Research Article Open Access
Modular Ultrasonic Lysis System for Rapid Nucleic Acid Extraction and Sample Transfer of Bacillus Spores
Matthias Geissler1*#, Sandra Isabel2#, Benoît Voisin1, Chantal Fauvel2, Maurice Boissinot2, Michel G. Bergeron2* and Teodor Veres1,3
1National Research Council of Canada, Boucherville (Quebec) J4B 6Y4, Canada
2Centre de recherche en infectiologie de l’Université Laval, Quebec (Quebec) G1V 4G2, Canada
3INRS-EMT, Varennes (Quebec) J3X 1S2, Canada
#These authors contributed equally to this work
- *Corresponding Author:
- Matthias Geissler
National Research Council of Canada, 75 de Mortagne Boulevard
Boucherville (Quebec) J4B 6Y4, Canada
Tel: +1 450 641-5388
Fax: +1 450 641-5105
E-mail: matthias.geissler@cnrc-nrc.gc.ca - Michel G. Bergeron
Centre de recherche en infectiologie de l’Université Laval
Centre de recherche du CHUQ, 2705 Boulevard Laurier
Quebec (Quebec) G1V 4G2, Canada
Tel: +1 418 654-2705
Fax: +1 418 654-2715
E-mail: Michel.G.Bergeron@crchul.ulaval.ca
Received Date: August 13, 2012; Accepted Date: November 05, 2012; Published Date: November 10, 2012
Citation: Geissler M, Isabel S, Voisin B, Fauvel C, Boissinot M, et al. (2012) Modular Ultrasonic Lysis System for Rapid Nucleic Acid Extraction and Sample Transfer of Bacillus Spores. J Bioterr Biodef 3:119. doi: 10.4172/2157-2526.1000119
Copyright: © 2012 Geissler M, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Bioterrorism & Biodefense
Abstract
This paper describes the design, functioning and use of an ultrasonic modular system intended for rapid extraction and fragmentation of DNA from microbial organisms following sample collection in the field. PCR assessment of the DNA extracts revealed that the system can disrupt Bacillus atrophaeus spores, a simulant for Bacillus anthracis, in less than 1 min, providing a DNA yield equivalent to that of a commercial nucleic acid extraction method. Simulation of the transfer from a contaminated to a secure area confirmed that the sample remained confined within the module while the exterior surface can be decontaminated through immersion in a disinfectant solution.
Keywords
Bacillus spores; Sonication; Spore lysis; PCR; Sample transfer
Abbreviations
DARE: Dual-filter method for Applied Recovery of microbial particles from Environmental and powdery samples; FITC: Fluorescein Isothiocyanate; FULM: Fluidic Ultrasonic Lysis Module
Introduction
Bacillus anthracis is the causative agent of anthrax disease and was classified by the Centers for Disease Control and Prevention in the category A of high-priority biological agents [1]. B. anthracis forms dormant endospore structures (spores) to survive in harsh environments over long periods of time. It has been estimated that the release of 100 kg of aerosolized anthrax spores over a metropolitan area could cause between 130,000 and 3 million deaths in favorable environmental conditions [2]. Hence the rapid detection of B. anthracis spores is crucial to public health or life stock management at a contaminated site.
Prior to the detection of biothreat agents, a number of steps involving sample collection, disinfection and transportation as well as concentration and purification of targeted analyte are typically required [3,4]. These steps can complicate the detection process and render on-site testing cumbersome. Also, sensitive lysis and detection equiments are often incompatible with decontamination procedures that follow deployment in the field. For these reasons, first responders inspecting a potentially contaminated area are commonly performing risk assessment rather than in-depth analysis, and samples collected at the scene are preferably transferred to a secured environment equipped with biological safety cabinets for analysis and follow-up investigation [4,5]. The work presented in this paper is responding to this task: we devised a fluidic ultrasonic lysis module (FULM) for collecting samples in the field and bringing them to a designated laboratory facility for further processing with a decontamination step in-between.
Sonication has found increasing attention in the development of portable, point-of-care instrumentation since it promotes the extraction of nucleic acids from bacterial spores or other microbial forms in a timely and technologically convenient fashion [6-11]. This technology exploits ultrasound waves (with frequencies typically ranging from ~20 kHz to the MHz regime) dissipating within a liquid medium. The exerted stress can disrupt cell envelopes, giving way for intracellular content to be released in the surrounding (mostly aqueous) buffer solution. Lysis performed in a closed and miniaturized system typically requires coupling of ultrasound waves into a liquid reservoir via a flexible interface. Such systems commonly employ polymer materials as the interface in conjunction with microparticles (e.g., glass beads) in the liquid phase to assist the lysis process [6,7,9,11]. However, microbeads render the design of microfluidic systems more complex. A bench-top sonication instrument that circumvents the need of micobeads has been devised by Chandler et al. [8] and Warner et al. [11]. In their set-up, acoustic energy is transmitted from a piezoelectric transducer to a water reservoir in which a tubular flow-through system is embedded that contains the sample. Marentis et al. [10] have developed a fluidic device entailing a microfabricated piezoelectric sonicator. The authors have demonstrated that spores of Bacillus subtilis can be lysed efficiently using ultrasound without the need of microbeads, though the miniaturized format mainly suits microvolume analysis. In practice, suspicious powders harvested at a potentially contaminated site are often suspended in larger volumes, as presented for eight different DNA extraction methods that have been tested in this context (e.g., 350 μL–2 mL) [12]. Herein, we describe an ultrasonic modular system using a stainless steel interface intended for rapid (<1 min) and efficient extraction and fragmentation of DNA from a 230 μL suspension of Bacillus spores to perform PCR analysis. We used spores of Bacillus atrophaeus subsp. globigii which is mostly inoffensive, yet produces spores similar to those of B. anthracis [13], making it a suitable candidate for research and development of detection technologies.
Materials and Methods
Equipment design and function
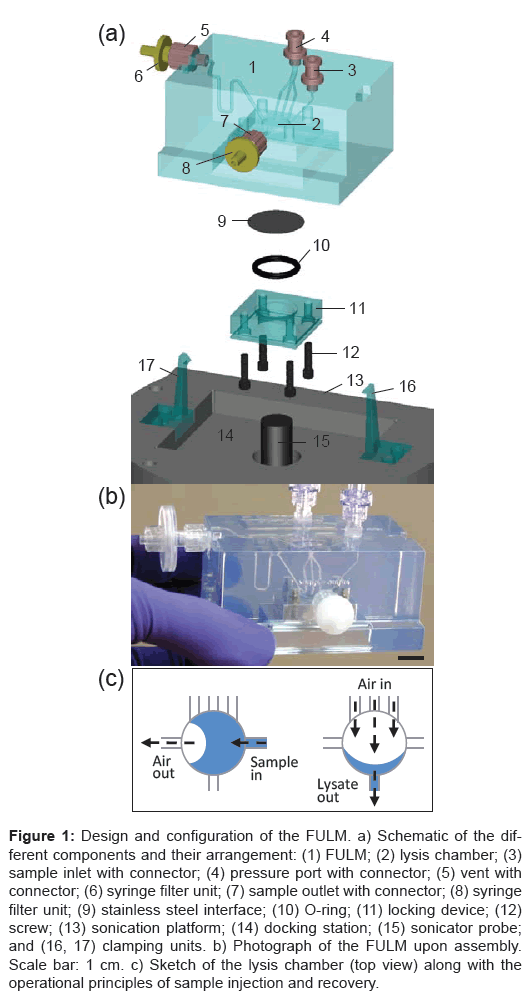
The set-up comprises two principal components —; the FULM and a sonication platform – whose design and arrangement are depicted in Figure 1a. The FULM constitutes a block (measuring 80 mm in length, 38 mm in height, and 51 as well as 59 mm in width at the top and bottom, respectively) that was custom-fabricated from transparent polymer resin (SLA 11122) using high-precision 3D stereolithography (Axis Prototypes, Saint-Léonard, QC). Internal supply channels (ranging from 0.5 to 1.0 mm in diameter) connect the lysis chamber with macroscopic access points located at the periphery of the FULM. Sample inlet and pressure port are equipped with a check valve connector (Qosina, Edgewood, NY) for receiving a luer lock tip syringe. The vent and sample outlet both entail luer lock connectors adapted for mounting a filter unit. We used a 17-mm-diameter, 0.2-μm-pore-size PTFE filter unit (Qosina) and a 13-mm-diameter, 0.22-μm-pore-size Millex-GV filter unit (Millipore, Billerica, MA) for vent and sample outlet, respectively. The exit of the filter unit was enclosed by a cap to seal the interior of the FULM. The FULM contains a circular lysis chamber which is enclosed by an interface punched out of 0.001˝ stainless steel shim stock (corresponding to 25.4 μm in thickness, Daemar Canada, Dorval, QC). The interface is held in place by an O-ring and a designated locking device fastened to the FULM using four screws. The locking device provides a vertical through-hole (15 mm in diameter) so that the interface can be accessed from below by a sonicator probe.
Figure 1: Design and configuration of the FULM. a) Schematic of the different components and their arrangement: (1) FULM; (2) lysis chamber; (3) sample inlet with connector; (4) pressure port with connector; (5) vent with connector; (6) syringe filter unit; (7) sample outlet with connector; (8) syringe filter unit; (9) stainless steel interface; (10) O-ring; (11) locking device; (12) screw; (13) sonication platform; (14) docking station; (15) sonicator probe; and (16, 17) clamping units. b) Photograph of the FULM upon assembly. Scale bar: 1 cm. c) Sketch of the lysis chamber (top view) along with the operational principles of sample injection and recovery.
The sonication platform entails a commercial ultrasound generator (Sonics & Materials, Newtown, CT) and a titanium alloy probe (13 mm in diameter, Sonics & Materials) fixed inside a free-standing socket. The interior of the socket, custom-fabricated from aluminum parts, is surrounded with a vibration dampening material (foam) to reduce noise during sonication. The tip of the sonicator probe extends vertically from the docking station located at the top of the socket destined to accommodate the FULM. Two flexible clamps (SLA 11122) are placed on opposite sides of the docking station to interlock with the FULM upon its placement, preserving contact between the sonicator probe and the stainless steel interface for the duration of the lysis process. An external power supply (Vibra-Cell ultrasonic processor, Sonics & Materials) was used as a control unit to regulate the sonication process.
Preparation of spores
Spores of B. atrophaeus subsp. globigii strain CCRI-9827 (Collection du Centre de recherche en infectiologie, Quebec, QC; https://www.wfcc.info) were prepared according to published procedures [14] and diluted in 1X phosphate-buffered saline (PBS, 137 mM NaCl, 6.4 mM Na2HPO4, 2.7 mM KCl, 0.88 mM KH2PO4, pH=7.4, Sigma-Aldrich Canada, Oakville, ON). Fluorescence labelling was performed by incubating spores with fluorescein isothiocyanate (FITC) as described previously [15].
DNA extraction
For DNA extraction, 230 μL of suspension of B. atrophaeus CCRI-9827 (3.0×105 spores mL-1 in PBS) were injected into the FULM using a 1-mL syringe. Lysis experiments were performed at a frequency of 20 kHz and 30 μm amplitude using variable exposure times to optimize the yield of extracted nucleic acids (Figure 2). The BD GeneOhmTM Lysis Kit (BD Diagnostics, Quebec, QC) was used according to the recommendations provided by the supplier, serving as a DNA extraction reference method for each series of experiments. Specifically, 100 μL of the diluted spore suspension were transferred into a designated lysis tube followed by agitation on a Vortex-Genie 2 (Scientific Industries, Bohemia, NY) at the maximum adjustable speed for a period of 5 min. For comparison and conservation purposes, all DNA extracts were heated immediately at 95°C for 2 min as recommended by the supplier of the BD GeneOhmTM Lysis Kit [16]. Highly purified genomic DNA serving as a PCR reference sample was prepared from mid-log-phase cells of B. atrophaeus strain CCRI-9827 and B. subtilis ATCC 15841 (American Type Culture Collection, Manassas, VA) using MagneSil KF isolation kit (Promega, Madison, WI) on a KingFisher ML instrument (Thermo Fisher Scientific, Waltham, MA).
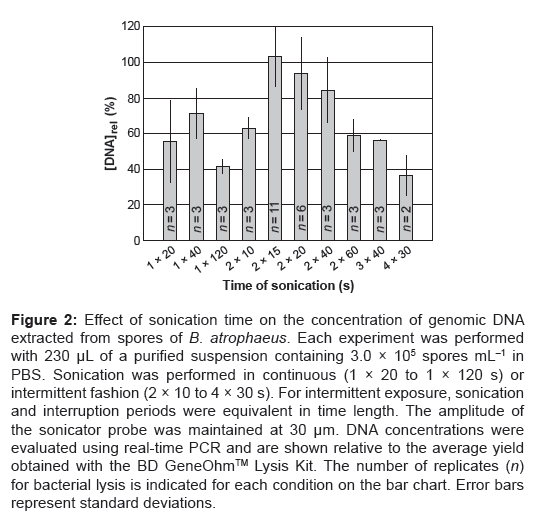
Figure 2: Effect of sonication time on the concentration of genomic DNA extracted from spores of B. atrophaeus. Each experiment was performed with 230 μL of a purified suspension containing 3.0 × 105 spores mL–1 in PBS. Sonication was performed in continuous (1 × 20 to 1 × 120 s) or intermittent fashion (2 × 10 to 4 × 30 s). For intermittent exposure, sonication and interruption periods were equivalent in time length. The amplitude of the sonicator probe was maintained at 30 μm. DNA concentrations were evaluated using real-time PCR and are shown relative to the average yield obtained with the BD GeneOhmTM Lysis Kit. The number of replicates (n) for bacterial lysis is indicated for each condition on the bar chart. Error bars represent standard deviations.
Analysis of DNA extracts
Real-time PCR assays to amplify a 212-bp fragment of the atpD gene of B. atrophaeus were performed using primers (ABgl158 and ABgl345a) along with a TaqMan probe (ABgl-T1-B1) as disclosed in previous work [14]. PCR reactions were performed with 25-μL mixtures each of which contained 5 μL of the lysate using conditions [17] and a standard curve of purified DNA [15] that are described elsewhere. Negative controls containing no DNA template were included in each series of PCR analysis. For all purified spore preparations that were used, non-lysed spores were tested as control samples. ANOVA statistical analysis of PCR data was performed using JMP statistical software from SAS (Cary, NC).
DNA fragmentation
To study the degree of DNA fragmentation, 230 μL of a solution containing 15 ng μL-1 purified genomic DNA of B. subtilis ATCC 15841 was sonicated using the FULM for the duration that provided the highest DNA yield (i.e., 2×15 s). Analyte solutions (20 μL) were subjected to gel electrophoresis using 0.8% agarose with 0.25 μg mL-1 of ethidium bromide in Tris-borate-EDTA buffer (89 mM Tris, 89 mM boric acid, 2 mM EDTA). A 1X DNA mass ladder and a 1-kb DNA ladder (Invitrogen, Carlsbad, CA) were used for quantification and estimation of DNA fragment lengths, respectively.
Fluorescence imaging
Fluorescence imaging was done using an Olympus FV300 confocal laser scanning microscope (Olympus Canada, Markham, ON) equipped with an argon-ion laser for excitation at 488 nm. Suspensions of treated (sonication using the FULM for 2×15 s) and untreated spores were diluted 1:1 (v/v) with a warm solution of 1% agarose (VWR International, West Chester, PA) in 1X PBS and sandwiched between a glass microscope slide and a cover slip prior to inspection [15].
Decontamination of the FULM
Following sample injection, the FULM was closed and immersed in a 0.5% solution of sodium hypochlorite for variable periods of time (i.e., twice for 5, 15 and 60 min) to evaluate any possible effect on PCR detection. Upon removal from the bath, the FULM was rinsed twice with sterile water and dried using cotton gauze before proceeding with the lysis and PCR experiments as described above. In addition, 230 μL of a more concentrated suspension of B. atrophaeus (0.5 McFarland, ~1×107 spores mL-1) were injected in the FULM using a 1-mL syringe. Once closed, the module was immersed in 0.5% sodium hypochlorite solution for 5 min with gentle agitation, followed by rinsing with sterile water and drying with sterile cotton gauze. To evaluate the presence of viable spores on the exterior of the FULM, we swabbed five selected sites of the FULM - stainless steel interface, sample inlet, sample outlet, pressure port, and vent - before and after the decontamination process, as well as after subsequent sonication. The collected samples were immediately inoculated on brain heart infusion agar and incubated overnight at 37°C under aerobic conditions to monitor the number of colony-forming units (CFU).
Simulated detection of Bacillus spores in suspect powders
In the simulated contaminated area, 20 mg of bentonite (Aldrich Chemical Company, Inc., Milwaukee, WI) were suspended in 1 mL PBS containing 3.0×105 spores mL-1. Recovery of clean bacterial spores from these powdery samples (DARE procedure) was performed twice as described previously [17]. The FULM was decontaminated in 0.5% sodium hypochlorite solution for 5 min and then transferred to a simulated secure area where lysis (using the FULM and BD GeneOhmTM Lysis Kit) and PCR experiments were performed as described above.
Results
Operation of the FULM and volume handling
The handy size of the FULM is intended to facilitate transport and manipulation by first responders wearing bulky rubber gloves as part of their protective gear. The assembled module shown in Figure 1b requires only few manual steps including injection of a bacterial spore sample, sonication, and recovery of the lysate. The sample (230 μL) is inserted to completely fill the lysis chamber with a 1-mL syringe at the injection port with the vent in the open position, as illustrated in Figure 1c. During this step, the pressure port and the outlet are closed. The opposite arrangement of incoming and outgoing supply channels facilitates displacement of air and promotes uniform filling of the lysis chamber. The shallow reservoir (1.5 mm in depth over 14 mm in diameter) thereby helps distribute the sample evenly in the acoustic field (without the need for microbeads). Recovery of the sample is performed by aspirating the lysate through the 0.22-μm-pore-size Millex-GV filter unit at the sample outlet using a 1-mL syringe while injection port and vent both remain closed. The recovery process can be assisted by pushing air into the system using a syringe at the pressure port. Volumes of recovered lysates were typically 175–185 μL without filtration and 90–125 μL with filtration. Hence, the dead volumes were estimated to be 55–45 μL in the FULM and 50–95 μL in the Millex-GV filter.
DNA extraction and fragmentation
We studied the effect of sonication time on the yield of amplifiable DNA (Figure 2) compared to a high-yield-providing, benchmark DNA extraction process (BD GeneOhmTM Lysis Kit). Negative controls without added DNA template as well as non-lysed spores, used as another control, did not produce positive PCR signals. Continuous sonication ranging from 20 s to 2 min (i.e., 1×20 to 1×120 s) resulted in moderate amounts of DNA, yielding 71.4 ± 13.9% at best when the sample was treated for 40 s consecutively. Interestingly, the DNA yield increased to 93.7 ± 20.0% when the sample was sonicated twice for 20 s with a 20-s interruption in between (i.e., 2×20 s). Two intermittent sonication steps of 15 s each with 15 s interruption time (i.e., 2×15 s) turned out to be the most productive condition, providing a relative DNA yield of 103.1 ± 16.5%, on average, which is comparable to that of the BD GeneOhmTM Lysis Kit. Statistical analysis of PCR average cycle thresholds (Ct) revealed no significant difference between results for DNA extracts obtained from sonication for 2×15 s (Ct=30.37 ± 1.09) and BD GeneOhmTM Lysis Kit (Ct=30.52 ± 1.24). Since ultrasounds are known to fragment DNA, longer lysis durations may lead to higher fragmentation of genomic material [18] which, in turn, may contribute to the overall decrease in the concentration of nucleic acids detected by PCR. Independent tests performed with purified nucleic acids revealed that 95% of the DNA strands were fragmented during sonication for 2×15 s using the FULM. The size range of the DNA fragments was estimated between 400 and 5,000 bp.
Imaging of disrupted spores
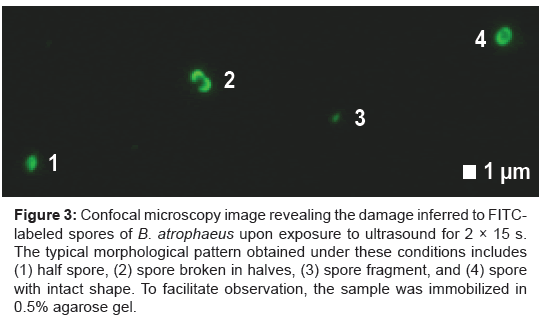
We performed confocal microscopy to visualize FITC-labeled spores in 3D and monitor possible changes in cell morphology as a result of the lysis process. Prior to sonication, non-lysed spores are elliptical in shape, measuring 1.2–1.8 μm in length and 0.9–1.0 μm in width as described previously [15]. In comparison, 21 fluorescent objects observed after sonication for 2×15 s were classified as such: 3 (14%) half spores, 4 (19%) spores broken in halves, 6 (29%) small spore fragments and 8 (38%) spores with intact shape. A section entailing an example of each specimen within relatively close proximity is shown in Figure 3.
Figure 3: Confocal microscopy image revealing the damage inferred to FITClabeled spores of B. atrophaeus upon exposure to ultrasound for 2 × 15 s. The typical morphological pattern obtained under these conditions includes (1) half spore, (2) spore broken in halves, (3) spore fragment, and (4) spore with intact shape. To facilitate observation, the sample was immobilized in 0.5% agarose gel.
Decontamination of the FULM
We tested decontamination of the FULM to simulate the transfer from a contaminated site to a secure area equipped with a biological safety cabinet containing the sonication platform and other detection equipment. We relied on immersion in a solution of sodium hypochlorite, which is known to be efficient against bacterial spores and hence is used for on-site decontamination [4]. Six immersion tests were performed to evaluate whether or not the interior of the FULM could be effectively protected from the disinfectant. The presence of sodium hypochlorite in the lysate, even at relatively low concentration, following penetration into the FULM is expected to interfere with subsequent DNA amplification and detection. Statistical analysis of PCR Ct revealed no significant difference between DNA extracts at each immersion time (Ct 5 min=30.45 ± 0.26, Ct 15 min=30.56 ± 0.23, Ct 60 min=30.36 ± 0.23), indicating that the interior of the FULM remained effectively sealed.
We further evaluated the efficiency of the decontamination procedure by testing viability of bacterial spores recovered from the surface of the FULM. Prior to immersion of the FULM in the hypochlorite bath, we recovered a significant amount of viable spores at the sample inlet (i.e., up to a maximum of ~300, and ~200 CFU on average). In contrast, the quantity of viable spores was negligible after decontamination had been completed: we found 1 CFU (max. 3 CFU) at the sample inlet as a statistical average deriving from four independent tests. After sonication for 2×15 s, an average of 0.5 CFU (max. 1 CFU) was recovered at the sample inlet, while no CFU was recovered from all other sites that had been swabbed. Agitation of the FULM during immersion seemed necessary to remove air bubbles being trapped in small-scale cavities (e.g., at the locking device or between connected units) which otherwise would prevent access of the disinfectant to these restrained sections of the module.
Dauphin and Bowen [19] have used filtration to remove viable spores of B. anthracis from lysates and preserve DNA extracts. We validated this possibility by collecting five different lysates from the FULM through Millex-GV filter units (0.22 μm pore size). Filtration indeed removed all viable spores from lysates as confirmed by microbial culture assays. Statistical analysis of PCR Ct performed with five replicates further revealed no significant difference between results for filtered (Ct=31.72 ± 0.31) and non-filtered (Ct=31.73 ± 0.44) lysates, suggesting no (or minimal) loss of DNA during filtration.
Simulated detection of Bacillus spores in suspect powders
We simulated a complete on-site Bacillus detection procedure using 3×105 B. atrophaeus spores and ~20 mg of bentonite. This powder has been reported as a potential additive in powdery bioweapons [20] while interfering with PCR detection of nucleic acids in lysates without sample preparation [17]. We previously developed a protocol termed dual-filter method for applied recovery of microbial particles from environmental and powdery samples (DARE) [17]. This sample preparation method allowed for rapid separation of bacteria from 23 different matrices using syringes and filter units. Using this method, bentonite resulted in the lowest spore recovery (i.e., 16%), hence we used it as a paradigm for detection simulation. Harvesting using swab and processing of the powder sample (<2 min) was performed twice using the DARE procedure. The cleaned spores deriving from this process were immediately injected into the FULM for spore lysis and DNA extraction (2×15 s). PCR analysis of the lysates clearly confirmed the presence of DNA from B. atrophaeus (data not shown), emphasizing the ability to recover cleaned microbial nucleic acids within 3 min.
Discussion
Performance of FULM for spore lysis and DNA extraction
Our ultrasonic lysis system provided DNA yield extracted from B. atrophaeus comparable to that of the BD GeneOhmTM Lysis Kit within a lysis time of 45 s (i.e., 2×15 s of sonication with 15-s interruption) (Figure 2). The BD GeneOhmTM Lysis Kit served as a benchmark process since it effectively promotes lysis (with 98.8% yield or more) of several microbial species, including B. anthracis and B. subtilis spores as determined by PCR amplification of genomic DNA in lysates [14,21]. Moreover, this kit has been shown to provide the highest yield of DNA from Mycobacterium tuberculosis with respect to five other relatively fast nucleic acid extraction methods [22]. Our ultrasonic system produced DNA extracts with concentrations comparable to those obtained with a validated commercial DNA extraction method in one fifth of the time. In addition, the yield is higher than that of a previous bead-based mechanical lysis performed on a chip under optimized conditions [15]. Furthermore, structural damage to sonicated spores (Figure 3) contrasts with our previous study that showed only subtle spore modifications after bead-based mechanical lysis using the same microscopic methodology [15]. These findings also support the capacity of the system to efficiently disrupt bacterial spores, making it possible to rapidly extract genomic DNA with high yield.
Effect of the stainless steel interface
We equipped the module presented herein with a thin, metallic interface as opposed to preceding sonication systems which primarily employed flexible, polymer-based membranes. Stainless steel provided a plausible choice for a number of reasons. Unlike carbon steel or brass which were both tested in the course of this study (data not shown), stainless steel does not corrode when coming in contact with aqueous buffer solutions prior to or during sonication. Chemical stability is important as metal ions or particles contaminating the lysate could interfere with PCR amplification and detection of extracted nucleic acids [23,24]. In addition, stainless steel can effectively promote propagation and transfer of acoustic energy. The energy flux E can generally be described as E=4π2f2ZA2T where f is the frequency, Z is the acoustic impedance, A is the amplitude of vibration, and T is the transmission coefficient of the interface. Due to its high acoustic impedance (e.g., Z=45.7×106 kg s-1 m-2 as a standard value) stainless steel can outperform a soft polymer compound such as polypropylene (e.g., Z=2.4×106 kg s-1 m-2) despite the fact that T is comparatively small for a stainless steel/water interface as a result of their impedance mismatch. Stainless steel further is preferable when considering the cost of other high-impedance materials such tungsten, iridium or gold. The lysis experiments performed in the course of this work suggest that a thickness of 0.001˝ (25.4 μm) constitutes one (though not the only) possible compromise in the trade-off between flexibility and robustness. When thin enough, the stainless steel interface can adapt to the shape of the sonicator probe which is important for effective transfer of the acoustic energy. On the other hand, an increased ability to conform renders the interface fragile which can constrain reliable and safe operation. For example, a stainless steel interface with a reduced thickness of 0.0005˝ (12.7 μm) seems impractical for it ruptures more easily, especially when performing lysis experiments in a sequential fashion. An increased thickness of 0.003˝ (76.2 μm), on the other hand, provides excellent stability during sonication, yet its higher rigidity hampers smooth adaptation of the interface to the sonicator probe. In our preliminary experiments, we evaluated a number of polymer-based interfaces and obtained decent DNA yields (data not shown). However, the yield improved noticeably when we exchanged the polymer by a stainless steel interface. Preceding work suggests the use of a polymer interface may need to be accompanied by the presence of microbeads in the suspension to enhance lysis of Bacillus spores and increase DNA yields [6,7,9,11]. We believe that the absence of beads is generally preferable for it helps maintain simplicity in the layout and assembly of microfluidic units.
DNA fragmentation
A certain level of DNA fragmentation can be useful for downstream molecular detection [18]. Using control experiments with purified nucleic acids, we estimated that 95% of DNA was fragmented during sonication. The size range of the DNA fragments was approximatly between 400 and 5,000 bp and compares well to those achieved by Warner et al. [11] (i.e., 100–600 bp) for subsequent PCR amplification and detection. By contrast, Warner et al. [11] evaluated DNA yields using a relativly short PCR amplicon (e.g., 66 bp). Short DNA fragments can limit the possibility to detect longer PCR amplicons. For example, reports on the PCR detection of B. anthracis rely on amplicon sizes between 91 and 596 bp [25-28]. We evaluated the yield of our DNA extracts with a 212-bp PCR amplicon, hence the DNA size range obtained by sonication using the FULM provides a relatively high flexibility for subsequent PCR assay selection.
Simulation of field testing
A number of requirements need to be taken into account when designing and evaluating an instrument for the detection of biothreat agents in the field. As emphasized by Lim et al. [3], an ideal detection platform should be able to detect biothreat agents from different environmental matrices (e.g., powder, liquid, or food) while providing portability, user-friendliness and resistance to unfavorable conditions in the field (e.g., shock, temperature, and decontamination, among others) [3]. We simulated the detection of B. atrophaeus spores from suspect powder under the constraints of a laboratory setting devided into a contaminated site and a secure area. The primary objectives were to evaluate the performance of our procedure when using powdery samples and identify any possible safety issues. Our findings suggest that the module can indeed be used to transfer a pathogenic sample for further processing in a biological safety cabinet respecting recommended packing and transportation protocols for hazardous materials, such as a triple-layer system [4]. Nevertheless, the instrumental setting may need further improvement to facilitate handling. For example, it should be desirable to streamline the exterior design of the FULM and reduce the number of sites where air bubbles can be trapped to facilitate its decontamination. Modification of the docking mechanism is likely to improve robustness of the system and prevent interface fracture. Likewise, automation of sample injection, adjustment of the sonication probe to the interface and recovery of the lysate would shorten process times, and increase user-friendliness and safety. Finally, miniaturization of the sonication platform would facilitate its integration into a portable total-analysis system.
We further used the prototype instrument in complementation to a recently developed protocol for fast (<2 min) separation of bacterial spores from suspicious powder samples using two filtration steps (DARE) [17]. Combined, these methodologies allowed for detecting B. atrophaeus processed with 20 mg of bentonite as a potential aerosolizing additive [20]. This important aspect had not been covered previously for the development of a sonication platform to detect B. anthacis spores [6,7,9-11]. Our analyte preparation from suspect powders (DARE and FULM) could potentially facilitate on-site detection of B. anthracis and other pathogenic species. While a combination of the two methods is in itself an appealing possibility, integration in a single biothreat detection device would reveal their full potential to facilitate sample preparation in a field setting.
Conclusions
Time, efficiency and safety are crucial parameters while testing for infectious agents, especially in the event of a bioterrorist attack. The concept presented in this work is responding to this challenge. First, an ultrasonic lysis modular system allows for rapid (<1 min) extraction of nucleic acids from bacterial spores, while providing yields that are equivalent to those obtained using a commercial DNA extraction method over a 5-min period. This performance is remarkable and qualifies the tool as one of the fastest and most efficient spore lysis systems currently available. The implementation of a stainless steel interface was a key factor in this context, marking a distinction in design with respect to other sonication systems that typically rely on a polymer membrane as the interface. Second, the need for safe portability was equally included in the design of the tool described herein. The interior of the module can be sealed, thus preventing aerosol production and propagation, while the exterior surface can be decontaminated without inducing any adverse effect on the tightness of the FULM and the integrity of the collected sample inside. The FULM, less expensive and more resistant, could be used in the field and be decontaminated, while the sonication platform, containing more expensive and sensitive electromechanical parts, could stay in a secure area.
Acknowledgement
This work was supported in part by Defence Research and Development Canada, Centre for Security Science, Chemical, Biological, Radiological/ Nuclear, and Explosives Research and Technology Initiative (CRTI) under the program Portable Biological Agent Detection System (06-0187TD). Fluorescence micrographs were acquired through the Bioimaging facility at CRI. We thank our colleagues Jean-Guy Allard (NRC), Liviu Clime (NRC), Karel Boissinot (CRI), Sébastien Chapdelaine (CRI), Isabelle Charlebois (CRI), Gale Stewart (CRI), Julie- Christine Lévesque (CRI) and Manon Tétreault for useful discussion and technical assistance. Ian Summerell and the first responder team of the Royal Canadian Mounted Police (Orleans, ON) are acknowledged for conceptual input to this work.
References
- Rotz LD, Khan AS, Lillibridge SR, Ostroff SM, Hughes JM (2002) Public health assessment of potential biological terrorism agents. Emerg Infect Dis 8: 225-230.
- Inglesby TV, Henderson DA, Bartlett JG, Ascher MS, Eitzen E, et al. (1999) Anthrax as a biological weapon: medical and public health management. Working Group on Civilian Biodefense. JAMA 281: 1735-1745.
- Lim DV, Simpson JM, Kearns EA, Kramer MF (2005) Current and developing technologies for monitoring agents of bioterrorism and biowarfare. Clin Microbiol Rev 18: 583-607.
- Emanuel P, Roos JW, Niyogi K (2008) Sampling for biological agents in the environment. ASM Press, Washington DC.
- Wade MM, Campbell MS, Niyogi K, Rogers JE, Coughlin J, et al. (2006) Evaluation of an inexpensive field test for ruling out the presence of biological threat agents in suspicious powders. ECBC-TR-459.
- Belgrader P, Hansford D, Kovacs GT, Venkateswaran K, Mariella R Jr, et al. (1999) A minisonicator to rapidly disrupt bacterial spores for DNA analysis. Anal Chem 71: 4232-4236.
- Belgrader P, Okuzumi M, Pourahmadi F, Borkholder DA, Northrup MA (2000) A microfluidic cartridge to prepare spores for PCR analysis. Biosens Bioelectron 14: 849-852.
- Chandler DP, Brown J, Bruckner-Lea CJ, Olson L, Posakony GJ, et al. (2001) Continuous spore disruption using radially focused, high-frequency ultrasound. Anal Chem 73: 3784-3789.
- Taylor MT, Belgrader P, Furman BJ, Pourahmadi F, Kovacs GT, et al. (2001) Lysing bacterial spores by sonication through a flexible interface in a microfluidic system. Anal Chem 73: 492-496.
- Marentis TC, Kusler B, Yaralioglu GG, Liu S, Haeggström EO, et al. (2005) Microfluidic sonicator for real-time disruption of eukaryotic cells and bacterial spores for DNA analysis. Ultrasound Med Biol 31: 1265-1277.
- Warner CL, Bruckner-Lea CJ, Grate JW, Straub T, Posakony GJ, et al. (2009) A flow-through ultrasonic lysis module for the disruption of bacterial spores. J Lab Autom 14: 277-284.
- Rose HL, Dewey CA, Ely MS, Willoughby SL, Parsons TM, et al. (2011) Comparison of eight methods for the extraction of Bacillus atrophaeus spore DNA from eleven common interferents and a common swab. PLoS One 6: e22668.
- Carrera M, Zandomeni RO, Fitzgibbon J, Sagripanti JL (2007) Difference between the spore sizes of Bacillus anthracis and other Bacillus species. J Appl Microbiol 102: 303-312.
- Picard FJ, Gagnon M, Bernier MR, Parham NJ, Bastien M, et al. (2009) Internal control for nucleic acid testing based on the use of purified Bacillus atrophaeus subsp. globigii spores. J Clin Microbiol 47: 751-757.
- Geissler M, Beauregard JA, Charlebois I, Isabel S, Normandin F, et al. (2011) Extraction of nucleic acids from bacterial spores using bead-based mechanical lysis on a plastic chip. Eng Life Sci 11: 174-181.
- Package insert of BD GeneOhmTM Lysis Kit. BD Diagnostics, Quebec QC (2009) DOPS04-07-V1M2.
- Isabel S, Boissinot M, Charlebois I, Fauvel CM, Shi LE, et al. (2012) Rapid filtration separation-based sample preparation method for Bacillus spores in powdery and environmental matrices. Appl Environ Microbiol 78: 1505-1512.
- Larguinho M, Santos HM, Doria G, Scholz H, Baptista PV, et al. (2010) Development of a fast and efficient ultrasonic-based strategy for DNA fragmentation. Talanta 81: 881-886.
- Dauphin LA, Bowen MD (2009) A simple method for the rapid removal of Bacillus anthracis spores from DNA preparations. J Microbiol Methods 76: 212-214.
- Shoham D, Jacobsen SM (2007) Technical intelligence in retrospect: The 2001 anthrax letters powder. International Journal of Intelligence and CounterIntelligence 20: 79-105.
- Ménard C, Picard FJ (2003) Universal method for the rapid lysis of cells for the release of nucleic acids and their detection. WO/2003/008636.
- Aldous WK, Pounder JI, Cloud JL, Woods GL (2005) Comparison of six methods of extracting Mycobacterium tuberculosis DNA from processed sputum for testing by quantitative real-time PCR. J Clin Microbiol 43: 2471-2473.
- Primorac D (2004) The role of DNA technology in identification of skeletal remains discovered in mass graves. Forensic Sci Int 146: S63-S64.
- Alaeddini R (2012) Forensic implications of PCR inhibition--A review. Forensic Sci Int Genet 6: 297-305.
- Ellerbrok H, Nattermann H, Ozel M, Beutin L, Appel B, et al. (2002) Rapid and sensitive identification of pathogenic and apathogenic Bacillus anthracis by real-time PCR. FEMS Microbiol Lett 214: 51-59.
- Ryu C, Lee K, Yoo C, Seong WK, Oh HB (2003) Sensitive and rapid quantitative detection of anthrax spores isolated from soil samples by real-time PCR. Microbiol Immunol 47: 693-699.
- Kim K, Seo J, Wheeler K, Park C, Kim D, et al. (2005) Rapid genotypic detection of Bacillus anthracis and the Bacillus cereus group by multiplex real-time PCR melting curve analysis. FEMS Immunol Med Microbiol 43: 301-310.
- Yang Y, Wang J, Wen H, Liu H (2012) Comparison of two suspension arrays for simultaneous detection of five biothreat bacterial in powder samples. J Biomed Biotechnol 2012: 831052.
Relevant Topics
- Anthrax Bioterrorism
- Bio surveilliance
- Biodefense
- Biohazards
- Biological Preparedness
- Biological Warfare
- Biological weapons
- Biorisk
- Bioterrorism
- Bioterrorism Agents
- Biothreat Agents
- Disease surveillance
- Emerging infectious disease
- Epidemiology of Breast Cancer
- Information Security
- Mass Prophylaxis
- Nuclear Terrorism
- Probabilistic risk assessment
- United States biological defense program
- Vaccines
Recommended Journals
Article Tools
Article Usage
- Total views: 15880
- [From(publication date):
November-2012 - Dec 20, 2025] - Breakdown by view type
- HTML page views : 11084
- PDF downloads : 4796