Research Article Open Access
Development of a Separation and Detection System for Bacillus anthracis Spores Based on Peptide Conjugates Identified from Peptide Library
Gi Wook Kim1, Jong Pil Park2, Sang Yup Lee3 and Tae Jung Park1*
1Department of Chemistry, Chung-Ang University, 84 Heukseok-ro, Dongjak-gu, Seoul 156-756, Republic of Korea
2Department of Pharmaceutical Engineering, Daegu Haany University, 290 Yugok-dong, Gyeongsan, Gyeongbuk 712-715, Republic of Korea
3Department of Chemical and Biomolecular Engineering, Department of Bio and Brain Engineering, Department of Biological Sciences, BioProcess Engineering Research Center, Bioinformatics Research Center, Center for Systems and Synthetic Biotechnology, and Institute for the BioCentury, KAIST, 291 Gwahak-ro, Yuseonggu, Daejeon 305-701, Republic of Korea
- Corresponding Author:
- Tae Jung Park
Department of Chemistry, Chung-Ang University
84 Heukseok-ro, Dongjak-gu, Seoul 156-756, Republic of Korea
Tel: +82-2-820-5220
Fax: +82-2-825-4736
E-mail: tjpark@cau.ac.kr
Received Date: November 18, 2014; Accepted Date: December 29, 2014; Published Date: January 05, 2015
Citation: Kim GW, Park JP, Lee SY, Park TJ (2015) Development of a Separation and Detection System for Bacillus anthracis Spores Based on Peptide Conjugates Identified from Peptide Library. Biosens J 4:112. doi:10.4172/2090-4967.1000112
Copyright: © 2015 Kim GW, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Biosensors Journal
Abstract
A simple, specific and rapid identification system against Bacillus anthracis spores was developed using specific capture peptides conjugated with a bead-based biosensor, which was tightly and specifically bound onto the spore surface of B. anthracis. We successfully detected and separated B. anthracis spores from the spores of B. thuringiensis and B. cereus using this peptide-magnetic bead conjugates by fluorescence and anthrax-specific analyses. For more convenient mediation of high-throughput detection against B. anthracis spores, streptavidin-biotin interactions of spore-peptide and spore-peptide-magnetic bead conjugates were performed, and we demonstrated the separation and detection of B. anthracis spores using this method. In the presence of mixed condition of several types of Bacillus spores, the B. anthracis spores were easily separated using the oligopeptide-conjugated magnetic beads, thus allowing the clear detection of B. anthracis spores from B. cereus, B. subtilis, and B. thuringiensis spores. This oligopeptidebased strategy was rapidly and unambiguously identified as little as one viable B. anthracis spore in less than 1 h with a simple binding assay format. When assessed for its effectiveness for specifically and selectively detection of environmental spores phylogenetically similar to B. anthracis spores, such as B. thuringiensis and B. cereus spores, the system was free of false-positive signals.
Keywords
Bacillus anthracis; Anthrax detection; Magnetic separation; Capture peptide; Oligopeptide-conjugated ligand
Abbreviations
PCR: Polymerase Chain Reaction; PS-SPCLs: Positional Scanning-Synthetic Peptide Combinatorial Libraries; FITC: Fluorescein Isothiocyanate; HPLC: High-Performance Liquid Chromatography; SULFO-NHS: N-HydroxySulfosuccinimide
Introduction
With recent threats of biological terrorism and outbreaks of microbial pathogens, a great need for biosensors that can quickly and accurately identify infectious agents has arisen. Bacillus anthracis is a gram-positive, spore-forming bacterium that causes anthrax, a lethal disease for both humans and animals [1-5]. Because of its potential use for bioterrorism, B. anthracis has become a critical threats to many countries [6,7]. Once exposed to internal tissues, B. anthracis spores germinate and enter a vegetative state, often resulting in the death of the host within several days. Furthermore, great similarity between B. anthracis and other members of the Bacillus genus such as B. cereus, B. thuringiensis, and B. mycoides, exists. Therefore, the rapid and specific detection of environmental B. anthracis spores prior to infection is of extreme importance for the preservation of human safety and national security.
Various biological and chemical techniques have been developed. Nanoparticle-based colorimetric or fluorometric assay are most common detection methods. Recently, direct and specific detection studies with the help of hydrogel-based materials and physicsbased methods were developed to detect virus and bacteria [8,9]. Furthermore, label-free methods implemented for the specific detection of DNA strands were developed [10,11]. Among the most important biological methods for the detection of anthrax, Polymerase Chain Reaction (PCR) [5-7] and immunoassay [12-14] have been used as representatives. PCR, a primer-mediated enzymatic DNA amplification method, requires expensive reagents and takes 3-4 h or longer several hours to obtain the result. Moreover, PCR needs considerable effort in sample preparation processing prior to analysis. The detection limit of PCR, which is based on the detection of bacterial pagA and lef genes encoding the protective antigen toxic protein, has been ~103 spores. Immunoassays, which rely on the interaction between B. anthracis spore surface antigens and their specific antibodies, can detect 105 spores in approximately 12 h. However, in the case of immunoassays, specific antibodies and additional molecular fluorophores and/or enzymatic reactions must be employed for the desired agents to be detected, and mobile-phase conditions must also be adjusted depending on their capture, elution, and separation properties. In addition, although this direct spore detection system is relatively fast, the lack of accuracy and limited sensitivity of current antibody-based detection methods result in unacceptably high levels of both false-positive and false-negative responses [12-14]. Therefore, the development of a more robust detection system is required.
Many biological actions such as ligand-receptor interactions are based on the specificity of proteins conferred by the primary sequence of amino acids as well as the secondary and tertiary structures dictated by the primary sequence. Of particular importance is the formation of local structures, such as the active site and motif, which play a key role in protein function. Among the random sequences of short peptides, there may be sequences that can ideally act on such local environments, and also serve as the focal point for the development of new and more effective drugs. Recently various methods have been developed for rapid and easily identification of the sequences of interest from vast mixtures of random peptide sequences or polymers (template) with various side chain groups (chemical diversity libraries) [15-19]. Successful screening of these libraries has dbeen described not only for epitopes recognized by their antibodies [20-22], but also for the identification of the biologically active peptides, such as antibacterial and antifungal peptides [16], human immunodeficiency virus protease inhibitors [23], substrate-analog trypsin inhibitors [24], and interleukin-8-specific antagonists [25].
In this study, we developed a high-throughput detection system for B. anthracis spores with oligopeptides using fluorescence, magnetic beads, and positional scanning-synthetic peptide combinatorial libraries (PS-SPCLs). Specific capture oligopeptides were developed to specifically bind against the surface of B. anthracis spores. Several peptide ligands were conjugated with Fluorescein Isothiocyanate (FITC) to detect fluorescent signals, and magnetic beads were used to facilitate rapid isolation by magnetic polarity conjugated with streptavidin to bind with biotin-conjugated oligopeptides. The resulting system was capable of the detection of specific spores with high sensitivity in less than 1 h. We also optimized the system to maximize its binding affinity to spores of two B. anthracis strains, namely, B. anthracis ΔSterne (pXO1-, pXO2-) and B. anthracis Sterne 34F2 (pXO1+, pXO2-), by using fluorescence assay and magnetic separation. Based on genome sequence comparisons, two species of Bacillus, B. cereus and B. thuringiensis, which are most similar to B. anthracis, were used in this study as model strains for comparison before the actual diagnosis of B. anthracis. Finally, to further improve the performance of the capture peptides, we newly designed their amino acid sequences using PS-SPCLs that can bind B. anthracis spores with greater specificity. The detection system based on these specific capture peptides is a high-throughput fluorescence-based assay that employs quantum dots. These specific binding systems can be employed to rapidly, specifically and selectively detect and separate B. anthracis spores in mixed conditions.
Materials and Methods
Materials
Streptavidin-conjugated magnetic beads (~ 1-2 μm in diameter) and a portable handheld magnetic separator were purchased from PureBiotech (San Diego, CA). The capture peptides used in this study (Table 1) were synthesized by Peptron (Korea). For rapid separation of B. anthracis spores from the spore mixture, the magnetic microbeads was functionalized with the capture peptides and used for detection of spores.
| Capture peptide | Sequence (N-terminus to C-terminus) | Reference |
|---|---|---|
| BA-1 | biotin-ATYPLPIRGGGC | 5 |
| B-Negative | biotin-SLLPGLPGGGC | 5 |
| Negative-amine | SLLPGLPGGGC-NH2 | 5 |
| BaS-01 | EWWWNW-NH2 | This study |
| BaΔS-01 | DWDHDW-NH2 | This study |
| BaΔS-02 | DWDSEW-NH2 | This study |
| BaΔS-03 | DWDKDW-NH2 | This study |
Table 1: Capture peptides used in this study.
Bacterial strains and spores
The Bacillus strains used in this study are listed in Table 2. Experiments with B. anthracis ΔSterne (pXO1-, pXO2-) and B. anthracis Sterne 34F2 (pXO1+, pXO2-) were carried out at Professor Y.G. Chai’s laboratory at Hanyang University to avoid transportation and handling problems.
| Strain or plasmid | Relevant characteristics | Reference or source |
|---|---|---|
| Strains | ||
| B. cereus 1092 | KCTC 1092 | KCTCa |
| B. subtilis DB104 | His nprR2 nprE18 ΔaprA3 (BGSC 1E75) | BGSCb |
| B. thuringiensis 4Q7 | Plasmidless mutant of B. thuringiensis var. israelensis | BGSCb |
| B. anthracis��?Sterne | (pXO1-, pXO2-) | Prof. Chaic |
| B. anthracisSterne 34F2 | (pXO1+, pXO2-) | Prof. Chaic |
aKorean Collection for Type Cultures, Daejeon, Korea
bBacillus Genetic Stock Center, Columbus, OH
cProf. Y.G. Chai’s laboratory, Hanyang University, Ansan, Korea
Table 2: Bacterial strains and plasmids used in this study.
Synthesis and conjugation of individual capture peptides
The capture peptides were chemically synthesized and purified by High-Performance Liquid Chromatography (HPLC) according to the manufacturer’s protocol (Peptron, Daejeon, Korea). Biotin was attached for conjugation at the N-terminus of capture peptides. The coupling efficiency was determined by analyzing the peptide solution, before and after coupling, by prominence HPLC (Shimadzu, Japan) and then reading the absorbance at a wavelength of 220 nm. The peptides were characterized by chromatography on a C18 column (Shiseido Capcell Pak, 22×300 mm). The peptides were eluted with a 0-30% gradient of water and acetonitrile in 0.1% (v/v) trifluoroacetic acid. The peptide composition was confirmed by amino acid analysis [26], and the peptides were sequenced using an Applied Biosystems® 473A protein/peptide microsequencer (Life Technologies, Grand Island, NY).
Preparation of positional scanning-synthetic peptide combinatorial libraries
Libraries were synthesized according to the protocol of Houghten et al. [16] and Pinilla et al. [19]. Briefly, PS-SPCLs, consisting of six-residue peptide sequences having free N termini and amidated C-termini, were synthesized. A single position in each peptide mixture was individually and specifically defined with 19 of the 20 natural L-amino acids (cysteine excluded), while the five remaining positions consisted of mixtures of the same 19 amino acids. Defined positions with known amino acids are represented by O and the positions with mixed amino acids are represented by X. The six sets of PS-SPCLs (a total of 114 pools) are represented by the following formulae: O1XXXXX-NH2, XO2XXXX-NH2, XXO3XXX-NH2, XXXO4XX-NH2, XXXXO5X-NH2, and XXXXXO6-NH2. Libraries of peptides were constructed on Rapidamide resin beads as described elsewhere called fluorenylmethyloxycarbonyl (Fmoc) chemistry [27]. The resin beads were distributed into different reaction vessels for each amino acid at each coupling step, then pooled, washed, and thoroughly mixed for randomization. Beads were then deprotected and redistributed into the various vessels again for the next coupling step and so on. The amount of each amino acid used to yield approximately equimolar coupling was determined empirically. The completeness of each reaction was checked with ninhydrin [26]. Side chains were deprotected with a mixture of trifluoroacetic acid (ethanedithiol:water:thioanisole; 90:5:4:1, vol/vol). The 114 peptide mixtures were individually extracted with water, lyophilized, and dissolved in water at a final concentration of 27 nM for a peptide sequence in each pool.
Cultivation of cells and purification of spores
Cells were cultivated in CDSM media at 37°C or 30°C and 250 rpm for 48-60 h [28]. Spores mixed with vegetative cells were harvested from 50 mL of the culture by centrifugation (5,000 rpm) and then resuspended in 0.2 mL of 20% (w/v) urografin (Sigma, St. Louis, MO). This suspension was gently layered over 1 mL of 50% (w/v) urografin in a 1.5 mL microcentrifuge tube, and then centrifuged for 10 min at 4°C and 13,000 rpm. The collected pellet contained only free spores.
Polymerase chain reaction
PCR experiments were performed with a PCR Thermal Cycler (Bio-rad, Hercules, CA) using the High Fidelity PCR System. DNA fragments encoding lethal factor 2 were obtained by PCR using the genomic DNA of B. cereus, B. anthracis Sterne and ΔSterne as templates, and the universal primers as follows: Forward primer: 5’-AAGCTTTGAGCAAGTTCATTCAAAAGC-3’; Reverse primer: 5’-ATTGGAAAGTTTTCGGAGCA-3’.
Fluorescence analysis
Fluorescence assays were performed using a spectrofluorometer (Model VICTOR2, PerkinElmer, Shelton, CT). Green fluorescence samples were excited by a 488/543 nm HeNe laser and were filtered by a long pass 505/575 nm filter.
Results and Discussion
Peptide labeling with biotin
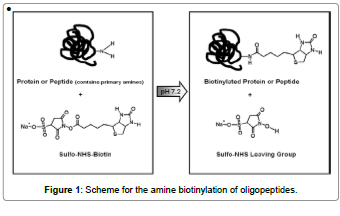
Each of the capture peptides created from the PS-SPCL peptide library have free N-termini and amidated C-termini. These peptide termini can be reacted with N-hydroxysulfosuccinimide (sulfo- NHS)-biotin reagents to enable simple and efficient biotin labeling of peptides, proteins and any other primary amine-containing molecules (Figure 1). Differing only in their spacer arm lengths, this reagent offers the possibility of optimizing labeling and detection experiments where steric hindrance of biotin binding is an important factor. Sulfo-NHS esters of biotin are the most popular type of biotinylation reagent. NHSactivated biotins react efficiently with primary amine groups (-NH2) in pH 7-9 buffers to form stable amide bonds. Several different NHS esters of biotin are available, with varying properties and spacer arm lengths. The sulfo-NHS ester reagents are water soluble, enabling reactions to take place in the absence of organic solvents such as dimethyl sulfoxide or dimethylformamide.
Proteins, including antibodies, generally have several primary amines in the side chain of lysine (K) residues and the N-termini of each polypeptide that are available as targets for labeling with NHS-activated biotin reagents. However, because these molecules dissolve readily in polar solutions and are charged by the sulfonate groups, they cannot penetrate the cell and spore. As long as the spore remains intact, only primary amines of peptides bound on the anthrax spore surface will be biotinylated with sulfo-NHS-biotin reagents. Biotinylated-peptide or protein can be detected by streptavidin-conjugated fluorescent dye.
We have a library of 400 peptides (Peptide Library Support Facility of Pohang University of Science and Technology, Korea). These pools were individually dissolved in PBS solution at a final concentration of 27 nM per peptide having free N-termini and amidated C-termini. The peptide pools first react with anthrax spores in each 96-well, and are then washed three times to remove unbound peptides. These peptidebound spores react with sulfo-NHS-biotin reagents, and streptavidin- FITC, and from these samples, we can detect specific bound peptides against anthrax spores. Next, the peptide sequences can be designed.
Screening and identification of specific binding peptides
We screened peptides bound to the B. anthracis Sterne spores, which were generated from PS-SPCLs, to examine their corresponding peptide-spore reactions. A total of 114 peptide pools of PS-SPCLs were subjected to testing, and the most effective amino acid sequence at each of the six positions in a hexapeptide was determined. The results of the initial screening of the peptide library are shown in Figure 2. The peptide mixture, EXXXNX-NH2, was found to strongly bind to the B. anthracis Sterne spores. The amino acids with slightly less activity than glutamate (E) at the first position were aspartate (D) and valine (V). The active amino acids at the second, third and fourth positions were all tryptophan (W), with slightly more activity than others. The most active amino acid at the fifth position was asparagine (N). At the sixth position, tryptophan (W) was significantly more active than other amino acids. The amino acids chosen for reiterative synthesis of the peptide were as follows: 1st, E; 2nd, W; 3rd, W; 4th, W; 5th, N; and sixth, W. The final selected amino acid sequence was EWWWNWNH2, named BaS-01.
Figure 2: Screening of the PS-SPCLs to select peptides that bind to B. anthracis Sterne spores. Panels represent the results obtained with the peptide pools having a known amino acid at each of the six positions in the hexapeptide. One of the six positions (O1, O2 . . . and O6) is assigned with each of the 19 L-amino acids. The remaining five positions consist of mixtures (X) of 19 L-amino acids (cysteine excluded). The library consists of 114 peptide pools. The optimized binding peptide sequence was the EWWWNW hexamer, named BaS-01.
Likewise, we applied the same binding and screening procedure to the B. anthracis ΔSterne spores, and the test elucidated the most active amino acid from 114 peptide pools at each amino acid position in the hexapeptide. The results for the initial screening of the peptide library are presented in Figure 3. The peptide mixture, DXXXXX-NH2, was found to strongly bind to B. anthracis ΔSterne spores. The most active amino acid at the first position was aspartate (D). The amino acids with slightly less activity than tryptophan (W) at the second position were alanine (A) and aspartate (D). The most active amino acid at the third position was aspartate (D). Histidine (H), lysine (K), and serine (S) appeared to be more active than other amino acids at the fourth position. The active amino acids at the fifth position were aspartate (D) and glutamate (E), with aspartate being slightly more active than the others. Tryptophan (W) was more active than other amino acids at the sixth position. The amino acids chosen for reiterative synthesis of the peptide were as follows: 1st, D; 2nd, A, D, and W; 3rd, D; 4th, E, H, K, and S; 5th, D and E; and sixth, W. The selected amino acid sequences used were DWDHDW-NH2, DWDSEW-NH2, and DWDKDW-NH2 hexamers, named BaΔS-01, BaΔS-02, and BaΔS-03, respectively.
Figure 3: Screening of the PS-SPCLs to select peptides that bind to the B. anthracis ΔSterne spores. Panels represent the result obtained with the peptide pools having a known amino acids at each of the six positions in hexapeptide. One of the six positions (O1, O2 . . . and O6) is assigned with each of the 19 L-amino acids. The remaining five positions consist of mixtures (X) of 19 L-amino acids (cysteine excluded). The library consists of 114 peptide pools. The optimized binding peptide sequences are DWDHDW, DWDSEW, and DWDKDW hexamers, named BaΔS-01, BaΔS-02, and BaΔS-03, respectively.
Each active peptide pool was synthesized at Peptron and confirmed by using HPLC on a C18 column. We subsequently investigated if the newly designed peptides had specific binding effects on B. anthracis spores with a binding assay incubating the active peptide (EWWWNW-NH2) with B. anthracis Sterne spores for 15 min at room temperature (Figure 4). The binding affinity of the other three active peptides (DWDHDW-NH2, DWDSEW-NH2, and DWDKDW-NH2) was examined with B. anthracis ΔSterne spores by 15-min incubation at room temperature (Figure 5). From these results, we developed the specific binding peptides with about 70-100% higher binding affinity against B. anthracis spores via peptide library screening, when compared with the BA1 peptide reported by Williams et al. [5].
Figure 4: Binding affinity of the synthesized hexapeptide from the selected amino acids to B. anthracis Sterne spores. Negative, SLLPGLPGGGC as a negative control; BA1, ATYPLPIRGGGC; BaS-01, EWWWNW.
Specificity and selectivity for B. anthracis spores of specific binding peptides
In order to verify their specificity corresponding peptide-spore reactions, the capture peptides bound onto the B. anthracis Sterne and ΔSterne spores (1.0×108 CFU/mL) were checked with same amount of spores of other Bacillus strains, B. cereus, B. subtilis, and B. thuringiensis, respectively (Figure 6a). The newly designed peptides have been specific binding effects on B. anthracis spores after incubating the active capture peptide (EWWWNW-NH2) with B. anthracis Sterne spores and other three Bacillus spores for 15 min at room temperature, respectively (Figure 6a). The binding affinities of the other three active peptides (DWDHDW-NH2, DWDSEW-NH2, and DWDKDW-NH2) were examined with B. anthracis ΔSterne spores and other three Bacillus spores by 15-min incubation at room temperature (Figure 6b- 6d). From these results, we confirmed the specific and selective binding peptides with higher binding affinity against B. anthracis spores, when compared with other several bacilli spores. Finally, the detection limit of the method was calculated one more time by 3-sigma rule. As a result, the limit was 1.8×102 CFU/mL for B. anthracis Sterne spores, upto 1.7×102 CFU/mL for B. anthracis ΔSterne spores, respectively. The screened capture peptides could be specifically and sensitively bound onto the B. anthracis spore surface.
Figure 6: Specificity for B. anthracis spores of the capture peptides with binding affinity of (a) EWWWNW hexamer, named BaS-01, (b) DWDHDW, named BaΔS-01, (c) DWDSEW, named BaΔS-02, and (d) DWDKDW, named BaΔS-03. The purified spores (1.0 × 108 CFU/mL) of B. cereus, B. subtilis, B. thuringiensis, and B. anthracis were examined, respectively
Rapid separation of B. anthracis spores using oligopeptideconjugated magnetic microbeads
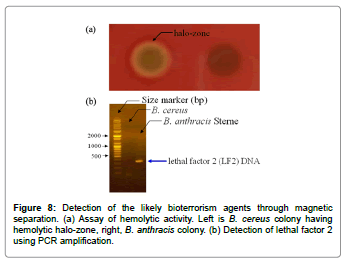
We next examined the possibility of detecting B. anthracis spores when mixed with other microorganisms, as is the case in natural environments. As shown in the schematic diagram of Figure 7, rapid separation of B. anthracis spores from the spore mixture was performed using magnetic microbeads. The BaS-01 peptides were incubated with the streptavidin-conjugated magnetic microbeads for 15 min at room temperature and washed with PBS buffer to remove unbound peptides. The peptide-bound magnetic microbeads were then incubated with a 1:1 mixture of B. cereus and B. anthracis Sterne spores. The sample mixture became slightly turbid with the formation of brown precipitate after incubation. The precipitate could be moved to the side wall of the tube by using a handheld magnet separator. To confirm the separation of the two types of spores (B. cereus and B. anthracis Sterne), the supernatant (unbound spores) and precipitate (bound spores) fractions were separately spread on a Mueller Hinton agar plate for the detection of hemolytic activity. As shown in Figure 8a, a halo-zone caused by hemolytic activity was observed with the unbound spores, but not with the bound spores. Since B. cereus, not B. anthracis, is supposed to show hemolytic activity, these results indicate that this method allows for simple and rapid isolation of B. anthracis spores from B. cereus spores. To further confirm the successful separation of the two types of spores, PCR was carried out with universal primers and genomic DNA of each fraction (B. cereus and B. anthracis after germination) as templates. We found that the PCR product encoding lethal factor 2 (ca. 400 bp) was only detected in B. anthracis Sterne (Figure 8a and 8b).
Figure 7: Schematic diagram for the rapid separation of B. anthracis spores using magnetic beads in the presence of other spores and microorganisms. The magnetic bead-peptide complex coupled by streptavidin-biotin interaction was specifically bound to B. anthracis spores. These magnetic bead-peptidespore complexes were separated by magnetic force.
Even with the success of this method, the major disadvantages, including false-positive results and non-specific bindings, must be prevented for urgent use during times of a biowarfare threat. Therefore, additional assays involving PCR and/or other antibodies that target different spore surface antigens should be used to separate and detect all forms of B. anthracis spores in environmental mixed samples. From a practical standpoint, it will be necessary to develop a sample preparation method for the detection of dilute B. anthracis spores present in water, powder, or other environmental conditions. When a suitable sample preparation method is developed in aqueous condition, it will then be possible to employ this system for the early detection of a number of real-life situations encountering anthrax, a dangerous bacterium (e.g., for personal checks against anti-terrorismtargeted activities).
Conclusion
The capture peptide-nanobead-based method developed in this study allows rapid, simple and accurate detection of B. anthracis spores. The high affinity to B. anthracis Sterne and ΔSterne spores was obtained by generating high-affinity-peptides against B. anthracis Sterne and ΔSterne spores via screening assays from the peptide library. Although our fluorescence-based methods require relatively expensive materials and sophisticated laboratory equipment for analysis, our assays described herein take less than 1 h to detect B. anthracis spores in the presence of other spores and microorganisms. We also developed a simple and rapid separation method for B. anthracis spores using magnetic microbeads; this method can specifically and sensitively detect B. anthracis spores with a detection limit of 1.7×102 CFU/mL when they are mixed with other microorganisms such as those characteristic of the natural environment.
Acknowledgement
We thank Professor Y.G. Chai (Hanyang University, Korea) for allowing us to use the B. anthracis Sterne and ΔSterne strains in his laboratory. The synthesis and analysis of peptides were conducted at the Peptide Library Support Facility of the Korea Science and Engineering Foundation at POSTECH. This work was supported by the IT R&D program of MOTIE/MISP/KEIT (10044580) and Advanced Production Technology Development Program, Ministry of Agriculture, Food and Rural Affairs (312066-3). Further supported by the Korean Health Technology R&D Project, Ministry of Health and Welfare (HI13C0862). The work of JPP was supported by the National Research Foundation of Korea(NRF) grant funded by the Korea government(MSIP) (No. NRF-2014R1A2A2A01005621). The work of SYL was supported by the Technology Development Program to Solve Climate Changes on Systems Metabolic Engineering for Biorefineries from the Ministry of Science, ICT and Future Planning (MSIP) through the National Research Foundation (NRF) of Korea (NRF-2012-C1AAA001-2012M1A2A2026556).
References
- Higgins JA, Cooper M, Schroeder-Tucker L, Black S, Miller D, et al. (2003) A field investigation of Bacillus anthracis contamination of U.S. department of agriculture and other Washington, D.C., buildings during the anthrax attack of October 2001. Appl Environ Microbiol 69:593-599.
- Oncu S,SelcenOncu, Sakarya S (2003) Anthrax--an overview. Med SciMonit 9:276-283.
- Das A, Das ND, Park JH, Lee HT, Choi MR, et al. (2013) Identification of survival factors in LPS-stimulated anthrax lethal toxin tolerant RAW 264.7 cells through proteomic approach. BioChip J 7:75-84.
- Wang SH, Wen JK, Zhou YF, Zhang ZP, Yang RF, et al. (2004) Identification and characterization of Bacillus anthracis by multiplex PCR on DNA chip. BiosensBioelectron 20:807-813.
- Williams DD, Benedek O, Turnbough CL Jr (2003) Species-specific peptide ligands for the detection of Bacillus anthracis spores. Appl Environ Microbiol 69:6288-6293.
- Hartley HA, Baeumner AJ (2003) Biosensor for the specific detection of a single viable B. anthracis spore. Anal BioanalChem 376:319-327.
- Reissman DB, Steinberg EB, Magri JM, Jernigan DB (2003) The anthrax epidemiologic tool kit: an instrument for public health preparedness. Biosecur Bioterror 1:111-116.
- Shin JO, Cherstvy AG, Metzler R (2014) Sensing viruses by mechanical tension of DNA in responsive hydrogels. Phys Rev X 4:021002.
- Ghosh SK, Cherstvy AG, Metzler R (2014) Deformation propagation in responsive polymer network films. J ChemPhys 141:074903.
- Abouzar MH, Poghossian A, Cherstvy AG, Pedraza AM, Ingebrandt S, et al. (2012) Label-free electrical detection of DNA by means of field-effect nanoplate capacitors: Experiments and modeling. Phys Status Solid A 209:925-934.
- Cherstvy AG (2013) Detection of DNA hybridization by field-effect DNA-based biosensors: mechanisms of signal generation and open questions. BiosensBioelectron 46:162-170.
- Bell CA, Uhl JR, Hadfield TL, David JC, Meyer RF, et al. (2002) Detection of Bacillus anthracis DNA by lightcycler PCR. J ClinMicrobiol 40:2897-2902.
- Oggioni MR, Meacci F, Carattoli A, Ciervo A, Orru G, et al. (2002) Protocol for real-time PCR identification of anthrax spores from nasal swabs after broth enrichment. J ClinMicrobiol 40:3956-3963.
- Welkos SL, Cote CK, Rea KM, Gibbs PH (2004) A microtiterfluorometric assay to detect the germination of Bacillus anthracis spores and the germination inhibitory effects of antibodies. J Microbiol Methods 56:253-265.
- Ostresh JM, Husar GM, Blondelle SE, Dorner B, Weber PA, et al. (1994) Libraries from libraries: chemical transformation of combinatorial libraries to extend the range and repertoire of chemical diversity. ProcNatlAcadSci USA 91:11138-11142.
- Houghten RA, Pinilla C, Blondelle SE, Appel JR, Dooley CT, et al. (1991) Generation and use of synthetic peptide combinatorial libraries for basic research and drug discovery. Nature 354:84-86.
- Lam KS, Salmon SE, Hersh EM, Hruby VJ, Kazmierski WM, et al. (1991) A new type of synthetic peptide library for identifying ligand-binding activity. Nature 354:82-84.
- Zuckermann RN, Kerr JM, Siani MA, Banville SC, Santi DV (1992) Identification of highest-affinity ligands by affinity selection from equimolar peptide mixtures generated by robotic synthesis. ProcNatlAcadSci USA 89:4505-4509.
- Pinilla C, Appel JR, Blanc P, Houghten RA (1992) Rapid identification of high affinity peptide ligands using positional scanning synthetic peptide combinatorial libraries. BioTechniques 13:901-905.
- Felici F, Castagnoli L, Musacchio A, Jappelli R, Cesareni G (1991) Selection of antibody ligands from a large library of oligopeptides expressed on a multivalent exposition vector. J MolBiol 222:301-310.
- Stephen CW, Lane DP (1992) Mutant conformation of p53. Precise epitope mapping using a filamentous phage epitope library. J MolBiol 225:577-583.
- Luzzago A, Felici F, Tramontano A, Pessi A, Cortese R (1993) Mimicking of discontinuous epitopes by phage-displayed peptides, I. Epitope mapping of human H ferritin using a phage library of constrained peptides. Gene 128:51-57.
- Owens RA, Gesellchen PD, Houchins BJ, DiMarchi RD (1991) The rapid identification of HIV protease inhibitors through the synthesis and screening of defined peptide mixtures. BiochemBiophys Res Commun 181:402-408
- Eichler J, Houghten RA (1993) Identification of substrate-analog trypsin inhibitors through the screening of synthetic peptide combinatorial libraries. Biochemistry 32:11035-11041.
- Hayashi S, Kurdowska A, Miller EJ, Albright ME, Girten BE, et al. (1995) Synthetic hexa- and heptapeptides that inhibit IL-8 from binding to and activating human blood neutrophils. J Immunol 154:814-824.
- Kaiser E, Colescott RL, Bossinger CD, Cook PI (1970) Color test for detection of free terminal amino groups in the solid-phase synthesis of peptides. Anal Biochem 34:595-598.
- Houghten RA (1985) General method for the rapid solid-phase synthesis of large numbers of peptides: specificity of antigen-antibody interaction at the level of individual amino acids. ProcNatlAcadSci USA 82:5131- 5135.
- Nicholson WL, Setlow P (1990) Molecular biological methods for Bacillus: John Wiley & Sons.
Relevant Topics
- Amperometric Biosensors
- Biomedical Sensor
- Bioreceptors
- Biosensors Application
- Biosensors Companies and Market Analysis
- Biotransducer
- Chemical Sensors
- Colorimetric Biosensors
- DNA Biosensors
- Electrochemical Biosensors
- Glucose Biosensors
- Graphene Biosensors
- Imaging Sensors
- Microbial Biosensors
- Nucleic Acid Interactions
- Optical Biosensor
- Piezo Electric Sensor
- Potentiometric Biosensors
- Surface Attachment of the Biological Elements
- Surface Plasmon Resonance
- Transducers
Recommended Journals
Article Tools
Article Usage
- Total views: 16659
- [From(publication date):
June-2015 - Sep 02, 2025] - Breakdown by view type
- HTML page views : 11827
- PDF downloads : 4832