Research Article Open Access
Heme Activates Macrophage Hepcidin Expression via Toll like Receptor 4 and Extracellular Signal-Regulated Kinases Signaling Pathway
Naveen Kumar Tangudu and Maja Vujić Spasić*Institute of Comparative Molecular Endocrinology, Ulm University, Ulm, Germany
- *Corresponding Author:
- Prof. Dr. Maja Vujić Spasić
Institute of Comparative Molecular Endocrinology
Ulm University, 89081 Ulm, Germany
Tel: +49 731 50 32635
Fax: +49 731 50 32609
E-mail: maja.vujic@uni-ulm.de
Received date: December 21, 2016; Accepted date: January 16, 2017; Published date: January 21, 2017
Citation: Tangudu NK, Vujic Spasic M (2017) Heme Activates Macrophage Hepcidin Expression via Toll like Receptor 4 and Extracellular Signal-Regulated Kinases Signaling Pathway. Clin Pharmacol Biopharm 6:166. doi: 10.4172/2167-065X.1000166
Copyright: © 2017 Tangudu NK, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Clinical Pharmacology & Biopharmaceutics
Abstract
Tight regulation of systemic and cellular iron levels is required for good health. This control is ensured by hepcidin, a small peptide hormone produced predominantly by the liver. Lack of hepcidin expression or mutations affecting regulators of hepcidin expression, cause common genetic iron disorders. Hepcidin is also expressed in myeloid cells and its expression is increased after infections and in response to lipopolysaccharide. Our study uncovers that macrophages rapidly increase hepcidin expression in response to excess of heme. Moreover, we demonstrate that the underlying mechanism by which heme triggers hepcidin activation in macrophages depends on the Toll Like Receptor (TLR)-4 and the contribution of Extracellular Signal-Regulated Kinases (ERK) pathway. Our data propose the contribution of hepcidin, locally produced by macrophages, to the pathology of disorders characterized by excess of free heme, such as certain bacterial infections and hemolytic disorders. Finally, using macrophages from Hfe-deficient mice, we demonstrate that the lack of Hfe is not critical for the hepcidin induction by heme but is required to maintain basal hepcidin expression in macrophages. The findings that the levels of hepcidin in macrophages are directly controlled by the actions of Hfe in these cells expand our view on Hfe beyond the liver and as mere regulator of iron levels.
Keywords
Heme; Macrophages; Hfe; Hepcidin expression
Introduction
Hepcidin is a small peptide hormone produced by the liver, which ensures for a tight balance of systemic and cellular iron levels in the body. Although its function appeared to be related to drosophila’s antimicrobial peptides like defensins, hepcidin is nowadays established as the key systemic regulator of iron homeostasis [1-5]. Hepcidin orchestrates systemic iron fluxes by blocking iron absorption form the duodenum and iron release from macrophages [6]. It does so by binding to ferroportin, the sole iron-exporter predominantly expressed on macrophages and enterocytes, causing its internalization, degradation and subsequent iron retention within the macrophages [6]. The principal function of hepcidin in the regulation of systemic iron levels has been revealed using genetically engineered mouse models with overexpression or loss-off hepcidin function [7-10]. Lack of hepcidin expression or mutations affecting regulators of hepcidin expression (e.g. Hfe) cause hemochromatosis, a common genetic iron overload disorder [11].
Although hepcidin is produced mainly by the hepatocytes, many other cells/tissues are able to synthesize the hormone [12-17]. However, the exact physiological role of the extra-hepatocytic hepcidin is still unknown. For example, it was shown that increased hepcidin expression by macrophages contributed to down regulation of ferroportin protein levels in an autocrine manner [18] and that hepcidin expression in myeloid cells was increased after infections and in response to lipopolysaccharide, LPS [12]. LPS is a potent proinflammatory molecule, which affects the innate immune responses through Toll Like Receptor-4 (Tlr-4) mediated signaling. Recently, heme (an iron-porphyrin complex) was shown to act as an extracellular damage-associate protein with strict binding specificity for Tlr4 on macrophages, which differs from the binding sites occupied by LPS [19]. Binding of heme to Tlr-4 results in the activation of proinflammatory responses [19-21]. In addition to activating the proinflammatory Tlr4-dependent responses, heme exerts potentially harmful pro-oxidant and cytotoxic effects [22,23]. These actions of heme are reflected in its properties to increase the production of reactive oxygen species, which are generated through heme catabolism within the cells [21,24,25], and in the ability of heme to impair cell membrane integrity causing cell burst [26]. High levels of heme occur as a consequence of hemolysis (extensive cell damage) and are present in common oxidative stress conditions, certain pathological conditions such as hemorrhages, bacterial infections, malaria, hemoglobinopathies, trauma, hemolytic anemias, and in hemodialysis and blood transfusion.
Given the role of macrophages as the major inflammatory cells and a key agent in iron homeostasis, we hypothesized that heme, a potent pro-inflammatory iron-containing molecule, exerts its actions on hepcidin expression in macrophages. Moreover, in this study we address whether hepcidin expression in the macrophages is influenced by the lack of well-known hepcidin regulator, the hemochromatosis protein Hfe, which deficiency in mice results in inappropriately low hepcidin expression and systemic iron overload [27,28].
Materials and Methods
Animal experimentations
Wild type, Tlr4-/- and Hfe-/- mice, age between 8-12 weeks, were used in the experiments. All mice were maintained on a standard mouse diet containing 200 mg/kg iron (Ssniff, Soest, Germany), under a constant dark-light cycle, and were allowed access to food and water ad libitum . Mice were sacrificed by CO2 inhalation. All animal experiments were approved by and conducted in compliance with the guidelines of the Ulm University Animal Care Committee and the Federal Authorities for Animal Research (Regierungspraesidium Tuebingen, Baden-Wuerttemberg, Germany).
Preparation of bone marrow-derived macrophages (BMDMs)
BMDMs were obtained from wild type, Tlr4-/- and Hfe-/- mice as described previously [27]. In brief, bone marrows were cultured in Dulbecco’s Minimal Eagles Medium (Invitrogen, USA) supplemented with 10% Fetal Bovine Serum, 10 mM Sodium-Pyruvate, 10 mM LGlutamine, penicillin and streptomycin (Sigma Aldrich, St. Louis, USA) and L929 cell-conditioned medium as a source for macrophagecolony stimulating factor. Following 4 days culture, non-adherent cells were removed and adherent cells were washed with PBS and the medium was replaced daily up to day 7, when the cells were used for the experiments.
Treatments of bone marrow-derived macrophages (BMDMs) in vitro
Experiments were performed using Haemin (ferric-protoporphyrin IX that is reduced to ferrous-protoporphyrin IX heme within cells), LPS (L2630, E. coli, serotype 0111:B4), iron-free heme analog protoporphyrin IX (PPIX), all purchased from Sigma Aldrich (St. Louis, USA) and U0126 (MEK1/2 inhibitor; Cell Signaling, USA).
RNA isolation, reverse transcription and quantitative real-time PCR
RNA isolation, reverse transcription, quantitative real-time PCR (qPCR) and data analysis were performed as described previously [29]. In brief, total RNA was isolated using RNeasy Midi kit (Qiagen, Hilden, Germany) according to the manufacturer’s instruction. RevertAid H Minus (M-MulV) reverse transcriptase (Fermentas, USA), 5x RT reaction buffer, random primers (200 ng/μl, Invitrogen, USA) and 10 mM dNTPs were used to convert 1-2 μg of RNA to cDNA following the manufacturer’s instructions. Quantitative realtime PCR was carried out in 10 μl reaction volume using SYBR Green I dye (Invitrogen, USA) on ABI ViiA-7 system (Applied Biosystems, USA). The mRNA abundance of the investigated genes was calculated relative to the expression of the reference gene Gapdh and data analyzed using a model based on correction for exact PCR efficiencies. Primers used in the study are listed in Table 1.
| Gene Name | Gene Symbol | Forward Sequence (5’-3’) | Reverse Sequence (5’-3’) |
|---|---|---|---|
| Hepcidin | Hamp | ATACCAATGCAGAAGAGAAGG | AACAGATACCACACTGGGAA |
| Hemeoxygenase | Hmox1 | AGGCTAAGACCGCCTTCCT | TGTGTTCCTCTGTCAGCATCA |
| Glyceraldehyde-3-Phosphate Dehydrogenase | Gapdh | CCCATTCTCGGCCTTGACTGT | GTGGAGATTGTTGCCATCAACGA |
Table 1: Primers used for qPCR.
Protein analysis
Protein extracts were prepared from BMDMs after homogenization in RIPA lysis buffer (50 mM Tris-Cl pH 8.0/150 mM NaCl/1% NP-40/0.5% DOC/0.1% SDS) supplemented with the protease inhibitors (Complete Mini (25) ROC 11836153001, Roche Diagnostics, Mannheim, Germany) and phosphatase inhibitors (1 mM Na3O4Va/25 mM NaF/1 mM PMSF, Sigma Aldrich, Germany) for 30 min on ice, as previously described [29]. Cell debris was removed by centrifugation at 13,000 rpm for 15 min at 4°C, and protein concentration determined using the Pierce BCA660 nm Protein Assay Kit (Thermo Fisher Scientific, Rockford USA). Equal amounts of protein extracts (30 μg) were diluted in 5x Laemmli buffer (0.34 Tris-HCl pH 6.8/0.7% SDS/0.6 M DTT/34% glycerol/bromphenol-blue) and subjected to a 10% SDSpolyacrylamide gel electrophoreses, transferred to nitrocellulose membrane, blocked for 1 h in 3% milk and 1% BSA (Sigma Aldrich, Germany) blocking solution. Blots were incubated with rabbit antipErk1/ 2 antibody (1:1000, in blocking solution; Cell Signaling Technology, USA) overnight at 4 degrees. Membranes were further incubated with anti-rabbit HRP antibody (1:5000, Invitrogen, USA) for 1 h in blocking solution, membrane developed in presence of DAB substrate (Thermo scientific, USA) and visualized in chemiluminescence detector (BioRad, USA). Membranes were stripped and re-incubated with mouse antibody against β-actin (1:10,000; Sigma Aldrich, USA) used for normalization of protein loading. The signals were quantified by computer-assisted image analysis (ImageJ: https://imagej.nih.gov/ij/).
Statistical analyses
Data were analyzed using GraphPad Prism software and results are shown as mean ± SEM (standard error of mean). Statistical analysis was performed using the Student t-test (two-tailed; unequal variance). A probability value p<0.05 was measured as statistically significant.
Results and Discussion
The goal of this study was to investigate the properties of macrophages to mount hepcidin induction in response to ironporphyrin complex, heme, and to assess the contribution of the Hfe in this process.
Heme stimulates hepcidin induction in macrophages
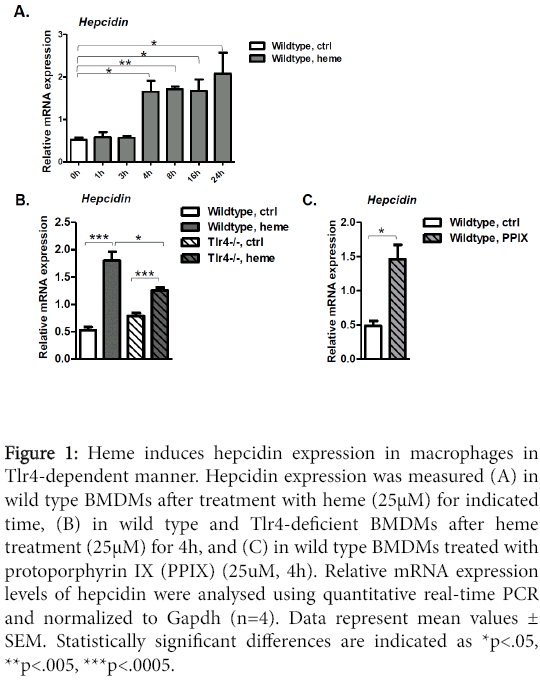
To evaluate whether hepcidin expression is modulated by heme in macrophages, we prepared bone marrow-derived macrophages (BMDMs) from wild-type mice and exposed them to in vitro treatment with heme (25 μM) for the indicated time (Figure 1A). Hepcidin mRNA levels were measured using real-time qPCR analysis. We showed that hepcidin expression rapidly increased in response to heme (by more than 3-fold, p<0.04) and was maintained elevated even 24 h after the heme stimulation (Figure 1A).
Figure 1: Heme induces hepcidin expression in macrophages in Tlr4-dependent manner. Hepcidin expression was measured (A) in wild type BMDMs after treatment with heme (25μM) for indicated time, (B) in wild type and Tlr4-deficient BMDMs after heme treatment (25μM) for 4h, and (C) in wild type BMDMs treated with protoporphyrin IX (PPIX) (25uM, 4h). Relative mRNA expression levels of hepcidin were analysed using quantitative real-time PCR and normalized to Gapdh (n=4). Data represent mean values ± SEM. Statistically significant differences are indicated as *p<.05, **p<.005, ***p<.0005.
Hepcidin induction in macrophages by Heme depends on Tlr4
To elucidate the contribution of Tlr4 to hepcidin activation by heme, we isolated BMDMs from wild type and Tlr4-deficient mice. Cells were stimulated in vitro with heme (25 μM; 4 h) and the expression levels of hepcidin were compared in regard to control cells. Our results showed that the basal mRNA expression levels of hepcidin were not affected in BMDMs from Tlr4-deficient mice when compared to the levels in wild type macrophages (Figure 1B). However, the observed hepcidin induction by heme in wild type BMDMs (by 3.4- fold, p<0.0007 in heme-treated versus control) was abolished in Tlr4- deficient macrophages. Namely, upon exposure to heme we detected 1.6-fold, p<0.0005 increase in hepcidin expression in heme-treated Tlr4-deficient BMDMs versus controls. In line with this, the levels of hepcidin induction by heme in Tlr4-deficient macrophages were by 50% reduced when compared to hepcidin levels in heme treated wild type macrophages (1.43-fold, p<0.02), suggesting that activation of hepcidin by heme was dependent on the functional Tlr4 signaling (Figure 1B). In addition, stimulation of BMDMs with an iron-free heme analog PPIX, which interacts with Tlr4 [19], increased hepcidin levels similarly to the levels in heme-treated cells (Figure 1C). Our findings demonstrate that heme employs the common proinflammatory Tlr4 receptor to signal for hepcidin activation in macrophages. However, since mild but significant induction of hepcidin was measured in heme-treated versus untreated Tlr4- deficient macrophages (1.6-fold, p<0.0005) (Figure 1B), we cannot fully exclude the possibility that additional signalling pathway(s) may enhance the overall hepcidin induction by heme.
Induction of hepcidin by heme is MAPK/ERK dependent
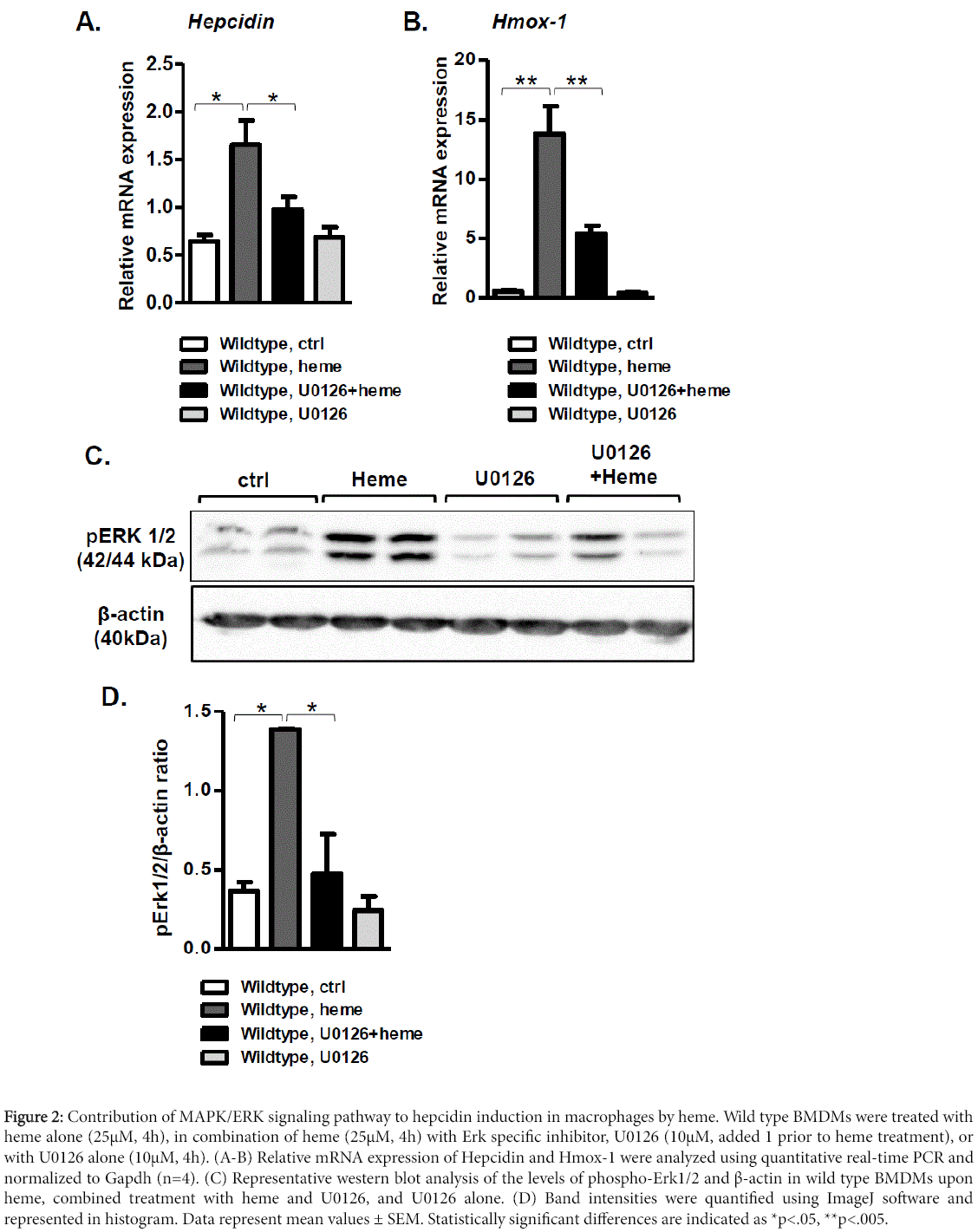
Based on the above findings, we postulated that MAPK/ERK (mitogen-activated protein kinases/extracellular signal-regulated kinases) signaling pathway might be the missing link, which contributed to hepcidin induction by heme. This hypothesis was based on the previous studies showing that: i) MAPK/ERK pathway was activated by heme [19,30]; and ii) activation of the MAPK/ERK signaling contributed to hepcidin induction by holotransferrin in primary hepatocytes [31]. We therefore pretreated wild type macrophages with specific MAPK/ERK inhibitor U0126 (10 μM) 1 h before stimulation with heme (25 μM; for 4 h). We show that hemetriggered hepcidin induction was almost fully suppressed upon combined treatment of macrophages with heme and MAPK/ERK inhibitor in regard to heme-treated cells (Figure 2A).
Figure 2: Contribution of MAPK/ERK signaling pathway to hepcidin induction in macrophages by heme. Wild type BMDMs were treated with heme alone (25μM, 4h), in combination of heme (25μM, 4h) with Erk specific inhibitor, U0126 (10μM, added 1 prior to heme treatment), or with U0126 alone (10μM, 4h). (A-B) Relative mRNA expression of Hepcidin and Hmox-1 were analyzed using quantitative real-time PCR and normalized to Gapdh (n=4). (C) Representative western blot analysis of the levels of phospho-Erk1/2 and β-actin in wild type BMDMs upon heme, combined treatment with heme and U0126, and U0126 alone. (D) Band intensities were quantified using ImageJ software and represented in histogram. Data represent mean values ± SEM. Statistically significant differences are indicated as *p<.05, **p<.005.
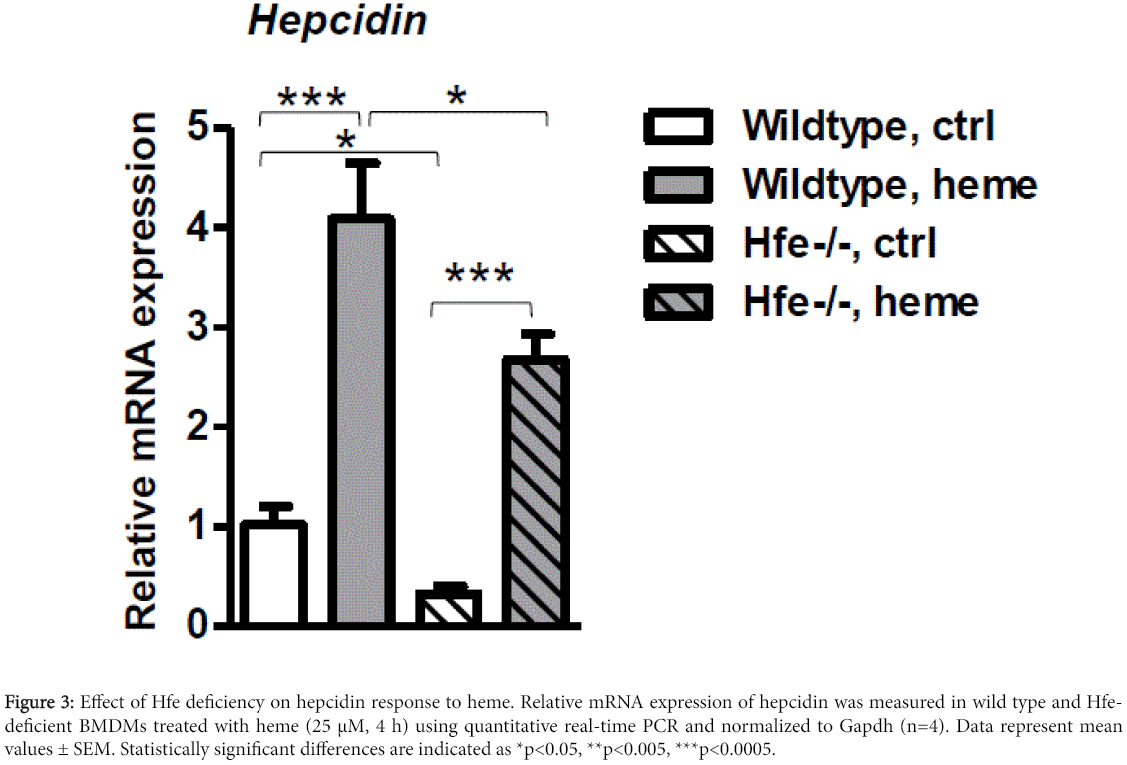
Figure 3: Effect of Hfe deficiency on hepcidin response to heme. Relative mRNA expression of hepcidin was measured in wild type and Hfedeficient BMDMs treated with heme (25 μM, 4 h) using quantitative real-time PCR and normalized to Gapdh (n=4). Data represent mean values ± SEM. Statistically significant differences are indicated as *p<0.05, **p<0.005, ***p<0.0005.
The presence of the MAPK/ERK inhibitor alone did not affect hepcidin expression (Figure 2A). The efficacy of MAPK/ERK inhibition was demonstrated by monitoring the expression of ERKtarget gene Hmox1 (which encodes for the heme oxygenase 1) (Figure 2B) and by using phospho-specific antibody to assess ERK activation by Western blot analysis (Figure 2C,D). These results demonstrate that MAPK/ERK pathway contributes to hepcidin induction by heme.
We suspect that activation of MAPK/ERK signaling by heme may not exclusively dependent on pro-inflammatory Tlr4-dependent activities of heme since mild but significant activation of hepcidin by heme was still observed in Tlr4-deficient cells (Figure 1B). One can postulate that pro-oxidant actions of heme, which are triggered upon heme catabolism within the cells and the production of reactive oxygen species, may add to overall activation of MAPK/ERK signaling and hepcidin activation by heme. Further studies are required to precisely delineate the contribution of pro-inflammatory, pro-oxidant and cytotoxic effects of heme to the regulation of hepcidin activation in macrophages.
Macrophage-Hfe is not required for the hepcidin induction by heme
We and others previously demonstrated that Hfe is a key component of iron-sensing regulatory complex in the liver, which controlled hepcidin production [27,28,32,33]. Correspondingly, mice with the lack of Hfe in the liver showed low hepatic hepcidin production and develop iron overload [27]. To determine to what extent Hfe is required for hepcidin regulation in macrophages, we isolated BMDMs from wild type and Hfe-/- mice. Cells were either left untreated or stimulated with heme (25 μM; 4 h) and hepcidin responses compared. We demonstrate that hepcidin expression by heme is induced to similar extent in BMDMs isolated from Hfe-/- mice and wild type mice (6.9-fold, p<0.0001 and 5.7-fold, p<0.0004, respectively), despite the basal hepcidin levels are significantly lower in Hfe-deficient macrophages in regard to wild type macrophages (1.9-fold, p<0.05).
These data imply that Hfe is required to maintain basal hepcidin expression in macrophages, whereas its contribution to heme-triggered hepcidin activation is dispensable. The findings that the levels of hepcidin in macrophages are directly controlled by the actions of Hfe in these cells correlates with the central role of Hfe in the regulation of hepcidin expression in hepatocytes. Thus the lack of Hfe, the main hepcidin regulator, affects hepcidin production both in hepatocytes and macrophages. These data expand our view on Hfe beyond the liver and hepatocytes as mere regulator of iron homeostasis and hepatic hepcidin expression. We propose that heme-Tlr4 signaling pathway dominates over Hfe for the regulation of hepcidin expression in macrophages and that the contribution of Hfe in macrophages is dispensable for hepcidin activation by heme.
Conclusion
The liver-derived peptide hormone hepcidin has been established as the key systemic regulator of iron homeostasis which expression greatly increased in response to iron accumulation and during bacterial infections and inflammation. By contrast, little is still known on the role of hepcidin produced by macrophages and other cells/ tissues. In this study we investigated the responses of macrophages upon exposure to heme and explored the signaling pathways governing hepcidin regulation.
We reveal that under high heme levels, macrophages increase hepcidin expression and this effect depends on binding of heme to Tlr4 receptor. The pro-inflammatory Tlr4-responses are equally activated upon LPS and previous study demonstrated that Tlr4-dependent activation was required for LPS-mediated hepcidin induction in macrophages [12]. These data imply that Tlr4-receptor dependent activities of heme mediate inflammatory responses and hepcidin activation in macrophages, resembling the effects of LPS [12]. Moreover, the findings that hepcidin expression in macrophages is positively regulated by heme suggest for the possible contribution of macrophage-hepcidin to the pathology of disorders characterized by excess of free heme such as common oxidative stress conditions, certain inflammatory and infectious diseases and chronic hemolytic disorders. We propose that in conditions with high heme levels, the Tlr4-dependent actions of heme affect macrophage immune responses and hepcidin expression proposing that blocking the Tlr4-receptor actions may be beneficial in resolving the inflammation and normalizing hepcidin levels. However, the exact physiological role of hepcidin produced by macrophages under physiological and pathophysiological conditions, such as those characterized by high levels of heme, will only be provided through the generation of mice with selective lack of hepcidin in macrophages and other extrahepatocytic cells.
Acknowledgments
We thank the staff of the Animal facility at the Ulm University for their contribution to this work. This work was supported by the German Research Foundation (DFG, VU75/2-1, to M.V.S.), and in part by the Ulm University (to M.V.S.).
Authorship Contributions
N.K.T., M.V.S. performed research and analyzed data; M.V.S. designed research, analyzed data, and wrote the paper.
Conflict of Interest
The author(s) declare no competing financial interests.
References
- Krause A, Neitz S, Magert HJ, Schulz A, Forssmann WG, et al. (2000) LEAP-1, a novel highly disulfide-bonded human peptide, exhibits antimicrobial activity. FEBS Lett 480:147-150.
- Pigeon C, Ilyin G, Courselaud B, Leroyer P, Turlin B, et al. (2001) A new mouse liver-specific gene,encoding a protein homologous to human antimicrobial peptide hepcidin, is overexpressed during iron overload.J BiolChem 276:7811-7819.
- Park CH, Valore EV, Waring AJ, Ganz T (2001) Hepcidin, a urinary antimicrobial peptide synthesized inthe liver. J BiolChem 276:7806-7810.
- Camaschella C (2005) Understanding iron homeostasis through genetic analysis of hemochromatosisand related disorders. Blood 106:3710-3717.
- Hentze MW, Muckenthaler MU, Galy B, Camaschella C (2010). Two to tango: regulation of Mammalianiron metabolism. Cell 142:24-38.
- Nemeth E, Tuttle MS, Powelson J, Vaughn MB, Donovan A, et al. (2004) Hepcidin regulates cellulariron efflux by binding to ferroportin and inducing its internalization. Science 306:2090-2093.
- Lesbordes-Brion JC, Viatte L, Bennoun M, Lou DQ, Ramey G, et al. (2006) Targeted disruption of thehepcidin 1 gene results in severe hemochromatosis. Blood 108:1402-1405.
- Viatte L, Nicolas G, Lou DQ, Bennoun M, Lesbordes-Brion JC, et al. (2006) Chronic hepcidin inductioncauses hyposideremia and alters the pattern of cellular iron accumulation in hemochromatotic mice. Blood107:2952-2958.
- Roy CN, Mak HH, Akpan I, Losyev G, Zurakowski D, et al. (2007) Hepcidin antimicrobial peptidetransgenic mice exhibit features of the anemia of inflammation. Blood 109:4038-4044.
- Zumerle S, Mathieu JR, Delga S, Heinis M, Viatte L, et al. (2014) Targeted disruption of hepcidin in theliver recapitulates the hemochromatotic phenotype. Blood 123:3646-3650.
- Vujic M (2014) Molecular basis of HFE-hemochromatosis. Front Pharmacol5:42.
- Peyssonnaux C, Zinkernagel AS, Datta V, Lauth X, Johnson RS, et al. (2006) TLR4-dependenthepcidin expression by myeloid cells in response to bacterial pathogens. Blood 107:3727-3732.
- Kulaksiz H, Fein E, Redecker P, Stremmel W, Adler G, et al. (2008) Pancreatic beta-cells expresshepcidin, an iron-uptake regulatory peptide. J Endocrinol 197:241-249.
- Kulaksiz H, Gehrke SG, Janetzko A, Rost D, Bruckner T, et al. (2004) Pro-hepcidin: expression and cellspecific localisation in the liver and its regulation in hereditary haemochromatosis, chronic renal insufficiency, andrenal anaemia. Gut 53:735-743.
- Bekri S, Gual P, Anty R, Luciani N, Dahman M, et al. (2006) Increased adipose tissue expression ofhepcidin in severe obesity is independent from diabetes and NASH. Gastroenterology 131:788-796.
- Zechel S, Huber-Wittmer K, von Bohlen und Halbach O (2006) Distribution of the iron-regulating proteinhepcidin in the murine central nervous system. J Neurosci Res 84:790-800.
- Pinto JP, Dias V, Zoller H, Porto G, Carmo H, et al. (2010) Hepcidin messenger RNA expression inhuman lymphocytes. Immunology 130:217-230.
- Theurl I, Theurl M, Seifert M, Mair S, Nairz M, et al. (2008) Autocrine formation of hepcidin induces ironretention in human monocytes. Blood 111:2392-2399.
- Figueiredo RT, Fernandez PL, Mourao-Sa DS, Porto BN, Dutra FF, et al. (2007) Characterization ofheme as activator of Toll-like receptor 4. J BiolChem 282:20221-20229.
- Vinchi F, Costa da Silva M, Ingoglia G, Petrillo S, Brinkman N, et al. (2016) Hemopexin therapy revertsheme-induced proinflammatory phenotypic switching of macrophages in a mouse model of sickle cell disease.Blood 127:473-486.
- Belcher JD, Nath KA, Vercellotti GM (2013)Vasculotoxic and Proinflammatory Effects of PlasmaHeme: Cell Signaling and Cytoprotective Responses. ISRN Oxidative Med 2013.
- Balla J, Vercellotti GM, Jeney V, Yachie A, Varga Z, et al. (2007) Heme, hemeoxygenase, and ferritin:how the vascular endothelium survives (and dies) in an iron-rich environment. Antioxid Redox Signal 9:2119-2137.
- Dutra FF, Bozza MT (2014) Heme on innate immunity and inflammation. Front Pharmacol5:115.
- Arruda MA, Barcellos-de-Souza P, Sampaio AL, Rossi AG, Graca-Souza AV, et al. (2006) NADPHoxidase-derived ROS: key modulators of heme-induced mitochondrial stability in human neutrophils. Exp CellRes 312:3939-3948.
- Moraes JA, Barcellos-de-Souza P, Rodrigues G, Nascimento-Silva V, Silva SV, et al. (2012) Hememodulates smooth muscle cell proliferation and migration via NADPH oxidase: a counter-regulatory role for hemeoxygenase system. Atherosclerosis 224:394-400.
- Gatidis S, Foller M, Lang F (2009) Hemin-induced suicidal erythrocyte death. Ann Hematol 88:721-726.
- VujicSpasic M, Kiss J, Herrmann T, Galy B, Martinache S, et al. (2008)Hfe acts in hepatocytes toprevent hemochromatosis. Cell Metab 7:173-178.
- Schmidt PJ, Toran PT, Giannetti AM, Bjorkman PJ, Andrews NC (2008) The transferrin receptormodulates Hfe-dependent regulation of hepcidin expression. Cell Metab 7:205-214.
- VujicSpasic M, Sparla R, Mleczko-Sanecka K, Migas MC, Breitkopf-Heinlein K, et al. (2013) Smad6and Smad7 are co-regulated with hepcidin in mouse models of iron overload. BiochimBiophysActa 1832:76-84.
- Wang JY, Lee YT, Chang PF, Chau LY (2009) Hemin promotes proliferation and differentiation ofendothelial progenitor cells via activation of AKT and ERK. J Cell Physiol 219:617-625.
- Ramey G, Deschemin JC, Vaulont S (2009) Cross-talk between the mitogen activated protein kinaseand bone morphogenetic protein/hemojuvelin pathways is required for the induction of hepcidin by holotransferrinin primary mouse hepatocytes. Haematologica 94:765-772.
- Schmidt PJ, Andrews NC, Fleming MD (2010)Hepcidin induction by transgenic overexpression of Hfedoes not require the Hfe cytoplasmic tail, but does require hemojuvelin. Blood 116:5679-5687.
- D'Alessio F, Hentze MW, Muckenthaler MU (2012) The hemochromatosis proteins HFE, TfR2, and HJVform a membrane-associated protein complex for hepcidin regulation. J Hepatol 57:1052-1060.
Relevant Topics
- Applied Biopharmaceutics
- Biomarker Discovery
- Biopharmaceuticals Manufacturing and Industry
- Biopharmaceuticals Process Validation
- Biopharmaceutics and Drug Disposition
- Clinical Drug Trials
- Clinical Pharmacists
- Clinical Pharmacology
- Clinical Research Studies
- Clinical Trials Databases
- DMPK (Drug Metabolism and Pharmacokinetics)
- Medical Trails/ Drug Medical Trails
- Methods in Clinical Pharmacology
- Pharmacoeconomics
- Pharmacogenomics
- Pharmacokinetic-Pharmacodynamic (PK-PD) Modeling
- Precision Medicine
- Preclinical safety evaluation of biopharmaceuticals
- Psychopharmacology
Recommended Journals
Article Tools
Article Usage
- Total views: 4126
- [From(publication date):
February-2017 - Aug 30, 2025] - Breakdown by view type
- HTML page views : 3182
- PDF downloads : 944