Research Article Open Access
Bacillus subtilis Spore Surface Display System Protects Recombinant Proteins from Degradation-Verified Hypothesis
Tomasz Łęga1*, Paulina Weiher2, Monika Paszkiewicz3 and Dawid Nidzworski2,41Department of Medical Biotechnology, Intercollegiate Faculty of Biotechnology, University of Gdańsk and Medical University of Gdańsk, Dębinki 1, 80-211 Gdańsk, Poland
2Department of Recombinant Vaccine, Intercollegiate Faculty of Biotechnology, University of Gdańsk and Medical University of Gdańsk, Kładki 24, 80-822 Gdańsk, Poland
3Department of Environmental Analytics, Faculty of Chemistry, University of Gdansk, ul. Wita Stwosza 63, 80-308 Gdansk, Poland
4Institute of Biotechnology and Molecular Medicine, Trzy Lipy 3 St., 80-172 Gdańsk
- *Corresponding Author:
- Łęga T
Department of Medical Biotechnology
Intercollegiate Faculty of Biotechnology
University of Gdańsk and Medical University of Gdańsk
Dębinki 1, 80-211 Gdańsk, Poland
Tel: +48725108632
E-mail: tomasz.lega@biotech.ug.edu.pl
Received date: May 19, 2017; Accepted date: June 24, 2017; Published date: June 30, 2017
Citation: Łęga T, Weiher P, Paszkiewicz M, Nidzworski D (2017) Bacillus subtilis Spore Surface Display System Protects Recombinant Proteins from Degradation- Verified Hypothesis. J Biotechnol Biomater 7: 263. doi: 10.4172/2155-952X.1000263
Copyright: © 2017 Łęga T, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Visit for more related articles at Journal of Biotechnology & Biomaterials
Abstract
Endospores of Bacillus subtilis have been used extensively as a platform for recombinant protein display for nearly two decades. Main use of the spore surface display system is generation of oral vaccines against many human and animal pathogens. Formulation of oral vaccine based on spores is an attractive approach to alternative vaccination due to the timesaving and relative easiness of production. Another advantage of such formulation is stability of presented antigens. It is assumed that the spore coat structure prevents degradation of displayed proteins by many destructive factors such as heat, proteases or stomach environment. However, there is little scientific background and substantial lack of experiments verifying this statement. In our work, we tested protective properties of spores against degradation of displayed antigens in harsh environment. We constructed B. subtilis strains producing spores presenting highly conserved long α-helix (LAH) region of the influenza A virus hemagglutinin. The constructs were obtained by fusion of LAH antigen to protein CotB or CotZ from the spore coat. We treated recombinant spores with destructive agents such as heat, protease, low pH and high-energy irradiation to test protective features of the spore coat. After treatment, spore coat extracts were analyzed by western blot to study fate of the displayed antigen. Results that we publish indicate that spore coat protects displayed antigen from degradation. This work is a strong support of hypothesis stating protective properties of spore surface display system against antigen degradation.
Keywords
Vaccination; Bacillus subtilis; Fusion gene; Recombinant spores; Centrifugation
Introduction
Bacillus subtilis is a model organism of Gram-positive bacteria. It can be commonly found in many environments however it is considered mainly as a soil bacterium [1]. One of the interesting features of this bacterium is production of the endospores in response to unfavorable condition for vegetation. Endospores are multi-structural, metabolically dormant entities that serve as a “safety shuttle” for bacterial genetic material which expression can be restored in process called germination [2,3]. Endospores are resistant to numerous destructive agents and harsh environments what is maintained by their specific structure [4]. Extreme endurance of the spores drew attention of the scientist to use it as a platform for displaying heterologous proteins or peptides. In the 2001 Isticato et al. presented concept of presenting 459-aminoacid C-terminal fragment of the tetanus toxin (TTFC) on the surface of B. subtilis endospores [5]. Presentation of the TTFC on spore surface was accomplished by fusing tetC gene to the B. subtilis gene encoding CotB spore coat protein so called anchoring protein. This original, genetic approach in presented system becomes a standard method. The initial purpose of such recombinant spores was to construct a platform for generation of cost-effective, timesaving and easy to administer oral vaccines. This trend resembles until today as most of the studies using spore surface display are focused on generation of new vaccine candidates [6].
The most common arguments raised by the researchers to use spores as a platform for generation of oral vaccines is that spore-based vaccine is heat-stable and will prevent degradation of presented antigens in harsh environment in stomach. However, to our best knowledge, there are no published results showing what the fate of the presented antigen is after treating recombinant spores with heat or exposing them to the stomach environment. In our work, we fused gene encoding long α-helix (LAH) region of the influenza A virus hemagglutinin to the genes coding for B.subtilis CotZ and CotB spore coat protein. Obtained spores presenting LAH antigen were treated with heat, electron beam radiation (e-beam), lyophilisation and were exposed to simulated gastric fluid. As shown by the western blot analyses none of the agents, except e-beam radiation, did not influence antigen load in the spore coat.
Materials and Methods
Bacterial strains and transformation
All cloning experiments were done using Escherichia coli DH5α [7]. Bacterial strains were transformed using previously described procedures: CaCl2-mediated transformation of E. coli competent cells [8] and transformation of B. subtilis [9]. Bacterial strains used in this study are listed in Table 1.
Construction of fusion genes
To generate genetic fusion the LAH region (76–130aa TKIDLWSYNAELLVALENQHTIDLTDSEMNKLFEKTRRQLRENA) of the A/HK/1/68 HA2 was synthesized (Life Technologies) with codon optimization for Bacillus subtilis and restriction site BamHI was added at the 5’ and SacI at the 3’ end of the ORF. Synthesized fragment was digested with BamHI and SacI and cloned at the 3’ end of the cotB and cotZ gene carried by plasmids pCotB-C [11] and pAGW2 [11], respectively obtaining plasmids pTL32 and pTL35.
Chromosomal integration
Plasmids pTL32 and pTL35 were linearized by digestion with a BsmBI restriction enzyme and used to transform B. subtilis 168. Transformation resulted in the integration of the plasmids into bacterial chromosome at the amyE locus in a double homologous recombination manner. Obtained chloramphenicol-resistant (CmR) clones were PCRtested for the incorporation of the fusion gene at the amyE locus in B. subtilis 168 chromosome using oligonucleotide pair AmyS/AmyA (S1). Selected clones were called BTL32 and BTL35 (Table 1).
| Strain | Relevant genotype | References |
|---|---|---|
| E. coli DH5a | fhuA2 lac(del)U169 phoAglnV44 F8 lacZ(del)M15 gyrA96 recA1 relA1 endA1 thi-1 hsdR17 | Woodcock [7] |
| B. subtilis 168 | trpC2 | Anagnostopoulos and Crawford [10] |
| B. subtilis BTL32> | amyE::cotB-LAH | This work |
| B. subtilis BTL35 | amyE::cotZ-LAH | This work |
Table 1: Strain list.
Preparation of spores
Sporulation was induced using the previously described nutrient depletion method in Difco sporulation medium (DSM) [12]. Sporulating cultures were harvested 48 h after the initiation of sporulation and purified using a lysozyme treatment, followed by washing in 1 M NaCl, 1 M KCl and water (twice each), as described by Nicholson et al. [12]. The spore suspension was titrated immediately for determination of c.f.u./mL before freezing at -20°C. We could reliably produce 7 × 1010 spores per liter of DSM culture using this method.
Extraction of spore coat proteins
To extract spore coat proteins a buffer containing 0.1 M NaCl, 0.1 M NaOH, 0.1 M Dithiothreitol (DTT) and 1% (w/v) SDS was used. 5 × 108 spores were resuspended in 50 μL of decoating buffer and incubated for 30 min at 70°C with shaking (1000 rpm). Suspension was centrifuged (10000 RCF, 10 min, RT) and supernatant kept for further analysis. Extracted proteins were assessed for concentration using Quick Start™ Bradford 1x Dye Reagent (Bio-Rad, USA).
Western blotting analyses
15 μg of total coat protein extracted were separated in NuPAGE® Novex® 4-12% Bis-Tris pre-cast polyacrylamide gels (Life Technologies), electrotransferred on a nitrocellulose using iBlot® 2 Dry Blotting System (Life Technologies). Membranes were incubated overnight at 4°C with anti-Influenza A H3 mAb (SinoBiological Inc. Catalog No. 11056- MM03). Western blots were visualized developing with BCIP/NBT according to the manufacturer’s instructions (Thermo Scientific).
Exposing spores to harsh environment
For temperature treatment 5 × 108 spores resuspended in 1 mL of dH2O were incubated for 60 min in 95°C followed by centrifugation and extraction of coat proteins. Spores were lyophilized using the Modulyo (Edwards) lyophilizer and incubated for six months at 37°C. After incubation spores were reconstituated in 1 mL of dH2O followed by centrifugation and extraction of coat proteins. 5 × 108 spores were resuspended in 1 mL of dH2O and were irradiated with high-energy electron beam (10 MeV, 15 kW) with the 25 kGy irradiation dose. Spores were centrifuged and coat proteins were extracted. 5 × 108 spores were incubated in 2 mL tube with 1mL artificial gastric fluid described elsewhere [13] for three hours at 37°C with 1000 rpm shaking using Thermo-Shaker (Novazym). An independent set of tubes were prepared for time 0, 15, 30, 45, 60, 120 and 180 min. After incubation spores were washed three times with 1 mL of d H2O followed by extraction of coat proteins.
Results and Discussion
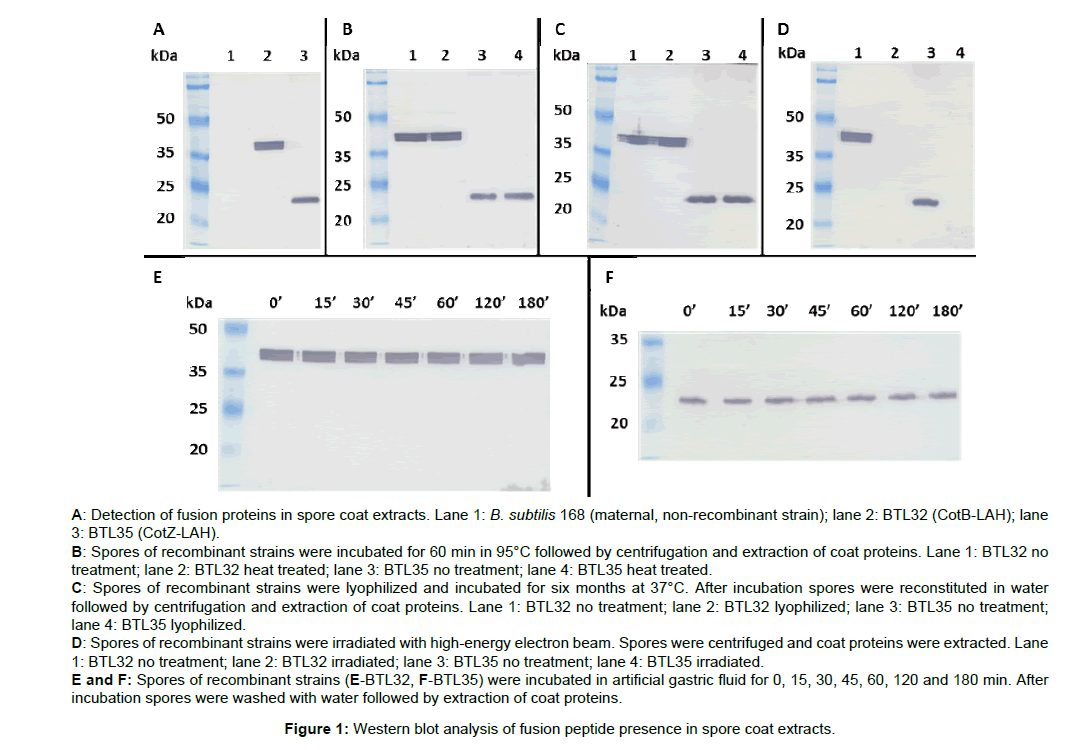
The presence of fusion proteins in the recombinant endospores coats was tested by western blotting with mouse monoclonal antiinfluenza A H3 antibodies. As demonstrated by western blotting CotZLAH and CotB-LAH fusion proteins were expressed and located in the spore coat (Figure 1A). Observed molecular weights of the fusion proteins CotZ-LAH and CotB were consistent with the calculated being 23 kDa and 38 kDa, respectively.
Proteinaceous coat is a multilayered, outermost structure of the spore and it provides resistance to enzymes, chemicals, and harsh environment. Thus, it is believed that heterologous proteins presented on spores are protected from degradation by destructive environment. In our work, we tested four agents to which recombinant spores can be exposed during their preparation or after application as an oral vaccine. Recombinant spores of both tested strains BTL32 and BTL35 remained their initial antigen load in the coat after treatment with 95°C for 60 min (Figure 1B). Recombinant spores were freeze-dried and stored for six months at 37°C and after reconstitution antigen load in the coat was checked. As proved by western blotting in case of both strains tested antigen load in the spore coat after treatment remained at their initial levels (Figure 1C). Sterilization of spores using electron irradiation resulted in total degradation of presented antigen in both recombinant strains (Figure 1D). To test protective properties of the spore coat in case of the stomach environment we performed in vitro test. Spores were incubated in artificial gastric fluid for three hours after which antigen content in the coat was checked with western blotting. After treatment spores produced by both recombinant strains showed antigen load on their initial levels (Figures 1E and 1F).
Authors of many publication presenting B. subtilis spores as a platform for generation of vaccine candidates state that such constructs are heat stable and can prevent degradation of antigen in the stomach environment [4,14-16]. However, none of the authors present experiments showing the direct effect of heat or peptidase treatment of recombinant spores and influence on the antigen content. Thus, protective properties of spore surface display in terms of antigen degradation in harsh environment remains a hypothesis. In our work, we tested this hypothesis and showed that in fact spores presenting heterologous peptide can protect it from degradation by harsh environment. We chose two anchor proteins: CotB form the outer coat and CotZ from the outermost layer called crust to check whether antigen localization in the spore coat influence protective properties of the system. In our study, we did not notice any significant differences in term of antigen protection between using CotB and CotZ as anchor proteins. Both fusion proteins CotB-LAH and CotZ-LAH remained at the same level after treatment with heat and artificial gastric fluid.
Conclusion
These results verify hypothesis that spore-based vaccine candidates are heat-stable and can be orally delivered without degradation of antigen in stomach environment. However, electron beam sterilization of spores resulted in total degradation of antigen. Such method of material preparation would not be appropriate for spore based vaccine. Nevertheless, our work clearly shows that Bacillus subtilis spore surface display is a robust and durable platform for generation of orally administered vaccine candidates.
Acknowledgement
The research was supported by the Polish National Center for Research and Development grant no LIDER/016/489/L-4/12/NCBR/2013.
References
- Earl AM, Losick R, Kolter R (2008) Ecology and genomics of Bacillus subtilis. Trends Microbiol 16: 269-275.
- Higgins D, Dworkin J (2012) Recent progress in Bacillus subtilis sporulation. FEMS Microbiol Rev 36: 131-148.
- Setlow P (2014) Germination of spores of Bacillus species: What we know and do not know. J Bacteriol 196: 1297-1305.
- Duc le H, Cutting SM (2003) Bacterial spores as heat stable vaccine vehicles. Expert Opin Biol Ther 3: 1263-1270.
- Isticato R, Cangiano G, Tran HT, Ciabattini A, Medaglini D, et al. (2001) Surface display of recombinant proteins on Bacillus subtilis spores. J Bacteriol 183: 6294âВ?В?6301.
- Wang H, Wang Y, Yang R (2017) Recent progress in Bacillus subtilis spore-surface display: Concept, progress, and future. Appl Microbiol Biotechnol 101: 933-949.
- Woodcock DM (1989) Quantitative evaluation of Escherichia coli host strains for tolerance to cytosine methylation in plasmid and phage recombinants. Nucleic Acids Res 17: 3469-3478.
- Green MR, Sambrook J (2012) Molecular cloning. A laboratory manual. Cold Spring Harbor Laboratory Press: Cold Spring Harbor.
- Julkowska D, Obuchowski M, Holland IB, Séror SJ (2005) Comparative analysis of the development of swarming communities of Bacillus subtilis 168 and a natural wild type: Critical effects of surfactin and the composition of the medium. J Bacteriol 187: 65-76.
- Anagnostopoulos C, Crawford IP (1961) Transformation studies on the linkage of markers in the tryptophan pathway in Bacillus subtilis. Proc Nat Acad Sci USA 47: 378-390.
- Iwanicki A, Piatek I, Stasia M, Grela A, Laga T, et al. (2014) A system of vectors for Bacillus subtilis spore surface display. Microb Cell Fact 13: 30.
- Nicholson WL, Setlow P (1990) Sporulation, germination and outgrowth. Molecular biological methods for Bacillus. John Wiley & Sons Ltd, England.
- Corcoran BM, Stanton C, Fitzgerald GF, Ross RP (2005) Survival of probiotic lactobacilli in acidic environments is enhanced in the presence of metabolizable sugars. Appl Environ Microbiol 71: 3060-3067.
- Uyen NQ, Hong HA, Cutting SM (2007) Enhanced immunisation and expression strategies using bacterial spores as heat-stable vaccine delivery vehicles. Vaccine 25: 356-365.
- Amuguni H, Tzipori S (2012) Bacillus subtilis: A temperature resistant and needle free delivery system of immunogens. Hum Vaccin Immunother 8: 979-986.
- Zhao G, Miao Y, Guo Y, Qiu H, Sun S, et al. (2014) Development of a heat-stable and orally delivered recombinant M2e-expressing B. subtilis spore-based influenza vaccine. Hum Vaccin Immunother 10: 3649-3658.
Relevant Topics
- Agricultural biotechnology
- Animal biotechnology
- Applied Biotechnology
- Biocatalysis
- Biofabrication
- Biomaterial implants
- Biomaterial-Based Drug Delivery Systems
- Bioprinting of Tissue Constructs
- Biotechnology applications
- Cardiovascular biomaterials
- CRISPR-Cas9 in Biotechnology
- Nano biotechnology
- Smart Biomaterials
- White/industrial biotechnology
Recommended Journals
Article Tools
Article Usage
- Total views: 3622
- [From(publication date):
July-2017 - Aug 16, 2025] - Breakdown by view type
- HTML page views : 2700
- PDF downloads : 922