to those that allow AIV to bind to the cell membrane producing these acids [6]. Therefore snails may promote or maintain the concentration of Influenza virions (notably AIV) in the environment, without necessarily being infected by the virus. Eaten by birds, they could be a link in the eco-epidemiological chain of bird transmission. Research in persistence conditions and existence of non-avian vectors or reservoir raises questions on the potential role of water snails in favoring the concentration and persistence of the virions in nature.
In Thailand, the golden apple snail Pomaceacanaliculata (Ampullariidae family) is widespread in the lake areas and wetlands including rice fields, canals, ponds, among others, which are furthermore visited by many wild and domestic birds, indeed, snails are an abundant food source for many wild waterfowl species (e.g.: open-bill storks) and domestic birds (e.g. free-range ducks) [7]. AI hits Thailand from 2003 to 2008 [8], and some communities of birds, with among them storks, were largely depopulated during the 2004 and 2005 epizootics [9]. Moreover, a correlation between AI and free-range ducks has also been identified in Thailand [10,11].
It was demonstrated that the influenza virus could persist in water for a few hours to several months depending mainly on the viral strain, the temperature and physico-chemical conditions of the water. At 17°C some AI strains remain infectious for more than 200 days, while AIV persistence in water is inversely proportional to the temperature and salinity [12,13]. Thus, at a temperature of 30° C (conditions frequently encountered in Thailand), the persistence of an AIV is less than 10 days [14], moreover H5N1 virus is inactivated after 30 minutes by sunlight with a temperature between 32°C and 35°C [15].
The objective of the present work is to assess the potential role of freshwater apple snails on the persistence and concentration of H5N1 virus particles in water. This was conducted under experimental conditions close to natural environmental conditions found in Thailand’s wetlands and swamps visited by birds (rice fields, lake areas, protected breeding sites).
Methods
The study was done using the AI A virus subtype H5N1 strain A/Chicken/Thailand/VSMU-3-BKK/2004 highly virulent in poultry (close to 100% lethality in infected flocks). Isolates were sequenced at the Faculty of Veterinary Science Laboratory, Mahidol University, Thailand [GenBank: EF593099;EF593106] and, virus suspension produced by cultivating the virus in embryonated eggs with an optimal titer of 2.14x106 TCID50/ml established by MDCK cell culture [17].
The animal model is an invasive common species of freshwater snail, Pomaceacanaliculata,a gastropod mollusk known as “Apple snail” (Figure1d), found in the resting sites of waterfowl and migratory birds in Thailand. Apple snails possess both a gill and a pseudo-lung, allowing them to live in and out of the water [18]. Apple snails can reach a large size (a shell up to 80 mm in females) and longevity can reach four years [19]. They have a high reproductive rate and the ability to adapt to the environment and difficult living conditions. Although Apple snail is herbivorous, in the absence of food it can become cannibalistic or simply feed on dead animals. Voracious, it is competitive with native species of snails [18-21]. Apple snail is native of South America, it was introduced to Asia in the 1980s in order to launch commercial production for human consumption [22,23] that finally failed, and farms were closed but many specimens escaped. Its presence in Thailand was notified in 1982, considered as an invasive species in 1988, and reported from 43 provinces, out of 72, by 1996 [18]. They cause havoc in many rice fields because they damage the young shoots [24]. However, they provide an important source of protein for wild birds and free-range ducks brought by farmers to the rice fields after the harvest. An minimum average of one snail per square meter can be found in this environment (Figure 1a, b) [10,11,25]. They are particularly numerous in house vicinity where they feed on waste, and tolerate polluted environments. They lay their eggs in large quantities on vegetation above water, seen as very characteristic pink clusters (Figure 1c).
For our study, we raised snails from wild eggs in order to have a significant number of adults with known characteristics. We had prepared five medium tanks (50cmx25cmx32cm) containing mineral water (15cm in depth) and the bottom lined with stones, gravel, and sand. The water was oxygenated with a pump to facilitate the development of bacterial flora and nitrogen cycle. After harvesting the eggs in the wild (clusters of hundred eggs attached to a plant stem), they were placed in an aquarium at room temperature (between 28°C and 32°C) above water.
After 2 to 6 days eggs fell naturally into the water where they hatched. Juvenile snails were fed with lettuce, cabbage, and a supplementation protein was added by a provision of commercial fish food. A supply of minerals was also provided by addition of chicken eggshells in the aquarium. Mortality was high in the early days but stabilized quickly (Table 1). After three months of rearing, the snails had reached adult size of 1 to 3 cm.
| Days before infestation | Mortality |
|---|---|
| -60 | 407 (0)a |
| -56 | 395 (8) |
| -53 | 389 (4) |
| -48 | 374 (11) |
| -43 | 349 (16) |
| -38 | 345 (4) |
| -33 | 340 (4) |
| -31 | 337 (4) |
| -14 | 278 (14) |
| -10 | 275 (4) |
a: Total (% mortality)
Table 1: Daily snail mortality during following a two months period before experimental infestation done on the water.
(a) Free grazing wild ducks in rice fields after harvesting (Thailand).
(b) Free grazing semi-domestic ducks in rice fields after harvesting (Thailand).
(c) Apple snail eggs laid on water grass in natura (Thailand).
(d) Pomaceacanaliculata (Apple snail)
Three plastic containers (39x27x18 cm) were introduced into a poultry isolator in ABSL-3 laboratory and prepared one week before the experiment in order to establish abiotic conditions. Each container contained ten liters of mineral water, stones and gravel, with diffusors to oxygenate the water. Non-hermetic lids closed the containers. The hand-raised snails were introduced into two containers (60 specimens per container), one container remaining only with water, without snails. The temperature was set at 30°C, close to natural conditions, and daily checked (Figure 2).
The same dose of H5N1 virions was added to the three containers and, samples taken from water and snails to test for presence of active virus. Snails were dissected to separately collect different organs including, gills, pseudo-lung, intestines and foot.
Samples (water and snails) were cultured by inoculation into the allantoic fluid of embryonated chicken eggs (fertilized and incubated for 9 to 11 days). Allantoic fluid was then collected and tested by Hemaglutination Test (HT) for the presence of the active virus. Quantitative RT-PCR (qRT-PCR) was performed to confirm positive HT and to measure the sample virus concentration.
In nature, it is estimated that infected ducks release 107 TCID50 per ml in their droppings [16]. Under natural conditions, the viral concentration in water after dejection can be estimated between 102and 103TCID50per ml based on the estimation that one dropping (20 to 30 g per day) contaminate between 100 liters (i.e.: in a rice field) and 1000 liters of water in a pond. The viral suspension was of 2.14x106 TCID50/ml, and with each container holding 10 liters of water, we introduced 0.5 ml of viral solution into each container to simulate natural infestation conditions of bird’s droppings after dispersion in water (102TCID50/ml). The air diffuser enabled homogenization of viral concentration in the container.
Every day, 3 ml of water were taken from each container. Water was taken from several locations in each container to optimize the presence of virions in the sample. The samples collected in the two containers with snails are mixed and treated as one single sample. The sample was clarified by centrifugation (8,000xg/10 min/4°C) and then filtered through a membrane of 0.45 µm. A sample of water was also taken before infestation. Sampling stopped when the active virus could not be detected for more than five continuous days1.
Five snails were also collected every two days2. For each snail specimen, mucus was taken from the foot, and then it was washed with distilled water and dissected to remove the organs used for respiration (gills and pseudo-lung) and digestive gut. Snails were killed before dissection by letting them 5’ in freezer. For the isolation of active virus, fragments of organs from the same specimen were put together in 800 µl of 1X PBS, and then ground with a pestle. The homogenate was clarified by centrifugation (8000xg /10 min / 4°C) and the supernatant decanted and filtered through a membrane of 0.45μm.
Samples were inoculated in embryonated chicken eggs incubated for three days before checking the presence of active virus by a HT [26]. Specific detection and titration was performed by qRT-PCR in order to confirm the presence of H5N1 virions and its link with positive HT, and to estimate the concentration of inactivated or not viral particles.
To increase the high sensitivity of the test for the presence of active virus (P>0.95), each water sample was inoculated to four embryonated chicken eggs, as well for snail sample using two embryonated eggs for each sample, leading to an overall sensitivity of the five snails collected with P>0.95. When the outcome was negative, we conducted a new test (blind passage) by inoculating the negative allantoic fluid into a new egg [27]. The result was considered positive when positive HT was observed for at least one of the eggs.
Minimum detectable virus concentration (HT sensitivity) was set up with a P>0.95 of HT after culturing the virus on eggs. For a test using four eggs, the sensitivity u of an egg must be at least 0.52 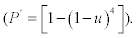 . We evaluated the minimum detectable concentration with a sensitivity>=0.5, by inoculating eggs to assess the percentage of positives. To obtain an accuracy of 0.15 in this assessment, with a type I error of 0.05, it is necessary to inoculate at least 40 eggs 3.
. We evaluated the minimum detectable concentration with a sensitivity>=0.5, by inoculating eggs to assess the percentage of positives. To obtain an accuracy of 0.15 in this assessment, with a type I error of 0.05, it is necessary to inoculate at least 40 eggs 3.
We tested the concentrations 10, 5 and 1 TCID50/ml. The 0.2 ml dose injected into each egg was respectively equal to 2, 1 and 0.2 TCID50. The results are summarized in Table 23.
| TCID50/ml | Positivea | First passageb | Dead after 1st dayc | Dead after 2nd dayd |
|---|---|---|---|---|
| 10 | 26/40 (65) | 21/40 (52) | 1 / 1 | 19/20d |
| 5 | 22/40 (55) | 17/40 (42) | 2 / 2 | 19/20 |
| 1 | 20/40 (50) | 11/40 (27) | 1 / 1 | 7/8 |
(a) positive / total tested (%); (b) Number of positive test the first egg inoculation (over 40 done); (c) Number of dead embryo one day after infection / total tested; (d) Number of positive dead embryo two days after infection / total tested
Table 2: Sensitivity Assessment between Hemaglutination test and RTPCR test.
(a) positive / total tested (%); (b) Number of positive test the first egg inoculation (over 40 done); (c) Number of dead embryo one day after infection / total tested; (d) Number of positive dead embryo two days after infection / total tested
At a starting dose of 2 TCID50 (10 TCID50/ml concentration) and 1 TCID50 (5 TCID50/ml concentration), the virus induced a very high mortality of egg embryos. With a starting dose of 0.2 TCID50 (concentration 1 TCID50/ml), 50% of the results was positive: in the condition of our study, the HT using eggs with two passes was 5 times more accurate than a cell culture test. Test applied to water samples (four eggs with two passes) allowed detecting the virus at a concentration of 10 TCID50/ml with a probability of 0.985, at a concentration of 5 TCID50/ml with a probability of 0.959, and at a concentration of 1 TCID50/ml with a probability of 0.937. The detection test used for snails (two eggs with two passes on five samples) allowed detecting the virus at a concentration 10 TCID50/ml with a probability of 0.995, at a concentration of 5 TCID50/ml with a probability of 0.982, and at a concentration of a 1 TCID50/ml with a probability of 0.969. Virus detection and titration by qRT-PCR were performed on all samples of water and snails, subjected to qRT-PCRto verify the presence of H5N1 viral genetic material and to determine its concentration (H5 Gene [GenBank: EF593102]). The technique required only a minimum of three RNA copies of the target gene [28]. False positives by external contamination may occur. HT gives the best specificity and provides the largest number of true negatives. qRT-PCRallows calculating the concentration of viral particles, active or inactivated. We used the kit SuperScript III OneStep RT-PCR System (Invitrogen®) according to manufacturer's recommendations. The amplicons obtained were analyzed by the program Rotor-Gene Real-Time Analysis Software 6.1®. In the absence of known variability for virus persistence in water, the significance of the difference in virus persistence between the two containers (with and without snails) could be analyzed from a model of virus survival in each container. From the results (water with and without snails), we modeled the probability of the event "presence of active virus" using a negative exponential function and used a Logrank test to compare the two models.
Results
Throughout all experiment, water temperature was maintained between 27°C and 30°C, and dailypH measure remained close to 7 (between 6.7 and 7.8). The Snail mortality of 12 % was recorded during the experiment as weel as before infestation (Table 1). Although, during virus isolation 22 % of embryonated eggs (75 over 339) did not survive the 72 h period post inoculation (embryos died after 24 to 48 hours), 77 % (40 over 52) of the eggs tested positive by HT did not survive the 72 h period.
(a) H = Hemaglutination; (b) copies / ml; (c) PCR = qRT-PCR; (d) + = number of positive embryonated egg (with four tests for each water sample +- one blind passage for each first negative test); (d) - = negative; (e) nt = not tested.
Control samples tested negative prior infestation. Active virus was detected up to 12 days from water without snails, and up to 14 days from water with snails. Over 90% of snail samples collected until seven days after infestation were tested positive and active virus was detected up to 11 days in snails. Of the 39 samples positive by HT, 23 (59%) were positive after two egg passages. Results are presented in Table 3 (water without snails), table 4 (water with snails) and table 5 (snails).
| Day | Container without snails | Container with snails | ||
|---|---|---|---|---|
| Ha | PCR(x106)b,c | H | PCR (x106) | |
| 0 | 0d | nte | 100 | 10 |
| 1 | 100 | 20 | 50 | 3 |
| 2 | 25 | 0.2 | 25 | 0.9 |
| 3 | 25 | 10 | 75 | 0.1 |
| 4 | 25 | 0.03 | 25 | 0.03 |
| 5 | 50 | 0.1 | 50 | 0.03 |
| 6 | 25 | 0 | 0 | 0 |
| 7 | 0 | 0.104 | 25 | 0 |
| 8 | 25 | 0 | 0 | 0.04 |
| 9 | 0 | 0.05 | 0 | 0 |
| 10 | 0 | 0.08 | 0 | -d |
| 11 | 25 | 0 | 0 | - |
| 12 | 25 | 0.1 | 0 | - |
| 13 | 0 | 0 | 0 | - |
| 14 | 0 | 0 | 0 | - |
| 15 | 0 | nt | 0 | - |
| 16 | - | nt | - | - |
| 17 | - | nt | - | - |
Table 3: Virus detection by Hemaglutination and qRT-PCR tests of H5N1 experimentally infested water with and without containing snails
| Day | Snail 1 | Snail 2 | Snail 3 | Snail 4 | Snail 5 | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | +a | 0b | + | 5x104 | + | 7x103 | + | 4x104 | + | 4x104 |
| 3 | + | 104 | + | 0 | + | ncc | + | nc | + | 4x104 |
| 5 | - | 5x104 | + | 10x104 | + | nc | - | nc | + | 15x104 |
| 7 | + | 0 | + | 90x104 | - | nc | - | nc | + | 63x104 |
| 9 | - | 1x104 | - | 1x104 | - | 2x104 | - | nc | - | 3x104 |
| 11 | + | nc | - | 16x104 | - | 1x104 | - | 33x104 | - | nc |
| 13 | - | nt | - | nt | - | nt | - | nt | - | nt |
| 15 | - | nt | - | nt | - | nt | - | nt | - | nt |
(a) HT positive (+) or negative (-); (b) qRT-PCR in genome copies/ml; (c) nc means no concentration detected. nt = not tested.
Table 4: Virus detection by Hemaglutination test and qRT-PCR test of snails sampled during the experiment.
Each snail sample was tested by qRT-PCR (Table 3, 4, 6). Note that for water samples, the observed concentration tends to decrease with time in the container with snails as in the container without snails. Virus particle concentration was detected among 18 snails of 30 tested (60%). Moreover, snail virus concentration tended to increase over time.
Discussion
Although, experimental conditions simulated natural environmental conditions, snail density was higher in the containers than in nature with absence of sunlight and chemical pollutants that could reduce survival of the virus. The low initial infectious viral concentration used in the experiment simulates unfavorable conditions for virus detection. Virus has no lethal effect on snails, while no excess mortality was observed after water infestation by the virus.
Three days culture on embryonated chicken eggs followed by a HT appears as the most sensitive method to detect active influenza virus [26]. Most influenza viruses replicate in less than three days in the allantoic fluid. Although, H5N1 virus induces high embryo mortality [28] as observed in the present study, it replicates actively before the death of the embryo.
Water samples were all positive until six days after infestation. Beside an initial low viral concentration in the containers (102 TCID50/ml), the water sampling optimizes the presence of viral particles containing at best 3x102 TCID50. The presence of active virus particles was observed respectively up to 12 and 14 days after infestation from water sampling without and with snails. Then virus became undetectable with an expected (P> 0.94) concentration below 0.2 TCID50/ml. However, the significance of this difference (12 days vs 14 days) cannot be objectively assessed because we lack knowledge on the variability of virus particle persistence in this type of experiment. In an attempt to develop a model by a negative exponential function, we obtain a function f1(j)=e-0.25(j-1) for the container without snails, and f2(j)=e-0.1875(j-1) for the container with snails and, a logrank test did not give significance to the difference observed (p value = 0.20). Ultimately, results of our study do not show over time a significant influence of apple snails on the persistence of virus in water.
We found active virus in the snail’s samples up to 11 days with an occurrence higher than for water samples. The virus particles may originate from intracellular compartments or from external integuments of the mollusk. The five samples from each session were tested separately (two eggs instead of four). The individual were positive up to seven days after infestation. Altogether, these results suggest that snails may have the ability to capture and concentrate the virus particles from the water and could therefore act as mechanical vectors. They could thus influence virus persistence in the environment, offering protection of viral particles from solar radiation or keeping them in a more favorable physico-chemical environment. Viral concentration in snail could also influence the epidemiology of the H5N1 virus when birds consume virus infested apple snails .Moreover viral RNA concentration detected for 60% of the samples taken from snail organs was higher than those from water samples. In addition, the viral RNA concentration did not decrease throughout time as observed in water samples, and instead tend to increase from five days after infestation. We can assume that the longer the snails are in contact with the virus particles, the more they capture and concentrate viral particles (active or not) in or on their bodies. Also some HT negative samples tested positive by qRT-PCR because PCR detects free viral RNA from inactivated viral particles.
Conclusions
In conclusion, although fresh water apple snails did not actively replicate H5N1, they could host and concentrate the virions over the time and protect it particularly from solar radiation or physico-chemical variations. Ultimately, they could play a role as a source of H5N1 virus in natura: firstly by an active role in concentrating the virus particles already present in water, and secondly, by improving the risk of infection for the birds that consume them. Additional studies are necessary to evaluate the impact of consumption of snails carrying the virus on bird infection.
Authors Contributions
MS and DG conceived and designed the study, and performed interpretation; MS prepared the draft of the manuscript; DG, WP, SC, LS performed manipulations and tests in ABSL3; WW, DS, MG, JPG help designing lab work and provided intellectual discussions; SK and SN were in charge of snail production; JPG and MG provided intensive review of the manuscript; PK contribute to the original idea, review the manuscript and gave a final approval for the manuscript to be submitted for publication. All authors edited and commented on the manuscript.
Acknowledgements
This work was conducted within the Faculty of Science and the Faculty of Veterinary Science of Mahidol University, Thailand. We acknowledge KanapornPoltep and ParutSuksai (Mahidol University) for their help in ABSL-3. We acknowledge Michele Ottman (Université Lyon 1), AnjiKodjo, Amélie Gautier (EcoleVétérinaire de Lyon), Remy Charrel (Université Aix-Marseille 2), Arunee Thitithanyanont and KamlangChumpobanchorn (Mahidol University) for intellectual discussions. This project was funded by ANR (The French Agency for National Research) as part of ECOFLU program (ANR 06SEST12), and by AIRD (Agence Inter-établissements de Recherche pour le Développement) as part of ARDIGRIP program. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
1The probability of not detecting active virus during five consecutive days (water or snail) is inferior to 10-6.
2Pr probability to detect the virus by sampling x snails : 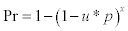 , where p is the probability (unknown) of virus presence in snails and, u the probability of detecting the virus (sensitivity of HA after culturing the virus in embryonated chicken eggs). “u” estimated at 0.75, except false positives (see further in text). For p=0.2 and x=5, Pr = 0.56. For p=0.6 and x=5, Pr = 0.95. For p=0.8 and x=5, Pr = 0.99. For p=1 and x=5, Pr=0.999.
, where p is the probability (unknown) of virus presence in snails and, u the probability of detecting the virus (sensitivity of HA after culturing the virus in embryonated chicken eggs). “u” estimated at 0.75, except false positives (see further in text). For p=0.2 and x=5, Pr = 0.56. For p=0.6 and x=5, Pr = 0.95. For p=0.8 and x=5, Pr = 0.99. For p=1 and x=5, Pr=0.999.
3Size m of the sample involves the accuracy e and the z-value εα corresponding to the risk of error : 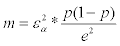
References
- Faust C, Stallknecht D, Swayne D, Brown J (2009) Filter-feeding bivalves can remove avian influenza viruses from water and reduce infectivity. ProcBiolSci 276: 3727-3735.
- Stumpf P, Failing K, Papp T, Nazir J, Böhm R, et al. (2010) Accumulation of a low pathogenic avian influenza virus in zebra mussels (Dreissenapolymorpha). Avian Dis 54: 1183-1190.
- Abbas MD, Nazir J, Stumpf P, Marschang RE (2012) Role of water fleas (Daphnia magna) in the accumulation of avian influenza viruses from the surrounding water. Intervirology 55: 365-371.
- Neumann G, Kawaoka Y (2006) Host range restriction and pathogenicity in the context of influenza pandemic. Emerg Infect Dis 12: 881-886.
- Naffakh N, Manuguerra JC, Van Der Werf S: Grippe : zoonose et transmission inter-espèces. Virologie 2002, 6: S73-S82.
- Bürgmayr S, Grabher-Meier H, Staudacher E (2001) Sialic acids in gastropods. FEBS Lett 508: 95-98.
- Cowie RH: Apple snails (Ampullariidae) as agricultural pests: their biology, impacts and management. In: Molluscs as crop pests. Edited by G.M. Barker, CABI Publishing. Wallingford; 2006, 145-192.
- Souris M, Gonzalez JP, Shanmugasundaram J, Corvest V, Kittayapong P (2010) Retrospective space-time analysis of H5N1 Avian Influenza emergence in Thailand. Int J Health Geogr 9: 3.
- Auewarakul P, The Past and Present Threat of Avian Influenza in Thailand. In Emerging Infections in Asia, Edited by Y. Lu et al. London: Springer, 2008: 31- 44.
- Gilbert M, Chaitaweesub P, Parakamawongsa T, Premashthira S, Tiensin T, et al. (2006) Free-grazing ducks and highly pathogenic avian influenza, Thailand. Emerg Infect Dis 12: 227-234.
- Gilbert M, Xiao X, Chaitaweesub P, Kalpravidh W, Premashthira S, et al. (2007) Avian influenza, domestic ducks and rice agriculture in Thailand. AgricEcosyst Environ 119: 409-415.
- Ito T, Okazaki K, Kawaoka Y, Takada A, Webster RG, et al. (1995) Perpetuation of influenza A viruses in Alaskan waterfowl reservoirs. Arch Virol 140: 1163-1172.
- Brown JD, Swayne DE, Cooper RJ, Burns RE, Stallknecht DE (2007) Persistence of H5 and H7 avian influenza viruses in water. Avian Dis 51: 285-289.
- Stallknecht DE, Brown JD (2009) Tenacity of avian influenza viruses. Rev Sci Tech 28: 59-67.
- Shahid MA, Abubakar M, Hameed S, Hassan S (2009) Avian influenza virus (H5N1); effects of physico-chemical factors on its survival. Virol J 6: 38.
- Webster RG, Yakhno M, Hinshaw VS, Bean WJ, Murti KG (1978) Intestinal influenza: replication and characterization of influenza viruses in ducks. Virology 84: 268-278.
- Siengsanan J, Chaichoune K, Phonaknguen R, Sariya L, Prompiram P, et al. (2009) Comparison of outbreaks of H5N1 highly pathogenic avian influenza in wild birds and poultry in Thailand. J Wildl Dis 45: 740-747.
- Joshi RC: Managing invasive alien mollusk species in rice. International Rice Research Notes, 30(2): 5-13.
- Joshi RC (2007) Problems with the management of the golden apple snail Pomaceacanaliculata: an important exotic pest of rice in Asia. Area-Wide Control of Insect Pests 3: 257-264.
- Estebenet AL, Martín PR (2002) Pomaceacanaliculata (Gastropoda: Ampullariidae): life-history traits and their plasticity. Biocell 26: 83-89.
- Chobchuenchom W, Bhumiratana A (2003) Isolation and characterization of pathogens attacking Pomaceacanaliculata. World Journal of Microbiology & Biotechnology, 19: 903-906.
- Teo SS (2004) Biology of the golden apple snail, Pomaceacanaliculata (Lamarck, 1822), with emphasis on responses to certain environmental conditions in Sabah, Malaysia. Molluscan Research 24: 139-148.
- Tanaka K, Watanabe T, Higuchi H, Miyamoto K, Yusa Y, et al. (1999)Density-dependent growth and reproduction of the apple snail, Pomaceacanaliculata: a density manipulation experiment in a paddy field. Researches on population ecology41: 253-262.
- Kenji I: Expansion of the golden apple snail, Pomaceacanaliculata, and features of its habitat. 2003, http://www.agnet.org/library/eb/540/
- Ichinose K, Wada T, Yusa Y, Kubota T (2000) Influence of habitat differences brought about by environmental changes on the densities of adults and eggs of Pomaceacanaliculata. Proceedings of the Association for Plant Protection of Kyush46: 78-84
- Manuguerra JC, Hannoun C (1999) Influenza and other viral repiratory diseases: surveillance and laboratory diagnosis. Inst Pasteur Coll Lab Refer Exper 186-189
- Manuel terrestre de l’oie, 2005, 2. 1. 14, 290-305.
- Chen W, He B, Li C, Zhang X, Wu W, et al. (2007) Real-time RT-PCR for H5N1 avian influenza A virus detection. J Med Microbiol 56: 603-607.
Relevant Topics
Recommended Journals
Article Tools
Article Usage
- Total views: 16853
- [From(publication date):
March-2015 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 12172
- PDF downloads : 4681


