Research Article Open Access
Separate and Joint Effects of Polycyclic Aromatic Hydrocarbons (PAH)and Polychlorinated Biphenyls (PCB) on Aromatase CYP19A Transcription Level in Atlantic Tomcod (Microgradus tomcod)
Adam Tulu1, Ali Ishaque1*, Egbe Egiebor1, Christopher Chambers2 and Rosemary Jagus3
1Department of Natural Science, University of Maryland, Eastern Shore, Princess Anne, MD 21853, USA
2NOAA-Fisheries, 74 Magruder Rd., Highlands, NJ 07732, USA
3Center of Marine Biotechnology, University of Maryland Biotechnology Institute.701 E Pratt st. Baltimore, MD 21202, USA
- *Corresponding Author:
- Ali Ishaque
Department of Natural Science
University of Maryland, Eastern Shore
Princess Anne, MD 21853, USA
Tel: (410) 651-2200
E-mail: Abishaque@umes.edu Abishaque@umes.edu
Received date: April 29, 2013; Accepted date: July 05, 2014; Published date: July 12, 2014
Citation: Tulu A, Ishaque A, Egiebor E, Chambers C, Jagu R (2014) Separate and Joint Effects of Polycyclic Aromatic Hydrocarbons (PAH) and Polychlorinated Biphenyls (PCB) on Aromatase CYP19A Transcription Level in Atlantic Tomcod (Microgradus tomcod). J Marine Sci Res Dev 4:151. doi:10.4172/2155-9910.1000151
Copyright: © 2014 Tulu A, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Marine Science: Research & Development
Abstract
Ovarian cytochrome 19A (CYP19A) expression is recognized as a useful biomarker for exposure of fish to environmental contaminants such as PAHs and PCBs. In this study, a laboratory approach using Atlantic tomcod (Microgradus tomcod) from the Hudson River was used to evaluate the additive and interactive effects of a PAH (benzo[a]pyrine) and a PCB mixture (Aroclor 1242) with respect to their effects on various metrics of reproduction. The experimental design was a two-way factorial with each treatment at 0, 0.1 and 1 ppm and replicated three times. Fish embryos were subjected to a short term aqueous exposure (DMSO as vehicle) whereas larvae were exposed via repeated feedings of contaminated prey (Artemia). Quantitative reverse transcription-polymerase chain reaction (RT-qPCR) was used to evaluate the mRNA expression level of the ovarian aromatase CYP19A as a biomarker to assess reproductive stress in Microgadus tomcod. When tested alone, expression of aromatase CYP19A was significantly up regulated at the higher level of PCBs but no effect was observed from benzo-a-pyrene (B[a]P). In the PCB/PAH combined treatment, both low-PCB with both low/high levels of PAH and high-PCB with both low/high levels of PAH treatment groups had no significant effect on aromatase CYP19A transcript levels compared with the control group. Gonadosomatic Index (GSI) of reproductively mature females showed that only the high-PCB treatment group exhibited significant gonadal loss. The results confirm the expectation that transcription of both PCB and PAH responsive genes are upregulated since they both exert their toxic effects through the aryl hydrocarbons receptor (AHR) pathway
Keywords
Aromatase; Biomarkers; Biotransformation; Endocrine disruption pollutants
Introduction
The potential adverse effects of environmental contaminants that function as endocrine disruptors have raised concerns in recent years. Endocrine disrupting chemicals (EDCs) in the environment are associated with adverse reproductive and developmental effects in wildlife by mimicing the actions of the female sex hormone estradiol [1]. Polychlorinated biphenyls (PCBs) and polycyclic aromatic hydrocarbons (PAHs) in the environment are associated with potential disruptive effects on vertebrate and invertebrate reproductive activity [2,3]. Many studies have shown that PCBs and PAHs are toxic to fish and other aquatic organisms [4-7]. PCBs and PAHs are thought to have contributed to diseases, deformities, and decline in some wild populations, including fish and birds of the Hudson River [8,9].
PCBs and PAHs can act through both receptor-dependent and receptor-independent (i.e., nonhormonal) pathways. Compared to PCBs, PAHs are biodegraded easily and accumulate in fish tissues in quantities that reflect recent exposure. PAH toxicity depends highly on the chemical structure of the PAH [10-12], and even PAH isomers with the same number of benzene rings may vary from nontoxic to extremely toxic depending on different steric positioning of these benzene rings [13,14]. As PCBs and PAHs are hydrophobic, they are found bound to the organic content of the sediments where they become bioavailable to the benthic organisms and bioaccumulate in the food chain [15]. The principal target organs for PAHs and PCBs are those that produce, regulate, or respond to estrogenic hormones such as: brain, gonad, liver, uterus, mammary, adrenals, prostate, and placenta, as well as organs in the developing embryo or fetus [16]. PCBs and PAHs are known to adversely affect estrogen receptor (ER) function either through direct binding to the receptor or by the activation of other receptor pathways that modulate ER activity [17-19].
Aromatase cytochrome P450 (CYP19A, aromatase), is the terminal enzyme in the steroidogenic pathway [20,21], and is responsible for the conversion of androgens to estrogens [22,23]. The product of the CYP19 gene catalyzes the conversion of androgens to estrogens (specifically, C19-androstenedione into C18-estrone and testosterone into E2) [17]. Its activity is an important modulator of plasma β-estradiol (E2) concentrations and is critical to the regulation of reproductive processes controlled by E2 [24]. Both aromatase CYP19A and E2 concentrations increase simultaneously with seasonal gonadal development [25-27].
In lower vertebrates, aromatase CYP19A is predominantly expressed in the ovary and plays important roles in sex-differentiation and oocyte growth, [28-31]. This enzyme complex is localized in the endoplasmic reticulum of cells in which it is expressed [29,32]. Studies have shown that transcript levels of gonadal aromatase (CYP19A) are increased in association with aromatase enzyme activity during vitellogenesis in Atlantic cod (Microgadus tomcod) [33], rainbow trout [34,35], tilapia [20], and red seabream [36]. Aromatase gene expression has been proposed as a key step of estrogen synthesis for ovarian differentiation [37]. Moreover, plasma E2 is directly related with the transcript levels and enzyme activity of aromatase CYP19A during oocyte development [2]. Studies have shown that activated aryl hydrocarbon receptor (AHR) induces the recruitment of estrogen receptor- (ERa) to AHR-regulated genes and that AHR is recruited to ERα-regulated genes [38-43].
Fish such as Microgadus tomcod are an appropriate model for studying the effects of EDCs because Microgadus tomcod have been exposed to multiple sources of EDCs in their environment including sewage, industrial effluent, and urban and agricultural runoff. The value of using local species and exposing them to two abundant xenobiotics in their natural environment and at ecologically relevant doses is that this allows inspection of possible interactive effects between PCBs and PAHs. Our logic in using two classes of contaminants – both are problematic in the Hudson River – is that such an approach permits a determination of whether the primary effect on the Microgadus tomcod population was mediated by bioaccumulation of PCB, PAH, or both. The laboratory-based study spans most of Microgadus tomcod life cycle, i.e., from embryos to feeding larvae to juveniles to reproductive maturation of adults.
Hence, by using both PCB and PAH on local species, we formulated a picture of the uptake and toxicity of contaminants within this ecosystem. We hypothesized that the effect of PCB and PAH together will be higher than their individual effects on CYP19A transcription and that both chemicals will cause reproductive stress. This study has two main objectives pertaining to the evaluation of the additive and/ or interactive sub lethal effects of early-life exposures of fish to PAHs and PCBs. First, to determine the interactive effects of PCB and PAH exposure on hepatic CYP19A transcript levels of Microgadus tomcod. Second, to quantify the toxic and morphological effects of PAHs and PCBs on the tissue, organ (ovary mass) and whole body size of female adult Microgadus tomcod.
Materials and Methods
Exposure of Microgadus tomcod
All female adult Microgadus tomcod (F0generation, N=6), were collected on the same day in mid-February of 2008 from the Hudson River just prior to their spawning. Fish were transferred to James J. Howard Marine Sciences Laboratory at Sandy Hook where they were maintained at 4ºC in clean laboratory water at salinity of 5 ppt for 21 days. The F1 generation (fertilized eggs) and larvae were relocated to the grow-out static water tanks (40-L) held at 8°C. Salinity was increased incrementally from 5 ppt to 25 ppt in order to mimic the downstream movement of tomcod larvae in nature.
All exposures were conducted in a temperature and light controlled cold room at 6 to 7ºC. During 12 wk post-hatching (wph) period, larvae were exposed to the PAH (benzo[a]pyrene), and the PCB (Aroclor 1242) at 0.1 (low), and 10 (high) times the observed concentration in the Hudson River. For embryo exposures, 100 μl of the high or low stocks was mixed with 100 ml of 5 ppt seawater in 150 ml glass beakers just prior to the addition of embryos. Thus, the final high and low exposures represented concentrations of 359 and 3.59 ng/ml (parts per billion), respectively. When both PAH and PCB were used, the PAH was administered to embryos for 48 h before the exposure to the PCB. The exposure medium was Dimethyl sulfoxide(DMSO) for the embryos and via a contaminated food source (Artemia) for the larvae for 10 sequential feedings. Vehicle control groups were exposed to 0.1% (v/v) acetone (100 μl/100 ml 5 ppt seawater).
At 14 wph, juveniles was transferred to 120-L flow-through tanks, weaned onto frozen foods, and tank temperatures were adjusted weekly to mimic seasonally changes in nature. Lastly, as juveniles achieved a size at which implant tags could be applied (~4 to 6 cm at 16 wph), they were marked with a subcutaneous injection of one of two colors of elastomer at one of six location associated with the dorsal fins (left or right side of anterior, medial, or posterior fins) according to treatment and pooled by replicate into 600-L tanks. The tomcod were fed ad libitum on thawed frozen bloodworms (chironomids), adult Artemia, and squid, and maintained at ambient temperature and salinity until they matured. The experiment was terminated at 36 wpf and maturing adults were sampled. During the rear-out period larvae and juveniles were checked daily for mortalities.
Toxic- interaction scale
A two parameter additive index approach model was used to determine the joint toxic action of PCB and PAH chemicals in a mixture. A Toxic Interaction Scale (S), was defined as (S)=[(Am/ Ai)+(Bm/Bi)], where, Am and Bm= toxicity of PCB and PAH when present in a mixture, and Ai and Bi = toxicity of PCB and PAH when they are tested separately. The values of S was measured in toxic unit (TU), and categorized into three groups: S=1 represents an additive effect, S<1 represent antagonistic effect, and S>1 represents synergistic effect.
Gonadosomatic Index (GSI)
Gonadosomatic index (GSI) quantifies the morphological effects of PAH and PCB on the ovary. GIS is calculated by the ratio of ovary weight to whole body weight; it is indicative of critical reproductive success in the life cycle of Microgadus tomcod population to determine its survival [44]. GSI was used for this study to serve as a timeintegrated indicator of the general wellbeing of Microgadus tomcod gonad development.
Condition Factor (CF)
Condition factor (also known as coefficient of condition, or length-weight factor) was computed to assess the relative robustness, or degree of well-being, of a fish. Condition Factor (CF)=[weight/ (length)] X 100%, where length was measured in centimeters (cm) and weight was measured in grams (g). Growth represents the integration of feeding, assimilation and energy expenditure over a period of time. Poor growth means less energy was available for reproduction, which will in turn reduce the species fitness and lead to a decline in the population. A high condition factor, therefore, reflects good environmental quality; while a low condition factor reflects poor environmental quality.
Aromatase CYP19A Transcription as an Indirect Assessment of E2 levels
In order to measure the mean difference of aromatase CYP19A transcription as indirect assessment of E2 level in Microgadus tomcod ovaries, three distinct phases of adult female Microgadus tomcod reproductive activity were evaluated. First, spawning stage was defined as fish whose ovaries are well developed and near peak mass, in which eggs are extruded from the body under slight pressure or when ovary sac is broken, and eggs were easily separated from each other. Second, matured stage was defined as fish whose ovaries had passed the peak spawning stage and begun to regress. At this stage, the ovaries contain less than 50% scattered clear eggs, and the eggs cannot be easily separated from each other. Third, spent stage was defined as fish whose the ovary had fully regressed and contained less than 5% white dry ova and reddish unspawned scattered eggs.
Isolation of Microgadus Tomcod CYP19A Complementary DNA (cDNA)
Approximately 100 mg of ovarian tissue were used as starting material for total RNA isolation with Ambion Pure Link total RNA extraction mini kit (Cat # 12183-018A) following the manufacture’s protocol. All RNA preparations were adjusted to 0.5 μg/μl and used as template for reverse transcription (RT). In all reactions, cDNA was synthesized using 1 μg of total RNA with 100 ng random hexamer primers(10 μl total volume), following the manufacturer’s instructions (Clontech). Reactions were terminated by heating at 70°C for 10min, then diluted 1:5 with DNase-free water and used as template for polymerase chain reactions (PCR).
Degenerate primers for Microgadus tomcod CYP19A PCR amplification were designed based on CYP19A cDNA sequences from the closely related Gadusmorhua along with six other most closely related teleost species (Table 1). Gadusmorhua, Mugilcephalus, Epinepheluscoioides, Epinephelusakaara, Latescalcarifer, Larimichthyscrocea, and Micropogoniasundulatus (Accession #s: DQ402370, AY859425, AY510711, AY547354, AY684256, FJ800566, and DQ184486 respectively) were selected from the NCBI database. Standard group codes were used to represent degenerate primers (W=A+C; S=G+C; R=A+G; K=G+T; V=A+G+C; Y=C+T).
Cloning and Sequencing of CYP19A Partial cDNA
The PCR amplification cycles were varied based on expected product size and the primer melting temperatures. A major RT-PCR product of approximately 1000 bp was isolated from 1.5% agarose/ethidium bromide (EtBr) gels and ligated into pGEM-T vector (Invitrogen). Sequencing was arranged using 250 to 400 ng concentration of plasmid DNA per 8 μl total volume on the ABI 3730xl 96-capillary automated DNA sequencers (Brisbane, Australia). Both strands of plasmid DNA containing the cDNA insert were sequenced in both directions using T7 (forward) and SP6 (reverse) sequencing primers. Thereafter, all positive sequence results were aligned with the CLUSTAL W method to generate a contig sequence using Mega4 software (http://www.megasoftware.net/mega4/.html).
Rapid Amplification of cDNA Ends (RACE)
Rapid amplifcation of cDNA ends (RACE)-ready cDNA libraries were generated using Microgadus tomcod ovarian total RNA and a commercial kit Clonetech SMARTer RACE cDNA Amplification Kit (Cat. Nos. 634923, CA, USA), following the manufacturer’s protocol. Microgadus tomcod CYP19A specific primers for 3’ RACE, outer primer: 5’-TGGTGGACGTCTCCAACACCCTCTTC -3’ and innerprimer: 5’- CTCAGGAGCTGCAGGACGCCATAGA -3’ were used. For nested PCR and for 5’ RACE, outer primer 5’-CGGTGCATGCGTCCCACGTT - 3’ and inner primer 5’- CTCAGGAGCTGCAGGACGCCATAGA -3’ were used. Primers were adjusted to 10 pmol for both 5’ and 3’ RACE reactions. Thermal parameters for RACE touchdown PCR were: 95 °C for 2 min; 5 cycles of 95°C for 5 s and 72°C for 180 sec; 5 cycles of 95°C for 5 sec, 70°C for 10 sec and 72°C for 180 sec; 25 cycles of 95°C for 5 sec, 68°C for 10 sec and 72°C for 180 sec using the Biorad i-cycler thermal cycler. Thermal parameters for RACE Nested PCR were: 95°C for 2 min; 35 cycles of 95° C for 30 sec, 55°C for 30 sec, and 72 min, followed by 72°C for 10 min final extension using the Biorad i-Cyclear thermal cycler. PCR products were gel purified as stated above and ligated into pGEM-T vector (Invitrogen) per the manufacturer’s protocol. Confirmation of the identified nucleotide sequence was achieved by comparing Microgadus tomcod CYP19 cDNA to Gadusmorhua aromatase CYP19A sequence using the Sequencer TM 4.0 Windows Version Package (Gene Codes Corporation).
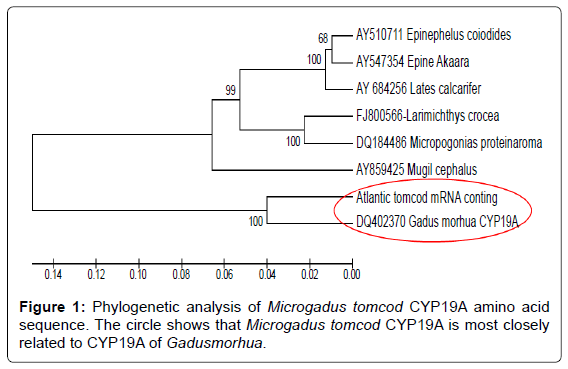
Deduced amino acid comparison was performed using the CLUSTAL W multiple sequence alignment aligorithm and a molecular phylogenetic analysis was performed using the neighbor joining method in MEGA version 5. To determine its relation to other aromatase proteins, the deduced Microgadus tomcod CYP19A protein was included in a molecular phylogenetic analysis with six other teleost sequences (Accession #s: DQ402370, AY859425, AY510711, AY547354, AY684256, FJ800566, and DQ184486), (Figure 1).
Real Time Quantitative PCR (RT-qPCR) Primers
To prevent genomic DNA (gDNA) amplification in RT-qPCR applications, primers were designed to anneal across a unique cDNA regions of splice junction. Primers spanning exon-exon junctions were designed to cross exon 8 and exon 9 boundaries by viewing the detailed alignment of the Microgadus tomcod CYP19A cDNA query sequence against zebra fish CYP19A genomic sequence [22]. Since CYP19A is a highly conserved gene in all vertebrates, the exon-exon boundaries of tomcod and zebra fish can be expected to be the same. Forward: 5’- TCCGGCAGTGTGTGC TGGAGATG -3’ and reverse: 5’- CCGGAGCTTCCTCTCGCCTATGA -3’ oligonucleotides were used for RT-qPCR. The validity of results obtained using RT-qPCR was improved by including Microgadus tomcod S2 ribosomal protein (S2RP) as a reference gene to correct for sample to sample variations in, RT-qPCR efficiency and errors in sample quantification (NCBI database Accession # AY292477). Forward- 5’-AGGGCTGGTCAGCGTACA-3’- and reverse -5’-CTGGAACGATGGACAGCTTG-3’- primers for S2RP were designed using primer3 software (Whitehead Institute, Cambridge, MA).
Real Time RT-PCR Analyses (TaqMan)
Real-time RT-PCR using Microgadus tomcod ovary was performed to determine abundance for aromatase CYP19A mRNA transcription. cDNA was produced using 500 ng total RNA primed with 100 ng random hexamer oligonucleotides and used as template for RT reactions. RT reactions were diluted (1:5) with DNase-free water and 2 μλ of each diluted RT was mixed with 23 μλ of a master mix containing a commercial qRT-PCR enzyme mix (Taqman). Each 25 μλ PCR reaction was set up as duplex with the threshold cycle (Ct) for the gene of interest (GOI) and the reference (REF) measured at the same time to minimize differences in reaction efficiency. The standard curve for CYP19A was generated using an in vitro transcribed Microgadus tomcod aromatase CYP19A plasmid DNA as described above. A serial dilution was prepared to be targeted in the range of 108 to 102 copies for the standard curve was prepared. Real Time RT-qPCR was performed on a BIORAD®iCycleriQ (Catalog number 170-8740). For each set of PCR analyses, the transcription levels were determined in a “standard sample” to determine interassay variability.
Statistical Analysis
The experimental design employed was a 3 x 3 factorial with 3 replicates per treatment combination. Equal variance and normality were checked and analyzed on the mean value by analysis of variance (ANOVA) with factorial design followed by Tukey’s pairwise comparisons. Polynomial contrasts were used for the actual spacing of the treatment levels for determining the existence and nature of trends in the treatment level means. The sums of squares for linear (degree 1), quadratic (degree 2), and cubic (degree 3) trends were computed if needed. Statistical analysis was performed using the statistix package, analytical software 9.0 (Tallahassee, FL 32317).
Results
Isolation of Microgadus tomcod CYP19A cDNA
The deduced cDNA and amino acid (AA) sequences of the isolated Microgadus tomcod aromatase CYP19A cDNA are presented in (Figure 2) Initially, a central part of the mRNA was amplified from degenerate primers and an ovarian total RNA. The sequence of this 1020 bp product was similar to other teleost CYP19A cDNA sequences (BLAST analysis, http://www.ncbi.nlm.nih.gov/blast/; data not shown). It is known that some genes are easily cloned as full-length forms and some are not. RACE reaction using Microgadus tomcod aromatase specific primers allowed identification of a 1415 bpcDNA, which encodes a gene product with a predicted molecular weight of 52.5, representing an almost complete CYP19A sequence. The deduced amino acid sequence of Microgadus tomcod CYP19A was aligned with G. moruha (DQ402370), E. coioides (AY859425), L. calcarifer (AY510711), and M.cephalus (AY547354) (Figure 2). The 1342 bp ORF encodes a protein of 521 amino acids which covers 96% of the ORF size compared with the cDNA for Atlantic cod and fully aligns with the other teleost CYP19A proteins. The sequence shares 86% identity with Atlantic cod and 65–78% identity with the other selected fish species CYP19A sequence. Figure 3 represent multiple alignments of Microgadus tomcod CYP19A mRNA with other teleost aroamatase CYP19A mRNA sequences.
Comparison of aromatase transcription levels in Microgadus tomcod reproductive development cycle
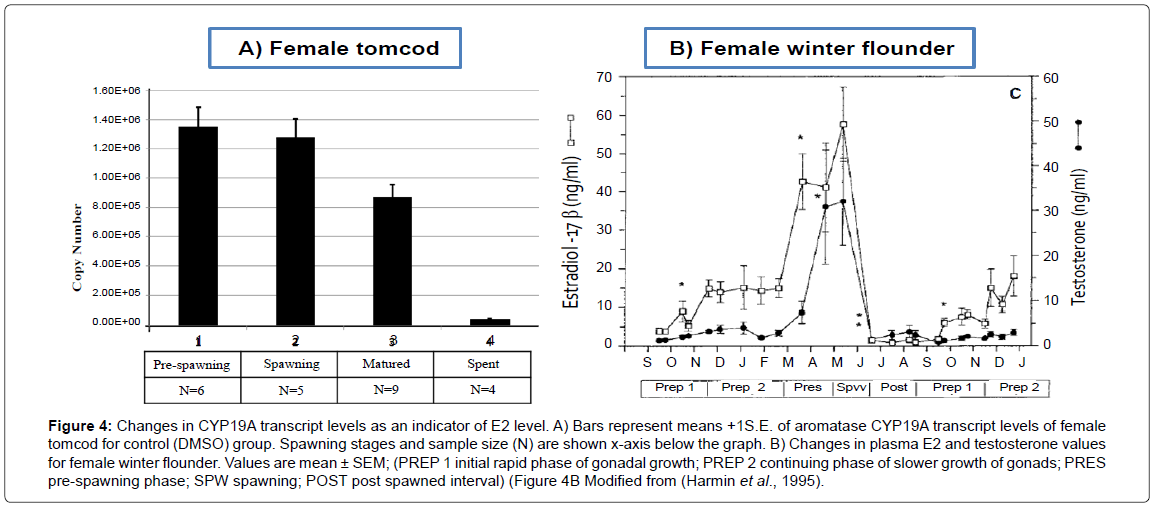
Changes in CYP19A mRNA as an indirect measure of E2 level in the control group and on PCB/ PAH separately treated groups were evaluated on Figures 4 and 5, respectively. Both the control group and PCB / PAH separately treated groups showed the highest aromatase CYP19A mRNA level at spawing, followed by matured and the lowest at the spent stage of the reproduction cycle. Neither PCB nor PAH affected the expected CYP19A transcription pattern, (Spawing>matured>spent). This finding suggested that the level of E2 steroid for PCB or PAH individually treated groups was at the highest seasonal values during pre-spawing and spawning stage. After spawning, the ovaries were regressed and the E2 level in the plasma continued to fall to very low levels.
Figure 4: Changes in CYP19A transcript levels as an indicator of E2 level. A) Bars represent means +1S.E. of aromatase CYP19A transcript levels of female tomcod for control (DMSO) group. Spawning stages and sample size (N) are shown x-axis below the graph. B) Changes in plasma E2 and testosterone values for female winter flounder. Values are mean ± SEM; (PREP 1 initial rapid phase of gonadal growth; PREP 2 continuing phase of slower growth of gonads; PRES pre-spawning phase; SPW spawning; POST post spawned interval) (Figure 4B Modified from (Harmin et al., 1995).
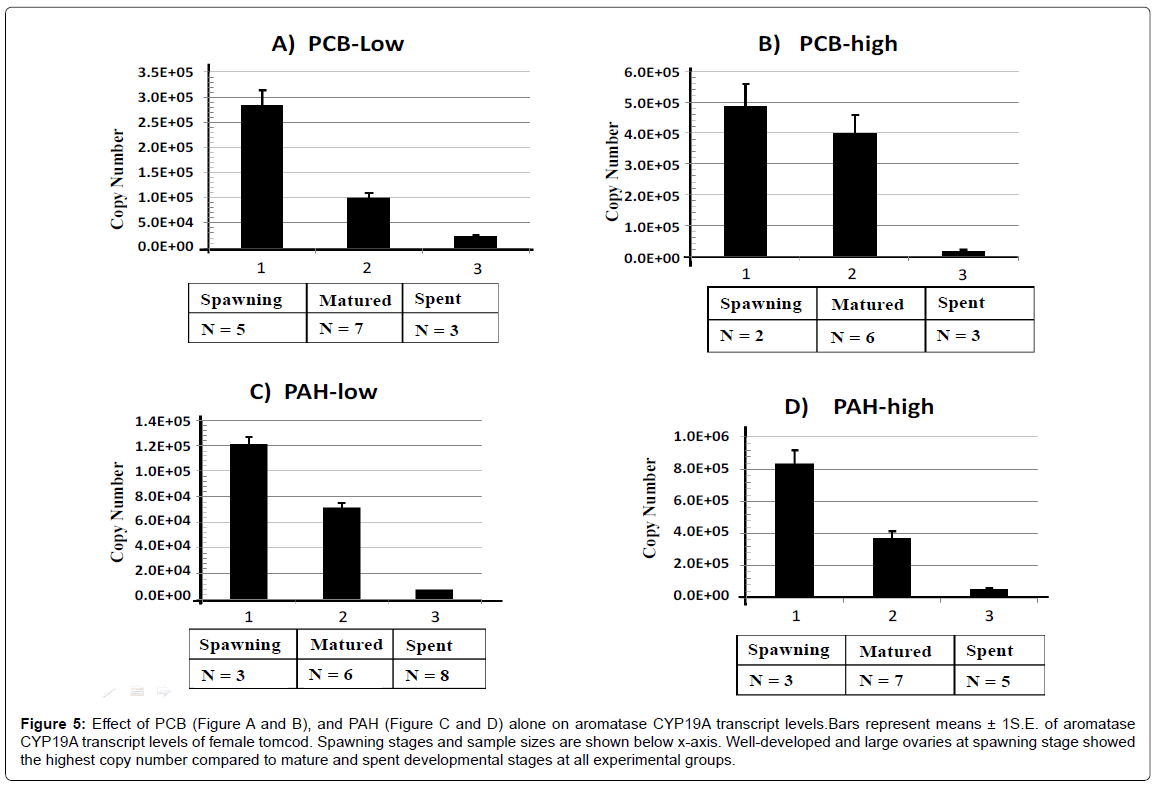
Figure 5: Effect of PCB (Figure A and B), and PAH (Figure C and D) alone on aromatase CYP19A transcript levels.Bars represent means ± 1S.E. of aromatase CYP19A transcript levels of female tomcod. Spawning stages and sample sizes are shown below x-axis. Well-developed and large ovaries at spawning stage showed the highest copy number compared to mature and spent developmental stages at all experimental groups.
Joint Effect of PCB and PAH on Aromatase CYP19A Transcript Level
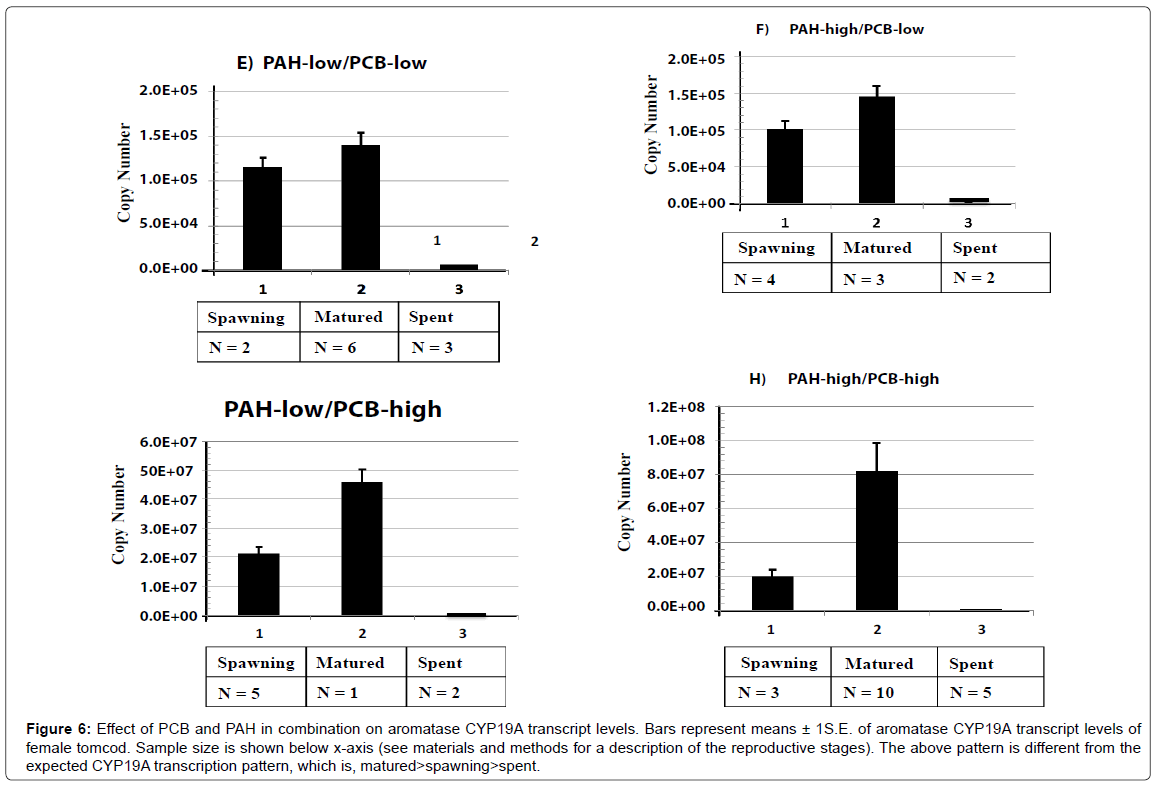
Treatment groups exposed to PCB and PAH in combination showed the highest level of aromatase CYP19A transcript at matured stage, followed by spawning stage and lastly spent stage of reproduction cycle (matured>spawning>spent). Due to the additive affect of PCB and PAH on the combined treatment groups, the level of CYP19A mRNA and E2 production were disrupted (Figure 6).
Figure 6: Effect of PCB and PAH in combination on aromatase CYP19A transcript levels. Bars represent means ± 1S.E. of aromatase CYP19A transcript levels of female tomcod. Sample size is shown below x-axis (see materials and methods for a description of the reproductive stages). The above pattern is different from the expected CYP19A transcription pattern, which is, matured>spawning>spent.
Analysis of CYP19A Transcript Level Using Absolute Quantification
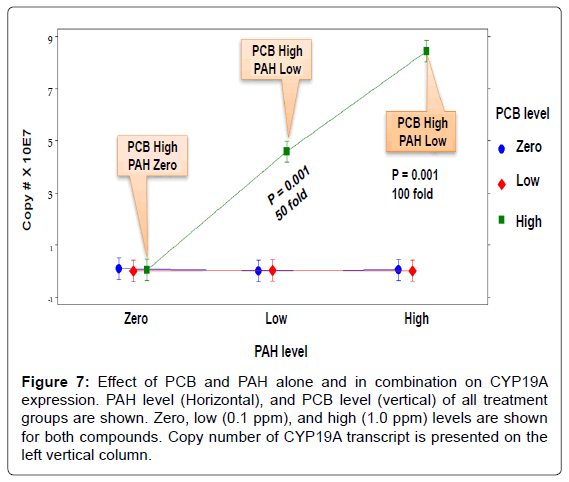
The interaction effect of PCB and PAH on CYP19A transcripts when reproductively mature female Microgadus tomcod aromatase CYP19A mRNA was analyzed using RT-qPCR assay (Figure 7). For individually treated groups, neither PCB nor PAH showed a significant effect on aromatase CYP19A mRNA level. For groups treated with both PCB and PAH, PCB significantly upregulated aromatase CYP19A transcript level at high-PCB/high-PAH (100 fold), and high-PCB/low-PAH, (50 fold) treatment groups. At the highest level of PCB, increasing the levels of PAH increased CYP19A mRNA expression because the toxicity of PCB was enhanced in the presence of PAH. In contrast, at zero and low levels of PCB, increasing level of PAH had no significant effect on CYP19A mRNA expression level. The toxic interaction effects (S) of PCB and PAH mixtures were mainly synergistic (S=1.9 to S=755 TU). The toxic interaction effects of the low-PCB/low-PAH and low- PCB/high-PAH treatment groups (4.5 and 1.9 TU, respectively) were not significantly high compared to those of the high-PCB/low-PAH, (437.9 TU), and high-PCB/high-PAH (754.5 TU) treatment groups.
Figure 7: Effect of PCB and PAH alone and in combination on CYP19A expression. PAH level (Horizontal), and PCB level (vertical) of all treatment groups are shown. Zero, low (0.1 ppm), and high (1.0 ppm) levels are shown for both compounds. Copy number of CYP19A transcript is presented on the left vertical column.
Gonadosomatic Index (GSI)
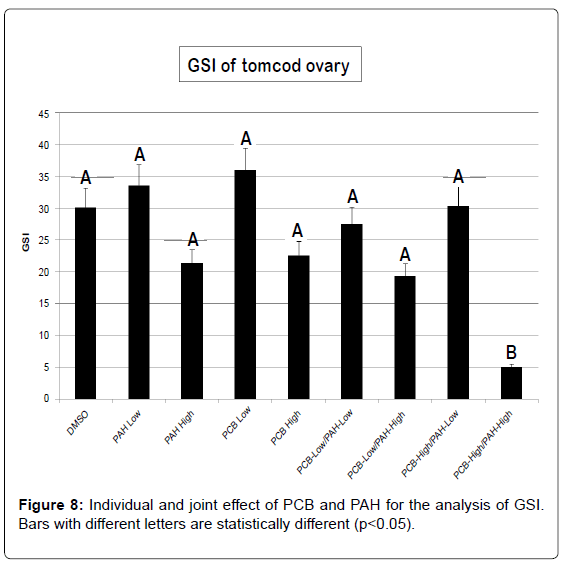
For either PCB or PAH individually treated groups, a significant gonadal loss occurred only in the high-PCB/high-PAH level of treatment (Figure 8). GSI did not change with low-PCB, and low or high-PAH individually treated groups compared to the control group. Exposure to estrogens mimicking compounds such as PAH and PCB can have various detrimental effects in fish. It can reduce general viability, induce gonadal malformations or feminization of genetic males, or lead to sterilization [45]. Even though there was no significant GSI for most treatment groups; we observed reproductive impairment ranging from underdeveloped gonads to decrease egg production and abnormal larval offspring. Morphological analysis was not conducted since it was beyond the scope of this study. However, abnormal morphological variation such as aplasias, atrophy, fusions, degeneration and intersex due to environmental pollutants need further study.
Condition Factor (CF)
The result of this study showed that there was no statistically significant difference in CF due to PCB or PAH exposure in any treatment group. There was also no significant PCB/PAH interaction observed in any treatment group. CF and the length-weight relationship comparison conclude that neither PCB nor PAH influence the wellbeing of fish reproductive cycle.
Discussion
This study demonstrated major differences in the way Microgadus tomcod CYP19A mRNA response to the effects of PCB and PAH at molecular and organismal levels. The individual effect of either PCB or PAH on Microgadus tomcod ovaries are not significant, but PCB caused higher CYP19A transcript levels in the PCB/PAH joint treatment groups. Moreover, low levels of aromatase CYP19A mRNA and lower GSI during the spawning stage of reproductive cycle was resulting from the additive effect of high-PCB and high-PAH. Previous studies on various teleosts found E2, testosterone, GSI, and oocyte diameter to peak during pre-spawning and spawning stages, and to be low or undetectable at spent stage [25,46-48]. This finding suggested that the level of CYP19A mRNA was compromised by the additive effect of PCB and PAH at any treatment group.
The results of all CYP19A transcription level, GSI, and CF tests revealed little or no significant effect of PAH exposure, suggesting that Microgadus tomcod from the Hudson River might have developed a protective mechanism against PAH. This result may be explained by the fact that PCBs caused reproductive impairment by reducing aromatase expression whereas the effect of PAH was reduced or compromised in Microgadus tomcod from the Hudson River.
Both PCBs and PAHs are known to mimic sexual hormones and/or disrupt steroidogenic enzyme function, [44,49], including aromatase CYP19A [22,50-52], PCBs and PAHs binding to the Aryl hydrocarbon receptor (AhR) can result in synergestic effects due to cross-talk between the AhR and the ER [53,54]. PCBs have also been shown to alter or inhibit hormone production by steroidogenic tissue in reproductive organs [16,55].
AhR is known to be involved in modulating estrogen-dependent transcription [34,39,56,57], to control target-specific regulation of ER [37,58], and to have crosstalk with several other important signal transduction pathways [9,24]. Some species are inherently capable of tolerating a contaminant through existing physiological acclimation mechanisms [59-61]. Previous exposure of environmental contaminants most likely decreases the sensitivity of the AhR pathway in Hudson River Microgadus tomcod population. The recent finding by Wirgin et al., [3] that a six-base deletion in the AhRreseptor 2 (AhR2) of Hudson River Microgadus tomcod is ths basis of six-base deletion as the bases for resistance. The finding suggests that the Hudson River Microgadus tomcod population has undergone rapid evolution, probably due to contaminant exposure. This finding of the basis of resistance in a vertebrate population provides evidence of evolutionary change due to selective pressure at a single locus. There is a possibility that Microgadus tomcod may instead develop a tolerance defense mechanism by which Microgadus tomcod do not reduce PAH toxicity or its action, but instead alleviate the negative consequences of such contaminats and action [62,63]. Previous studies have shown that wild caught Microgadus tomcod from the Hudson River have been developing various stages of resistance compared to Atlantic tomcod from relatively uncontaminated tributaries [1,3,9,61,64]. Tolerance to contaminants within cellular organelles is possible by preventing toxicants from exerting their toxic effects despite their elevated presence in receptor tissues. Further investigations are required to determine if resistance may be due to genetic or epigenetic modifications on Microgadus tomcod from Hudson River.
When EPA banded the production of PCB in 1977, it was estimated that 1.3 million pounds of PCBs were discharged in the Hudson River from two General Electric capacitor manufacture plants located in the town of Fort Edward and Hudson Falls, New York. This study supports the idea that resistance to pollution is a recent phenotype in Microgadus tomcod from the Huddson River, suggesting that Microgadus tomcod have rapid evolutionary responses with respect to their environment. Besides, future study is needed to determine if resistance is a common or rare response of chronically exposed fish populations with different selective pressure mechanisms. An important question is, how much does resistance among Microgadus tomcod is matter to the ecology of the contaminated Hudson River? Is an adapted population necessarily an “unhealthy” one? Whether resistance evolves or the population is eliminated depends on the rapidity of onset and severity of the stress, and the capacity of the population to adapt to it. Answering these difficult questions will require to be addressed by moving resistance studies beyond the species-contaminant pairings method. The evolutionary costs of resistance to Hudson River populations or to their communities remain to be experimentally determined.
In general, the polluted sites of the Hudson River may become well known for PCBs and PAHs contaminants, but it is rare that these compounds were released into the environment by industry or municipalities without the release of many other compounds. This issue may be particularly critical for a species such as Microgadus tomcod that appears to serve a unique position in estuarine food chains. Thus, exposures to single or two contaminants can be made in the laboratory, but their results are often confounded by the unknown effects of preexposures to a great variety of other contaminants in wild population. It is believed that studies of two major contaminants are the appropriate departures for future resistance research. The possibility exists that resistance is fostered by synergies among contaminants, at least in some instances. Future analysis will address whether AhR is potentially responsible for conferring resistance in population adapted to chronic exposure to chemical pollutants in the different superfound sites.
Acknowledgements
This research was supported and funded by National Oceanic and Atmospheric Administration (NOAA) – Living Marine Resources Cooperative Science (LMRCS). We thank to all of those at The Institute of Marine and Environmental Technology (IMET) at University of Maryland Center of Environmental Science (UMCES) and the James J. Howard Marine Sciences Laboratory at Sandy Hook.
References
- Wirgin I, Kreamer G, Grunwald C, Squibb K, Garte SJ, et al. (1992) Effects of prior exposure history on cytochrome P450IA mRNA induction by PCB congener 77 in Atlantic tomcod. Marine Environmental Research 34: 103-108.
- Swedenborg E, Pongratz I (2010) AhR and ARNT modulate ER signaling. Toxicology 268: 132-138.
- Wirgin I, Roy KN, Loftus M, Chambers RC, Franks D, et al. (2011) Mechanistic Basis of Resistance to PCBs in Atlantic Tomcod from the Hudson River. Science 331: 1322-1325.
- Bunton TE (1996) Experimental chemical carcinogenesis in fish. ToxicolPathol 24: 603-618.
- Chen S, Zhang F, Sherman MA, Kijima I, Cho M, et al. (2003) Structure–function studies of aromatase and its inhibitors: a progress report. J Steroid BiochemMolBiol 86: 231-237.
- Anderson JW, Lee RF (2006) Use of biomarkers in oil spill risk assessment in themarine environment." Human and Ecological Risk Assessment 12: 1992-1222.
- Buryskova B, Hilscherova K, Blaha L, Marsalek B, Holoubek I (2006) Toxicity and modulations of biomarkers in Xenopuslaevis embryos exposed to polycyclic aromatic hydrocarbons and their N-heterocyclic derivatives. Environ Toxicol21: 590-598.
- Laukers JD (1986) Disposal and destruction of waste PCBs. CRC Press: 83-152.
- Wirgin I, Chambers RC (2006) Atlantic tomcod Microgadus tomcod:A model of responses of Hudson River fish to toxicants. AFS Symposium 51: 331-365.
- Oost Ron, van der b, Jonny Beyer, Nico Vermeulen (2003) Fish bioaccumulation and biomarkers in environmental risk assessment: a review. Environmental Toxicology and Pharmacology 13: 57-149.
- Chavan H, Krishnamurthy P (2012) Polycyclic aromatic hydrocarbons (PAHs) mediate transcriptional activation of the ATP binding cassette transporter ABCB6 gene via the aryl hydrocarbon receptor (AhR)." J Biol Chem 287: 32054-32068.
- Liao C, Lv J, Fu J, Zhao Z, Liu F, et al. (2012) Occurrence and profiles of polycyclic aromatic hydrocarbons (PAHs), polychlorinated biphenyls (PCBs) and organochlorine pesticides (OCPs) in soils from a typical e-waste recycling area in Southeast China. Int J Environ Health Res 22: 317-330.
- Aas E, Beyer J, Jonsson G, Reichert L, Andersen O (2001) Evidence of uptake, biotransformation and DNA binding of polyaromatic hydrocarbons in Atlantic cod and corkwing wrasse caught in the vicinity of an aluminium works. Mar Environ Res 52: 213-229.
- Wurl O, Obbard JP (2004) A review of pollutants in the sea-surface microlayer (SML): a unique habitat for marine organisms.Mar Pollut Bull 48: 1016-1030.
- Lee J, Choe E (2009) Effects of phosphatidylcholine and phosphatidylethanolamine on thephotooxidation of canola oil. J Food Sci 74: C481-c486.
- Yan X, Richard M, Yu K, Zhang X, Giesy Margaret JG, et al. (2006) Effects of PCBs and MeSO2-PCBs on adrenocortical steroidogenesis in H295R human adrenocortical carcinoma cells. Chemosphere 63: 772-784.
- Simpson ER,Mahendroo MS, Means GD, Kilgore MW,Hinshelwood MM, et al. (1994) Aromatase cytochrome P450, the enzyme responsible for estrogen biosynthesis." Endocr Rev 15: 342-355.
- Frye CA, Bo E, Calamandrei G, Calzà L, Dessì-Fulgheri F, et al. (2012) Endocrine disrupters: a review of some sources, effects, and mechanisms of actions on behaviour and neuroendocrine systems. J Neuroendocrinol 24: 144-159.
- Stasinakis AS, Mermigka S, Samaras VG, Farmaki E, Thomaidis NS (2012) Occurrence of endocrine disrupters and selected pharmaceuticals in Aisonas River (Greece) and environmental risk assessment using hazard indexes. Environ SciPollut Res Int 19: 1574-1583.
- Chang XT, Kobayashi T, Kajiura H, Nakamura M, Nagahama Y (1997) Isolation and characterization of the cDNA encoding the tilapia (Oreochromisniloticus) cytochrome P450 aromatase (P450arom) changes in P450arom mRNA, protein and enzyme activity in ovarian follicles during oogenesis. J MolEndocrinol18: 57-66.
- Stocco C (2008) Aromatase expression in the ovary: Hormonal and molecular regulation. Steroids 73: 473-487.
- Trant JM, Gavasso S, Ackers J, Chung BC, Place AR (2011) Developmental expression of cytochrome P450 aromatase genes (CYP19a and CYP19b) in zebrafish fry (Daniorerio). J ExpZool 290: 475-483.
- Lara LA, Duarte AA, Reis RM, Vieira CS, Rosa-e-Silva AC (2012) Endocrine disrupters: potential risk factors affecting sexual function in both men and women. J Sex Med 9: 941-942.
- Carlson AE, Roy KN, Wirgin II (2009) Microarray Analysis of Polychlorinated Biphenyl Mixture-Induced Changes in Gene Expression Among Atlantic Tomcod Populations Displaying Differential Sensitivity to Halogenated Aromatic Hydrocarbons. Environ ToxicolChem 28: 759-771.
- Harmin SA, Crim LW, Wiegand DM (1995) Plasma sex steroid profiles and the seasonal reproductive cycle in male and female winter flounder, Pleuronectesamericanus. Marine Biology 21: 601-610.
- Shozu M, Zhao Y, Bulun SE, Simpson ER (1998) Multiple splicing events involved in regulation of human aromatase expression by a novel promoter. Endocrinology 139: 1610-1617.
- Maranghi F, Mantovani A (2012) Targeted toxicological testing to investigate the role of endocrine disrupters in puberty disorders. Reprod Toxicol 33: 290-296.
- Young G, Kagawa H, Nagahama Y (1983) Evidence for a decrease in aromatase activity in the granulosa cells of amago salmon (Oncorhynchusrhodurus) associated with final oocyte maturation. Biol Reprod 29: 310-315.
- Gelinas D, Pitoc GA, Callard GV (1998) Isolation of a goldfish brain cytochrome P450 aromatase cDNA: mRNA expression splicing events involved in regulation of human aromatase expression by a novel promoter. Endocrinology 139: 1610-1617.
- Tchoudakova A, Kishida M, Wood E, Callard GV (2001) Promoter characteristics of two cyp19 genes differentially expressed in the brain and ovary of teleost fish. J Steroid Biochem MolBiol 78: 427-439.
- Doering JA, Giesy JP, Wiseman S, Hecker M (2013) Predicting the sensitivity of fishes to dioxin-like compounds: possible role of the aryl hydrocarbon receptor (AhR) ligand binding domain. Environ SciPollut Res Int 20: 1219-1224.
- KazetoK, Place A, Trant MJ (2004) Effects of endocrine disrupting chemicals on the expression of CYP19 genes in zebrafish (Daniorerio) juveniles. Aquat Toxicol 69: 25-34.
- Kortner T, Arukwem A (2007) Effects of 17α-methyltestosterone exposure on steroidogenesis and cyclin-B mRNA expression in previtellogenic oocytes of Atlantic cod (Gadusmorhua). Comparative Biochemistry and Physiology 146: 569-580.
- Tanaka M, Telecky TM,Fukada S, Adachi S, Chen S, et al. (1992) Cloning and sequence analysis of the cDNA encoding P-450 aromatase (P450arom) from a rainbow trout (Oncorhynchusmykiss) ovary; relationship between the amount of P450arom mRNA and the production of oestradiol-17ß in the ovary." J Mol Endocrinol 8: 53-61.
- Fukada S, Tanaka M, Matsuyama M, Kobayashi D, Nagahama Y (1996) Isolation, characterization, and expression of cDNAs encoding the medaka (Oryziaslatipes) ovarian follicle cytochrome P-450 aromatase." Mol Reprod Dev 45: 285-290.
- Gen K, Okuzawa K, Kumakura N, Yamaguchi S, Kagawa H (2001) Correlation between messenger RNA expression of cytochrome P450 aromatase and its enzyme activity during oocyte development in the red seabream (Pagrus major). Biol Reprod65: 1186–1194.
- Heid SE, Pollenz RS, Swanson HI (2000) Role of heat shock protein 90 dissociation in mediating agonist-induced activation of the aryl hydrocarbon receptor. Mol Pharmacol 57: 82-92.
- Tanguay RL, Abnet CC, Heideman W, Peterson R. E (1999) Cloning and characterization of the zebrafish (Daniorerio) aryl hydrocarbon receptor. Biochim Biophys Acta 1444: 35-48.
- Barouki R, Aggerbeck M, Aggerbeck L, Coumoul X (2012) The aryl hydrocarbon receptor system. Drug Metabol Drug Interact 27: 3-8.
- Kiss EA, Diefenbach A (2012) Role of the Aryl Hydrocarbon Receptor in Controlling Maintenance and Functional Programs of RORgammat(+) Innate Lymphoid Cells and Intraepithelial Lymphocytes." Front Immunol 3: 124.
- MayatiA, Le Ferrec E, Lagadic-Gossmann D, Fardel O (2012) Aryl hydrocarbon receptor-independent up-regulation of intracellular calcium concentration by environmental polycyclic aromatic hydrocarbons in human endothelial HMEC-1 cells. Environ Toxicol 27: 556-562.
- He J, Hu B, Shi X, Weidert ER, Lu P, et al. (2013) Activation of the aryl hydrocarbon receptor sensitizes mice to non-alcoholic steatohepatitis by deactivating the mitochondrial sirtuindeacetylase Sirt3. Mol Cell Biol 33: 2047-2055.
- Mittelholzer C, Andersson E, Consten D, Hirai T, Nagahama Y, et al. (2007) 20b-hydroxysteroid dehydrogenase and CYP19A1 are differentially expressed during maturation in Atlantic cod (Gadusmorhua). J Mol Endocrinol39: 319-328.
- Hoffmann LJ, Oris TJ (2006) Altered gene expression: A mechanism for reproductive toxicity in zebrafish exposed to benzo[a]pyrene. Aquat Toxicol78: 332-340.
- Senior AM, Nakagawa S (2013) A comparative analysis of chemically induced sex reversal in teleosts: challenging conventional suppositions. Fish Fisheries 14:60-76.
- Stegeman JJ, Elskus AA, Pruell R (1992) Endogenously-mediated, pretranslational suppression of cytochrome P4501A in PCB-contaminated flounder. Marine Environmental Research 34: 97-101.
- McMaster ME, Munkittrick KR, Luxon PL, Van Der Kraak GJ (1994) Impact of low-level sampling stress on interpretation of physiological responses of white sucker exposed to effluent from a bleached kraft pulp mill. Ecotoxicol Environ Saf 27: 251-264.
- Williams JP, Courtena CS, Wilson EC (1998) Annual sex steroid profiles and effects of gender and season on Cytochrome P450 mRNA induction in Atlantic tomcod (Microgadus tomcod). Environmental Toxicology and Chemistry 17: 1582-1588.
- Jonny BJ, Johnson G, Porte C, Krahn MM, Ariese F (2010) Analytical methods for determining metabolites of polycyclic aromatic hydrocarbon (PAH) pollutants in fish bile: A review. Environ Toxicol Pharmacol30: 224-244.
- Halm S, Rand-Weaver M, Sumpter JP, Tyler CR (2001) Cloning and molecular characterization of an ovarian-derived (brain-like) P450 aromatase cDNA and development of a competitive RT-PCR assay to quantify its expression in the fathead minnow (Pimephalespromelas). Fish Physiology and Biochemistry 24: 49-62.
- Kishida M, Callard GV (2001) Distinct cytochrome P450 aromatase isoforms in zebrafish (Daniorerio) brain and ovary are differentially programmed and estrogen regulated during early development. Endocrinology 142: 740-750.
- Greytak SR, Champlin D, Callard GV (2005) Isolation and characterization of two cytochrome P450 aromatase forms in killifish (Fundulusheteroclitus): differential expression in fish from polluted and unpolluted. Aquatic Toxicology 71: 371-389.
- Bemanian V, Rune Male, Anders Goksøyr (2004) The aryl hydrocarbon receptor-mediated disruption of vitellogenin synthesis in the fish liver: Cross-talk between AHR- and ERα-signalling pathways. Comp Hepatol.
- Vogel CF, Matsumura F (2009) A new cross-talk between the Aryl hydrocarbon receptor and RelB, a member of the NF-κB family.Biochem Pharmacol 77: 734-745.
- Yuan Z (2003) Resistance of cytochrome P4501A1 mRNA to induction by halogenated aryl hydrocarbons in Hudson River Atlantic tomcod-its prevalence and mechanistic basis " Doctoral dissertation, Department of Environmental Medicine, New York University School of Medicine, Tuxedo, New York.
- O'Donnell EF, Kopparapu PR, Koch DC, Jang HS, Phillips JL, et al. (2012) The aryl hydrocarbon receptor mediates leflunomide-induced growth inhibition of melanoma cells. PLoS One 7: e40926.
- Rager JE, Fry RC (2012) The aryl hydrocarbon receptor pathway: a key component of the microRNA-mediated AML signalisome. Int J Environ Res Public Health 9: 1939-1953.
- Oda Y, Nakajima M, Mohri T, Takamiya M, Aoki Y, et al. (2012) Aryl hydrocarbon receptor nuclear translocator in human liver is regulated by miR-24." Toxicol Appl Pharmacol 260: 222-231.
- Klerks PL, Leberg PL, Lance RF, McMillin DJ, Means JC (1997) Lack of development of pollutant-resistance or genetic differentiation of darter gobies (Gobionellusboleosoma) inhabiting a produced-water discharge site. Marine Environmental Research 44: 377-395.
- Weis JS, Mugue N, Weis P (1999) Mercury tolerance, population effects, and population genetics in the mummichog, Fundulusheteroclitus, in: V.E " Forbes (Ed.), Genetics and Ecotoxicology, Taylor & Francis, Philadelphia, 1999: 31-54.
- Wirgin I, Waldman JR (2004) Resistance to contaminants in North American fish populations.Mutat Res 552: 73-100.
- Lefèvre T, Williams AJ, de Roode JC (2011) Genetic variation in resistance, but not tolerance, to a protozoan parasite in the monarch butterfly. Proc Biol Sci 7: 751-759.
- Shivanna B, Zhang W, Jiang W, Welty SE, Couroucli XI, et al. (2013) Functional deficiency of aryl hydrocarbon receptor augments oxygen toxicity-induced alveolar simplification in newborn mice. Toxicol Appl Pharmacol 267:209-217.
- Yuan Z, Courtenay S, Wirgin I (2006) Comparison of hepatic and extra hepatic induction of cytochrome P4501A by graded doses of aryl hydrocarbon receptor agonists in Atlantic tomcod from two populations. Aquat Toxicol 76: 306-320.
Relevant Topics
- Algal Blooms
- Blue Carbon Sequestration
- Brackish Water
- Catfish
- Coral Bleaching
- Coral Reefs
- Deep Sea Fish
- Deep Sea Mining
- Ichthyoplankton
- Mangrove Ecosystem
- Marine Engineering
- Marine Fisheries
- Marine Mammal Research
- Marine Microbiome Analysis
- Marine Pollution
- Marine Reptiles
- Marine Science
- Ocean Currents
- Photoendosymbiosis
- Reef Biology
- Sea Food
- Sea Grass
- Sea Transportation
- Seaweed
Recommended Journals
Article Tools
Article Usage
- Total views: 16284
- [From(publication date):
December-2014 - Aug 17, 2025] - Breakdown by view type
- HTML page views : 11534
- PDF downloads : 4750