Research Article Open Access
Mesoglein Expression During Aurelia aurita Life Cycle
Anastasiya Kotova1, Anastasiya Naiden1, Alexander Shumeev1, Tatiana Shaposhnikova1, Olga Podgornaya1 and Leonid Adonin2*1Department of Cytology and Histology, St-Petersburg State University, St.-Petersburg, Russia
2Institute of Cytology RAS, St.-Petersburg, Russia; Far Eastern Federal University, School of Biomedicine, Vladivostok, Russia
- *Corresponding Author:
- Leonid Adonin
Institute of Cytology RAS, St.-Petersburg
Russia; Far Eastern Federal University
School of Biomedicine, Vladivostok
Russia
Tel: 007 911155-4469
E-mail: leo.adonin@gmail.com
Received date: November 01, 2016; Accepted date: November 15, 2016; Published date: November 25, 2016
Citation: Kotova A, Naiden A, Shumeev A, Shaposhnikova T, Podgornaya O, et al. (2016) Mesoglein Expression During Aurelia aurita Life Cycle. J Marine Sci Res Dev 6: 211. doi: 10.4172/2155-9910.1000211
Copyright: © 2016 Kotova A, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Marine Science: Research & Development
Abstract
Cnidarian is thought to possess two tissue layers: endoderm (gastroderm) and ectoderm, which are separated by the layer of extracellular matrix (ECM) called mesoglea. Aurelia aurita complex life cycle consists of several stages including alternating generations of sexual adult stage medusa and asexual stage polyp (scyphistoma). The main difference between polyp and medusa is the degree of the ECM (mesoglea) development. The new protein “mesoglein” was determined as one of the main components of mesoglea. Mesoglein is the component of the mesoglea “elastic” fibers. Previously we found that according to reverse transcription PCR mesoglein is expressed in the mature medusa exclusively in the mesogleal cells. The aim of the current work was to find out at what stage of development the specific mesoglein expression occurs. Mesogleins’ expression have been checked by PCR with specific primers on the template of transcriptomes’ cDNA from different stages; by mesogleins’ antibody staining on SDSPAGE and on paraffin sections; by histochemistry staining. In A.aurita life cycle mesoglein synthesis begins at scyphystoma polyp stage at RNA and protein level in mesogleal cells (Mc) which separated at this stage as the cell type. Mesoglein stored in granules both in Mc of adult medusa and polyp on its’ way to ECM elastic fibers. Our data perfectly correspond to the recently reported de novo transcriptome assembled from Illumina RNA-Seq data generated from six stages throughout the Aurelia life cycle.
Keywords
Aurelia aurita; Semaeostomeae; Ulmaridae; Scyphozoa; Mesoglea
Abbreviations
ASW: Artificial Sea Water; BSA: Bovine Serum Albumin; DAB: 3,3’-Di Amino Benzidine Tetrahydrochloride; ECM: Extracellular Matrix; GAGP-HRP: Anti-Guinea Pig Igg-Horseradish Peroxidase Antibody produced in Goat; MC: Mesogleal Cell; PBS-Tw: Phosphate-buffered Saline with Tween 20; PVDF: Polyvinyl Difluoride Membrane; RA47: Polyclonal Antibody against Mesoglein, produced in Rabbit; RT-PCR: Reverse Transcription Polymerase Chain Reaction; SDS-PAGE (Laemmli): Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis; WsA: A.Aurita From White Sea; ZP: Zona Pellucida
Introduction
A.aurita belongs to the family Ulmaridae order Semaeostomeae class Scyphozoa type Cnidaria [1,2]. Cnidarian is thought to possess two tissue layers: endoderm (gastroderm) and ectoderm, which are separated by the layer of extracellular compound called mesoglea [3,4]. The cnidarian radiation is relatively modest, giving rise to two major body plans (polyp and medusa) and some ten thousand extant species. A.aurita complex life cycle consists of several stages including alternating, ciliated blastula develops and invaginates to become a gastrula, the latter, known as the planula larva, leaves the mother to swim in the plankton (Figure 1). The planula settles on the sea bottom and attaches itself onto a suitable hard substratum, growing into a polyp called a scyphistoma. The scyphistoma is a tiny, sessile animal that feeds and could be kept in marine aquarium. Polyps feed by predation like small anemones and resemble Hydra in appearance [5,6]. The scyphistoma undergoes an asexual process of transverse fission called strobilation. A scyphistoma undergoing strobilation is a strobila. When the strobila is fully formed it begins to fall apart, each disk become an ephyra with most of adult features present though in different proportions. Ephyrae can hibernate on or close to the bottom during the winter, so as to rise to the surface in spring and grow up to a sexually mature adult medusa with a diameter of 30-40 cm in several months in the White Sea [3,7].
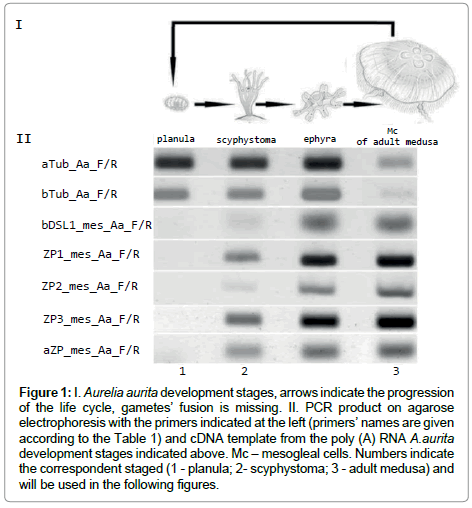
Figure 1: I. Aurelia aurita development stages, arrows indicate the progression of the life cycle, gametes’ fusion is missing. II. PCR product on agarose electrophoresis with the primers indicated at the left (primers’ names are given according to the Table 1) and cDNA template from the poly (A) RNA A.aurita development stages indicated above. Mc – mesogleal cells. Numbers indicate the correspondent staged (1 - planula; 2- scyphystoma; 3 - adult medusa) and will be used in the following figures.
The main difference between polyp and medusa is the degree of the extracellular matrix (ECM) or mesoglea development. A.aurita mesoglea contains a number of mesogleal cells (Mc). The population of Mc inside mesoglea was observed in other species of Scyphozoa and Anthozoa [8]. This feature is not unique but rather rare. Mesoglea, with its Mc, acquires a similarity with the connective tissues, derivate of mesoderm, of more advanced animals [7-9].
The new protein “mesoglein” was determined as one of the main components of mesoglea. Mesoglein mRNA cloned sequence is 1421 bp long [10]. The deduced amino acid (aa) sequence of 416 aa contains Zona Pellucida (ZP) domain and Delta/Serrate/Lag- 2 domain. The presence of the ZP domain in mesoglein allows to consider it as a member of the big family of ZP domain-containing proteins, which includes more than a thousand known members. The domain is thought to mediate the formation of extracellular protein fibers or meshworks [11], which is in full agreement with mesoglein’s position in the mesoglea: it is the component of the “elastic” fibers [12]. The name of the family originate from Zona Pellucida, which is built up of ZP domain proteins and play crucial role in fertilization [13]. Mesoglein happens to belong to ZP-domain protein family and therefore we paid attention to A.aurita oogenesis. Antibodies against mesoglein stain the plate in the place of contact of germinal epithelium and oocyte. The structure was named the ‘‘contact plate’’ and it could be the evolution precursor of the Zona Pellucida [14]. Contact plate is the external structure, kind if hat on the oocyte and ZP domain protein of contact plate differ from mesoglein [14], so this protein could not be involved in mesoglea formation. According to reverse transcription PCR mesoglein is expressed in the mature medusa exclusively in the mesogleal cells [10]. When the expression begins in development?
The aim of the current work was to find out at what stage of development the specific mesoglein expression occurs.
Materials and Methods
No specific permits were required for the described field studies; no specific permissions were required for these locations/activities; the location is not privately-owned or protected in any way; and the field studies did not involve endangered or protected species.
Animals
All experiments were carried out using A.aurita strain from the White Sea (WsA).
A.aurita medusa were collected in the vicinity of the White Sea Biological Station of the Zoological Institute RAS ‘‘Kartesh’’ (Chupa Inlet, Kandalaksha Bay in the White Sea (http://www.zin.ru/kartesh/ default_en.asp) during the summers of 2009-2013.
A.aurita (Linnaeus) has been considered a good example of such a cosmopolite [2,7]. However, recent molecular studies have revealed cryptic species in many marine taxa (Möller, 1980), suggesting that marine biodiversity is higher and opportunities for speciation have been more frequent than generally recognized. In this study, we shall call A.aurita species found in White and Barets seas, where it was first described by Linnaeus.
The methods of mesoglea and mesogleal cells (Mc) isolation have been published [12]. Planulae were collected when released from mature medusa and kept in sea water till sedimentation and transformation to scyphistoma.
Scyphistomae were cultured in artificial sea water (ASW, Reef Crystals) at 15-18ºC. Polyps were fed twice per week with freshly hatched Artemia nauplii. Strobilation in the WsA strain was induced by incubation of polyps (polyps age is about 3 years) at 10ºC for a week.
SDS-electrophoresis and immunoblot
Mesoglea polypeptides were separated by SDS-PAGE [15]. Acrylamide concentration was from 7 to 10%. Gels were stained with 0.1% Coomassie Brilliant Blue R-250.
Polypeptides separated by SDS-PAGE were transferred to polyvinylidene difluoride (PVDF, Sigma, USA) or nitrocellulose membrane (Sigma) in a Trans-blot. The membranes were blocked with 5% skimmed milk at PBS-Tw for 1 h or overnight. The membrane was then immersed in PBS-Tw with RA47 (1:5000 dilution) for 1 h, then washed with PBS-Tw three times (10 min each). The secondary antibody commercial stock (GAGP-HRP, Sigma, USA) was diluted 1/10000 (v:v) in TBS-Tw. After incubation for 1 h at room temperature with shaking, the membranes were washed twice with PBS-Tw (10 min each). The sites of enzyme binding were developed with DAB-H2O2. All operations were carried out at room temperature with shaking. Rabbit pre-immune serum instead of RA47 was used as the control and it did not produce any signal. Serum production has been published previously [12].
Immunohistochemistry
Small pieces (about 0.5 cm3) of medusas body including ectodermal layer and mesoglea, planulae and scyphistomae were fixed in 4% paraformaldehyde in filtered seawater for 24 h and dehydrate sequentially in 30%, 50% and 70% ethanol (1 h each). The fixed samples were embedded in paraffin blocks to be cut in 3–5 μm thick sections. The sections were preincubated with 5% skimmed milk in TBS-Tw for 1 h, washed 4 times for 10 min with TBS-Tw, incubated with RA47 1:5000 dilution (for 1 h), washed and incubated with HRP-conjugated goatanti- Rabbit Ig (GAR-HRP, Sigma), and visualized with microscope (Leica DM6000). For controls, some sections were incubated with nonimmune serum followed by the same subsequent steps.
Immunofluorescence (IF)
For IF study, fixed planulae were stored in 70% ethanol at -20°C for several months. All the treatment steps of immunolabeling were performed in 1.5 ml tubes on a rocker at RT. Before each step that demanded a liquid exchange, the tubes had been positioned vertically for 5 min to let the planulae settle on the bottom of the tubes. The stored whole-mount planulae were transferred through 30%, 50% and 70% ethanol (10 min each) into the 0.01 M phosphate buffered saline (PBS) and rinsed in 0.3% Triton X-100 in PBS (PBS-Tw) for 30 min. Then the planulae were incubated in 3% bovine serum albumin in PBS (PBSTw/ BSA) for 1 hour. α-Tubulin labeling was performed by overnight incubation with anti-α-Tubulin−FITC antibody (Sigma, F2168) that was diluted 1:150 in PBS-Tw/BSA; it was followed by 2 washes in PBS, 30 min each. Next the planulae’s nuclei were stained with 1 μg/ ml Hoechst 33342 (ImmunoChemistry Technologies, 639) in PBS and washed in PBS (2×15 min). Finally the planulae were mounted on glass slides in 80% glycerol in PBS. The whole-mount preparations were studied by using Leica TCS SP5 confocal microscope at the ”Taxon” Resource Center of the Zoological Institute of the Russian Academy of Sciences. Series of optical sections and maximum intensity projections were prepared in LAS AF Software (Leica Microsystems).
Histological staining of paraffin sections
Planula and scyphistoma were fixed in 4% formaldehyde solution and embedded in 1,5% agarose gel. Then the gel pieces with fixed material were dehydrated and embedded in paraffin mix Histomix (BioVitrum). Series of sections were prepared with 3-5 μm thickness on microtome Leica SM 2000R. Sections were deparaffinazed and stained with hematoxilin-eosin and Mallory’s method according standard protocols [16].
RNA isolation and cDNA synthesis
About 10 ml mesoglea jellyfish was cut into pieces and treated with 1 mg/ml Collagenase (ICN, cat N 150705) solution in sea water at 37°C for 30 min. The suspension of mesogleal cells (Mc) was centrifuged at 800 g for 20 min. Supernatant was discarded and RNA was isolated from pelleted cells by Trizol (Invitrogen) solution according to the manufacturer’s instructions. RNA samples were treated with RNAse free DNAse (Roche) to avoid DNA contamination. DNAse was removed by phenol:chloroform (2:1) extraction. About 1 μg of the RNA was reverse transcribed using oligo-dT primer (5′ T(12-20)V) primer by MMLV RT kit (Fermentas) according to the manufacturer’s recommendations. The RT reaction product was diluted 5 fold and stored at -20°C. For controls, PCR reaction with primers for A.aurita aTubulin and bTubulin have been used. The sequence have been obtained for the (http://www.compagen.org/aurelia) database (A.aurita beta-tubulin mRNA, partial cds, 1161 bp linear mRNA , Accession:AY226068.1, GI:32967427; A.aurita alpha-tubulin mRNA, partial cds 1152 bp linear mRNA Accession:AY226057.1, GI:32967405); the resulting primers are given in Table 1.
| No pairs |
Primer name | Expected length (bp) |
5'-3' – primer sequence |
|---|---|---|---|
| 1 | bDSL1_mes_Aa_F | 88 | ACAGAAGAGGCCCCTGGTGCA |
| bDSL1_mes_Aa _R | TGCAAGCAAGAAAACGATGGAGACA | ||
| 2 | ZP1_mes_Aa _F | 293 | CCCGCATCAGCTTGAAGGACGA |
| ZP1_mes_Aa _R | ACTGGAATGCGCCAAATCCATCT | ||
| 3 | ZP2_mes_Aa _F | 399 | GGATTCAGCACCACAGAATTGTGGA |
| ZP2_mes_Aa _R | GCGTTCCATGGTAGGTGTTGCAT | ||
| 4 | ZP3_mes_Aa _F | 277 | GCAACACCTACCATGGAACGCA |
| ZP3_mes_Aa _R | TCACCGCCAGCGTAGCTTGAA | ||
| 5 | aZP_mes_Aa _F | 112 | TGGTCGCTGGAGTCTGCAGTC |
| aZP_mes_Aa _R | TGTGGCTCAGCAATGACTTCATTCA | ||
| 6 | aTub_Aa_F | 322 | TGGAATTTGCAATCTACCCAGCACC |
| aTub_Aa_R | GGAGCATAGGTTGCCAGTGGG | ||
| 7 | bTub_Aa_F | 374 | GGTGGTGGTACTGGGTCTGGT |
| bTub_Aa_R | TGAAGACGTGGGAATGGCACC | ||
| No pairs |
Primer name | Expected length (bp) |
5'-3' – primer sequence |
| 1 | bDSL1_mes_Aa_F | 88 | ACAGAAGAGGCCCCTGGTGCA |
| bDSL1_mes_Aa _R | TGCAAGCAAGAAAACGATGGAGACA | ||
| 2 | ZP1_mes_Aa _F | 293 | CCCGCATCAGCTTGAAGGACGA |
| ZP1_mes_Aa _R | ACTGGAATGCGCCAAATCCATCT | ||
| 3 | ZP2_mes_Aa _F | 399 | GGATTCAGCACCACAGAATTGTGGA |
| ZP2_mes_Aa _R | GCGTTCCATGGTAGGTGTTGCAT | ||
| 4 | ZP3_mes_Aa _F | 277 | GCAACACCTACCATGGAACGCA |
| ZP3_mes_Aa _R | TCACCGCCAGCGTAGCTTGAA | ||
| 5 | aZP_mes_Aa _F | 112 | TGGTCGCTGGAGTCTGCAGTC |
| aZP_mes_Aa _R | TGTGGCTCAGCAATGACTTCATTCA | ||
| 6 | aTub_Aa_F | 322 | TGGAATTTGCAATCTACCCAGCACC |
| aTub_Aa_R | GGAGCATAGGTTGCCAGTGGG | ||
| 7 | bTub_Aa_F | 374 | GGTGGTGGTACTGGGTCTGGT |
| bTub_Aa_R | TGAAGACGTGGGAATGGCACC |
Table 1: Mesoglein and tubulin specific primers.
PCR amplification
All PCR-reactions were amplified in 25 μl reactions composed of 0.2 μM mesoglein (GenBank: ABK88269.1) [10] specific primers (tabl.1, pic.1.I), 2.5 μl of 10×PCR buffer, 3 mM MgCl2, 0.2 mM dNTPs, and 0.1 μl of Taq polymerase (Evrogen) on Eppendorf Mastercycler Personal 5332 Thermocycler. All PCRs began with a 5 min denaturation step at 94–95°C, followed by 25-30 thermal cycles including a 94°C for 20 s denaturation step, 57°C for 20 s primer annealing and elongation step – 72°C for 1 min. All PCRs terminated with a 3 min extension step at 72°C and refrigeration at 4°C or immediately used for agarose electrophoresis.
Mesoglein gene search in the transcriptome databases
The transcriptomes of the A.aurita stages exist in NCBI: mature jelly fish (SRX608573); ephyra (SRX608572); late strobila (SRX608570); early strobili (SRX608568); polyp (SRX603450); planula (SRX603448). Mesoglein sequence have been found in Transcriptome Shotgun Assembly (TSA) by tblastx [17]; its’ name is comp186661_c0_seq1 (GBRG01251588.1) in the current database. The same algorithm is used for alpha-tubulin (comp182108_c0_seq1) and beta-tubulin (comp166711_c0_seq1).
This sequences have been used for the search in developmental transcriptomes. Results are shown in Table 2.
| Sequence ID | Planula | Polyp | Early strobila | Late strobila | Ephyra | Medusa | Length seq. (bp) |
|
|---|---|---|---|---|---|---|---|---|
| comp186661_c0_seq1 | 0.9 | 2.25 | 1.5 | 1.47 | 8.56 | 26.86 | 1554 | |
| comp182108_c0_seq1 alpha-tubulin |
4,09 | 18,84 | 9,43 | 21,48 | 30,91 | 50,9 | 984 | |
| 56,57 | 105,83 | 58,85 | 50,88 | 20 | 23,51 | 631 | ||
Table 2: The amount of mesoglein transcripts in FPKM (fragments per kilobase per million fragments mapped) in each stage transcriptome. (comp186661_c0_seq1 – mesoglein sequence ID in the current A.aurita database; stages indicated above; Length seq – length of sequence in bp).
Results
Mesoglein mRNA synthesis begins at scyphystoma stage as evidenced by RT-PCR (Figure 1). Ephyra RT-PCR picture is similar to the one of scyphystoma. It happens that strobilation occurs in aquarium, but it is rather rare occation. We did not use ephyra in the following experiments for the scyphystoma is the key stage for the mesoglein mRNA occurance.
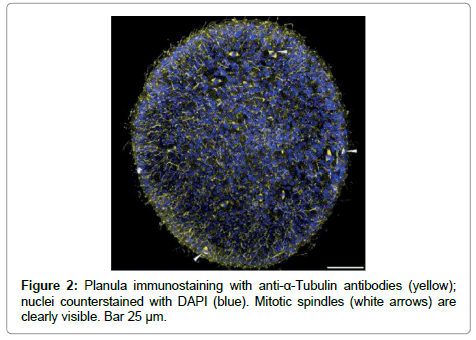
Tubulin gene and primers for it were used as the houskeeping gene control. It is visible that tubulin synthesis is more prominent in planula than in adult medusa. The prolifiration activity is supposed to be high in planula and low in adult medusa. The proliferarion activity was checked with tubulin AB. It is rather difficult to fins mitosis in Mc cells of adult medusa [18-20], but the number of mitosises is clearly visible on planula (Figure 2). So, the RT-PCR results are in accordance with the tubulin AB immunostaining.
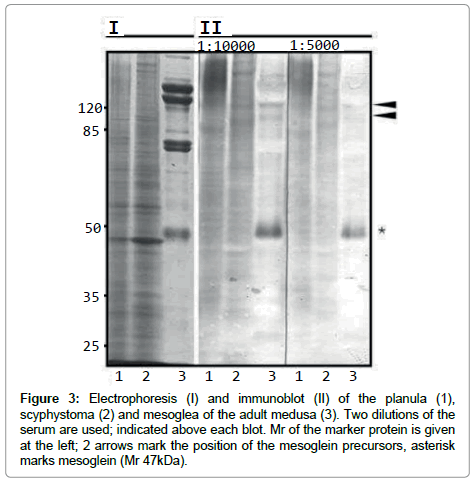
Mesoglea contains mesogleal cells (Mс). Cell proteins of the mature medusa mesoglea are not detected at the level of one-dimensional SDS-PAGE resolution (Figure 3). The major protein components of mature medusa’s mesoglea are likely to be components of ECM. One of the main mesoglea polypeptide is of apparent molecular mass (Mr) 45/47 kDa (p47) – mesoglein. Polyclonal AB raised (RA47 for Rabbit Antibodies) against mesoglein stain p47 itself in mesoglea preparations (Figure 3). In Mc p47 is a minor stained zone, but the most prominent are zones with higher Mr [12]. These high zones are stained in mesoglea when low dilution of serum with RA47 is used (Figure 3, II, arrows). We suppose that mesoglein is synthesised by cells (Mс) as a high molecular mass precursor and it undergoes restricted proteolysis during incorporation in ECM. Immunoblot reveals high Mr bands among the female gonads (Gf) germinal epithelium polipeptides. The high Mr bands in planula (~180 kDa) could be the maternal proteins’ remnant while mesoglein synthesis not yet switched on (Figure 3, II, 1). The mesoglein synthesis switched on at scyphistoma stage looks like 2 bands with the Mr similar to that of the Mc from mesoglea (Figure 3, II, arrows)
Figure 3: Electrophoresis (I) and immunoblot (II) of the planula (1), scyphystoma (2) and mesoglea of the adult medusa (3). Two dilutions of the serum are used; indicated above each blot. Mr of the marker protein is given at the left; 2 arrows mark the position of the mesoglein precursors, asterisk marks mesoglein (Mr 47kDa).
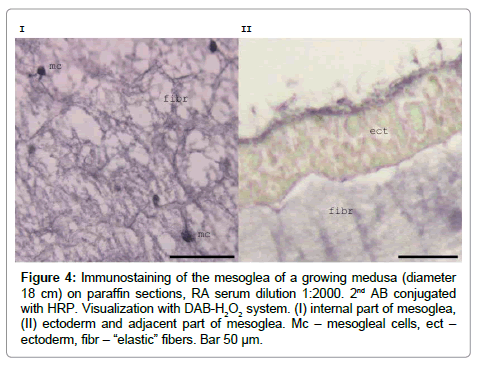
Immunolabelling with RA47 shows that fibrils in the mesoglea and Mc are stained intensely in the adult medusa (Figure 4). Granules in the Mc of the growing medusa have very pronounced staining in paraffin sections [12]. The cell contours are blurred and the cells look as though they are surrounded by weakly stained material. In the epidermis, the antigen is distributed throughout the cell cytoplasm but immunostaining is most prominent in the apical parts of the cells (Figure 4). We assumed that mesogleal cells of A.aurita produce mesoglein, which is a component of the ‘‘elastic’’ fibres. Mc definitely participates in mesoglein production and might play a role in mesoglein modification during ‘‘elastic’’ fibre formation [12].
Figure 4: Immunostaining of the mesoglea of a growing medusa (diameter 18 cm) on paraffin sections, RA serum dilution 1:2000. 2nd AB conjugated with HRP. Visualization with DAB-H2O2 system. (I) internal part of mesoglea, (II) ectoderm and adjacent part of mesoglea. Mc – mesogleal cells, ect – ectoderm, fibr – “elastic” fibers. Bar 50 μm.
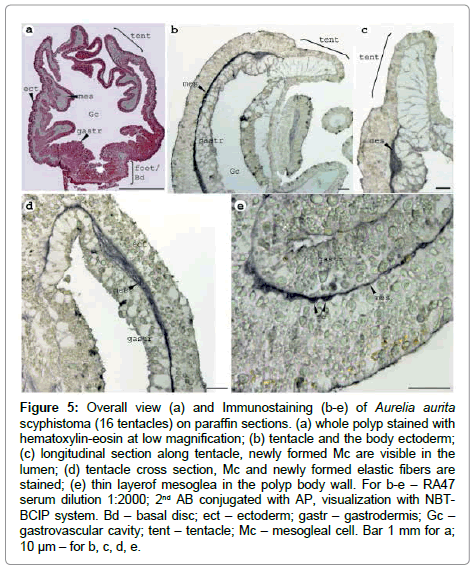
The mesoglea of polyps (16 tentacles stage) is a thin layer in the wall of the body and enters the tentacles (Figure 5). The first cells begin to appear probably at this stage, so immunostaining of the scyphistoma was done in order to determine the compartment of the initial mesoglea synthesis. It is visible that Mc arise at the border of the epidermis and gastroderma (Figure 5). They are collected at the newly formed lumen of the tentacle (Figure 5) and it is possible to find cell abut to the basal membrane (Figure 5). So, Mс are the cells first to express mesoglein and they arise as the cell type together with the synthesis beginning. The weak staining in the ectoderm is localised to the basal part (part close to the basal membrane) of the cells in contrast to the adult medusa (Figure 5, b versus Figure 4,II).
Figure 5: Overall view (a) and Immunostaining (b-e) of Aurelia aurita scyphistoma (16 tentacles) on paraffin sections. (a) whole polyp stained with hematoxylin-eosin at low magnification; (b) tentacle and the body ectoderm; (c) longitudinal section along tentacle, newly formed Mc are visible in the lumen; (d) tentacle cross section, Mc and newly formed elastic fibers are stained; (e) thin layerof mesoglea in the polyp body wall. For b-e – RA47 serum dilution 1:2000; 2nd AB conjugated with AP, visualization with NBTBCIP system. Bd – basal disc; ect – ectoderm; gastr – gastrodermis; Gc – gastrovascular cavity; tent – tentacle; Mc – mesogleal cell. Bar 1 mm for a; 10 μm – for b, c, d, e.
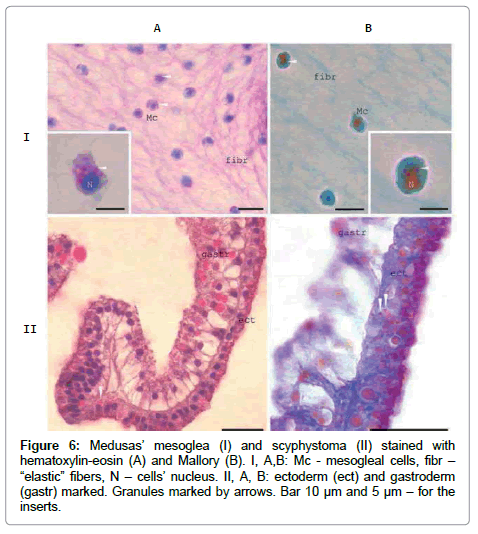
Antibodies exhibit cross-reactivity due to the presence of several ZP-domain proteins [14]. According to the sequencing of at least 5 [1]. density and at the ends of the endoplasmic reticulum tubes [12]. The very similar granules observed in newly formed Mc in scyphystoma polyp and in some ectodermal cells (Figure 6). The crimson / raspberryred colour of the granules in Mallory staining indicate high rate of the proteins’ glycosylation inside them.
Figure 6: Medusas’ mesoglea (I) and scyphystoma (II) stained with hematoxylin-eosin (A) and Mallory (B). I, A,B: Mc - mesogleal cells, fibr – “elastic” fibers, N – cells’ nucleus. II, A, B: ectoderm (ect) and gastroderm (gastr) marked. Granules marked by arrows. Bar 10 μm and 5 μm – for the inserts.
In A.aurita life cycle mesoglein synthesis begins at scyphystoma polyp stage at RNA (Figure 1) and protein level (Figure 3) in mesogleal cells (Mc) which separated at this stage as the cell type (Figure 5). Mesoglein stored in granules both in Mc of adult medusa and polyp on its’ way to ECM elastic fibers (Figure 6).
Disccussion
Cnidarians represent one of the most basal animal groups in which complex life cycles are present. The life cycle of Medusozoa (Hydrozoa, Cubozoa, and Scyphozoa) consists of two morphologically disparate generations with three well-defined life stages - planula larvae, polyp, and jellyfish (Figure 1). Transition from one stage into another is tightly regulated by environmental stimuli and depends on seasonal rhythms [21-24].
Nowadays the comparative “gene approach” studies make obvious a similar gene pattern in the same differentiation pathways in distant species to such an extent that the very term “tissue” becomes uncertain. For instance, if A.aurita possesses striated muscles with gene expression characteristic of the myogenic pathway [25,26] could they be designated as a special tissue in spite of their origin?
I. Metchnikoff in his “Lectures on the comparative pathology of inflammation” [27] made one of the first attempts to judge the evolutionary attitude of different tissues, especially tissues of the internal medium. He stressed the importance of the simple key link determination in complicated processes and the necessity to follow their evolution from the simplest forms into the higher orders. This way of thinking proved to be productive for it brings up the whole immunology that arises from phagocytes of starfish larva. Zawarzin [28] followed this ideology with his work on regularities of tissues evolution. The work was summarized in the theory of parallelism, which postulated, for example, a prominent similarity of external epitelia, in distant species. On morphological level the similarity between tissues with similar functions seems to be more prominent than between different tissues of one organism. This theory was not widely spread but it gave the basic paradigm for decades of research for Russian biologists. Zawarsin wrote “Comparative method is the main one to comprehend the evolution of the tissue system. But only well established subjects are worthy of comparison. If one would like to compare, for example, medusa mesoglea with tissues of vertebrate internal medium, the comparison should be done not with some specific jelly-like tissue such as external layer of tadpole tail, but with the complete system made up of the blood, connective tissue, cartilages and bones, on the whole” [28].
The “candidate gene approach” makes it possible to check whether there are any of the known vertebrate genes, from all of the tissues mentioned, expressed in medusa, but such a way leads nowhere if some special gene, absent in other animals, makes mesoglea so different from the rest of the internal medium tissues. Our attempts to determine the mesoglea protein composition lead to the description of mesoglein as one of the main mesoglea components [10]. Now we determined the beginning if its’ expression.
It has been found that during metamorphosis into a polyp, cells in the planula endoderm, but not in the ectoderm, become strongly caspase 3 immunoreactive, suggesting that the planula endoderm, in part or in its entirety, undergoes apoptosis during metamorphosis. The polyp endoderm seems to be derived from the planula ectoderm in Aurelia, implicating the occurrence of “secondary” gastrulation during early metamorphosis [29]. In contrast, the planula endoderm in hydrozoans may contain the future polyp endoderm as well as progenitors to the polyp ectoderm. Thus, the planula “ectoderm” and “endoderm” of one species cannot be assumed developmentally equivalent to the planula “ectoderm” and “endoderm” of another species in cnidarians. If this is the case, expression patterns of genes may be expected to differ in planulae of species with different modes of “gastrulation” [29].
The mesoglea is mostly acellular tissue, but some species (including A.aurita) contain amoebocytes - mesogleal cells (although this is quite rare). So, the long on-going discussion about possible A.aurita triploblasty, based on the similarity of mesoglea to the internal tissues of higher animals [28] and the expression pattern of target genes [27], could have the paradoxical solution – A.aurita is a one-layer organism since all adult tissues are the derivates of the ectoderm. All our results confirm that Mc and, probably, the whole mesoglea originate from ectoderm [10,12]. Still, mesoglein expression is specific for the mesogleal cells of adult medusa.
It is obvious that mesoglein RNA appears at planula-to-polyp transition (Figure 1). The successive stages of polyp-to-jellyfish are relatively well investigated [30]. The proteins, which are mesoglein AB reveal in polyp have Mr slightly higher that the ones in adult Mc (Figure 3). This could happens due to the different rate of glycosylation, which existence confirmed by histochemistry (Figure 6). We already observed mesogleins’ high Mr precursors in Hydra [12], which appearance is quite similar to the scyphystoma polyp. We assume that both animals, Hydra sp. and A.aurita medusa, synthesize mesoglein as a high molecular mass precursor, but posttranslational modifications leading to the appearance of the mesoglea are more extensive in the medusa. Obviously, such modifications not occur in scyphystoma polyp until strobilation, i.e. ephyra formation. A range of additional work would be merited to find out at what stage of development the specific post-translational modifications occur.
Previously we observed the similarity between the epidermal and Mc polypeptide patterns, which argues that Mc originate from the epidermal layer or it’s derivate [12]. Now we can add to these argument the fact that at the very beginning of Mc formation both ectotermal cells and Mc bear mesoglein label (Figure 5) and both cell types bear same type of granules (Figure 6). No such a staining was ever observed in the gastroderm.
Recently the de novo transcriptome was assembled from Illumina RNA-Seq data generated from six stages throughout the Aurelia life cycle [1]. The amount of mesoglein transcripts in each stages’ transcriptome have been counted (Table 2).
The figures perfectly correspond to our finding: mesoglein transcripts appear at the polyp stage, the transcripts’ amount rise in ephyra and reach its maximum in adult medusa.
In the current work we determined the beginning of mesoglein synthesis at planula-to-polyp transition and traced the synthesis to the newly formed mesogleal cells (Mc) which separated at this stage as the cell type. Mc do express gene different from ectodem or gastroderm, which make them an independent population. Still, the specific synthesis does not make them the founder of the third layer. The complication of the Cnidarian composition at cytological and molecular biology level just calls for the new biological language.
More and more answers come in molecular biology terms – with precise figures, gene patterns and sequences. This type of information does not agree happily with the way of thinking in classical terms such as “tissues” or “germ layers”. There is an urgent necessity of a new biological language, one combining the classical point of view with the formal and exact computer- and databases-based language of modern biology.
Conclusion
Mesoglein is one of the main A.aurita mesoglea proteins. In the current work we determined the beginning of mesoglein synthesis at planula-to-polyp transition and traced the synthesis to the newly formed mesogleal cells (Mc). The amount of mesoglein transcripts in each stages’ transcriptome [1] perfectly correspond to our finding. Mesoglein synthesis begins at scyphystoma polyp stage at RNA and protein level in mesogleal cells (Mc) which separated at this stage as the cell type. Our previous and current results do not support the idea of mesoglea being the third layer in spite of specific synthesis in Mc. It looks like all adult medusa tissues originate from ectoderm.
Competing Interest
The authors declare that they have no competing interests.
Acknowledgements
The authors wish to express gratitude to Dr. I.Matveev who provided the foundation for this work. We deeply appreciate the help that we have always received at the White Sea Biological Station of the Zoological Institute RAS ‘‘Kartesh’’. This work was supported by the Russian Foundation for Basic Research (grants 09-04-01145-a, 13-04-01739-a, 16-34-00603-youth), grant from presidium RAS (MCB) and Russian Science Foundation grant (15-15-20026).
References
- Brekhman V, Malik A, Haas B, Sher N, Lotan T (2015) Transcriptome profiling of the dynamic life cycle of the scypohozoan jellyfish A.aurita. BMC Genomics.
- Kramp PL (1961) Synopsis of the Medusae of the World. J Mar BiolAssoc UK 40: 469.
- Dogel VA (1937) Phylum Coelenterata. In: Manual in zoology (ed. VA Dogel, YIPoljanskii). Moscow-Leningrad: Nauka 1: 268-369.
- Chapman DM (1974) Cnidarian histology. In Coelenterate biology (ed L. Muskatine, H. Lenhoff). Reviews and new perspectives. New York - San Francisco - London: Academic Press pp: 1-93.
- Tardent P (1978)Coelenterata, Cnidaria. In: Morphogenesis der Tiere (ed. T. Seidel). New York: Gustav Fischer Verlagpp: 71-391.
- Purcell JE (1991) A review of cnidarians and ctenophores feeding on competitors in the plankton. Hydrobiologia216/217: 335-342.
- Arai MN (1997) A functional biology of scyphozoa. London: Chapman and Hall pp: 28-206.
- Chapman G (1966) The structure and function of the mesoglea. In: Cnidaria and their evolution (ed. WJ Rees). SympZoolSocLond 16: 147-168.
- Hyman LH (1940) The invertebrates: protozoa through Ctenophora. pp. 726. New York, London: McGraw Hill.
- Matveev IV, Shaposhnikova TG, Podgornaya OI(2007) A novelA.auritaproteinmesogleincontains DSL and ZP domains. Gene 399: 20-25.
- Jovine L, Darie CC, Litscher ES, Wassarman PM (2005) Zona Pellucidadomainproteins. AnnuRevBiochem 74: 83-114.
- Shaposhnikova T, Matveev I, Napara T, Podgornaya O (2005) Mesogleal cells of the jellyfishA.auritaare involved in the formation of mesoglealfibres. Cell BiolInt 29: 952-958.
- Bork P, Sander C (1992) A large domain common to sperm receptors (Zp2 and Zp3) and TGF-beta type III receptor. FEBS Lett 300: 237-240.
- Adonin LS, Shaposhnikova TG, Podgornaya O (2012) A.aurita(Cnidaria) oocytes' contact plate structure and development.PLoS One 7: e46542.
- Laemmli UK (1970) Cleavage of structural protein during the essambly of the head of the bacteriophage T4. Nature 4: 680-682.
- Pearse AGE (1958)Hystochemistry, Therioretical and Applied. Vol.1. 3rd ed. Edinburgh: Churchill Livingstone.
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25: 3389-3402.
- Napara TO, Oskol’sky AA, Chaga OY(1994) Mesogleal cells of A.aurita (Cnidaria, Scyphozoa). I. Morphology of mesogleal cells and of the matrix of mesoglea. Tsitologiia 36: 623-630.
- Napara TO, Oskol’sky AA, Chaga OY (1996a) Mesogleal cells of A.aurita (Cnidaria, Scyphozoa). II. Ultrastructural and histochemical analysis. Tsitologiia 38: 456-464.
- NaparaTO, Oskol’sky AA, Shaposhnikova TG, Chaga OY (1996b)Mesogleal cells of A.aurita(Cnidaria, Scyphozoa). III. An autoradiographic analysis of the synthesis of extracellular matrix of mesoglea. Tsitologiia 38: 465-474.
- Hofmann DK, Neumann R, Henne K(1978)Strobilation, budding andinitiation of scyphistomamorphogenesis inthe rhizostome\ Cassiopeaandromeda (Cnidaria, Scyphozoa). Mar Biol 47: 161-176.
- Leitz T, Wagner T (1993) The marine bacterium Alteromonasespejianainduces metamorphosis of the hydroid Hydractiniaechinata. Mar Biol 115: 173-178.
- Kroiher M, Siefker B, Berking S (2000) Induction of segmentation in polyps of A.aurita (Scyphozoa, Cnidaria) into medusa and formation of mirror-image medusa anlagen. Int J DevBiol 44: 485-490.
- oiher M, Siefker B, Berking S (2000) Induction of segmentation in polyps of A.aurita (Scyphozoa, Cnidaria) into medusa and formation of mirror-image medusa anlagen. Int J DevBiol 44: 485-490.
- Ruppert EE (1991) Introduction to the aschelminth phyla: a consideration of mesoderm, body cavities, and cuticle. In: Microscopic Anatomy of Invertebrates (ed. FW. Harrison), New York: Wiley-Liss V 4: 1-17.
- Spring J, Yanze N, Middel AM, Stierwald M, Groeger H, et al. (2000)Ancestral role of the mesoderm specification factor twist in the life cycle of jellyfish. DevBiol 228: 363-375.
- Miyake H, Terazaki M, KakinumaY (2002) On the polyps of the common jellyfish A.aurita in Kagoshima Bay JOceanogr 58: 451-459.
- Zawarzin A (1945) Essays on the evolution histology of blood and connective tissue. Moscow: Medgizpp: 130-200.
- Seipel K, Schmid V (2006) Mesodermal anatomies in cnidarian polyps and medusae. Int J DevBiol 50: 589-599.
- Yuan D, Nakanishi N, Jacobs DK, Hartenstein V (2008) Embryonic development and metamorphosis of the scyphozoan Aurelia. Dev Genes Evol 218: 525-539.
Relevant Topics
- Algal Blooms
- Blue Carbon Sequestration
- Brackish Water
- Catfish
- Coral Bleaching
- Coral Reefs
- Deep Sea Fish
- Deep Sea Mining
- Ichthyoplankton
- Mangrove Ecosystem
- Marine Engineering
- Marine Fisheries
- Marine Mammal Research
- Marine Microbiome Analysis
- Marine Pollution
- Marine Reptiles
- Marine Science
- Ocean Currents
- Photoendosymbiosis
- Reef Biology
- Sea Food
- Sea Grass
- Sea Transportation
- Seaweed
Recommended Journals
Article Tools
Article Usage
- Total views: 13615
- [From(publication date):
December-2016 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 12666
- PDF downloads : 949