Research Article Open Access
Analysis of Enzymes Activities on Domestic Waste Dump Sites
Callistus I Iheme*, Doris I Ukairo, Chiedozie O Ibegbulem, Olivia O Okorom and Kelechi ChibunduDepartment of Biochemistry, Federal University of Technology, Owerri, Nigeria
- *Corresponding Author:
- Callistus I Iheme
Department of Biochemistry
Federal University of Technology, Owerri
Nigeria
Tel: +2347031014133
E-mail: rabikally@gmail.com
Received date: June 15, 2017; Accepted date: June 26, 2017; Published date: June 30, 2017
Citation: IIheme C, Ukairo DI, Ibegbulem CO, Okorom OO, Chibundu K (2017) Analysis of Enzymes Activities on Domestic Waste Dump Sites. J Bioremediat Biodegrad 8: 400. doi: 10.4172/2155-6199.1000400
Copyright: © 2017 Iheme C, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
Introduction: Effects of physico-chemical parameters on microbial dehydrogenases from domestic waste dumpsites were studied.
Methodology: The microorganisms (S. aureus, E. coli, P. aeruginosa, C. albicans, T. Mentagrophytes and F. oxysoprium) were isolated from three domestic waste dumps in Owerri metropolis and subculture. Microbial dehydrogenases were extracted with acetone, partially purified by ammonium sulphate precipitation, dialysis, DEAEcellulose column chromatography and Sephadex G200 gel filtration chromatography. Dehydrogenase activity at each stage of purification was assayed with 2,3,5-triphenyltetrazolium chloride (TTC) and the absorbance of the formazan formed was measured and used to ascertain the total dehydrogenase activity. The impacts of effectors (Ca2+, Mg2+, Zn2+, Fe2+ and EDTA, ethanol and butanol) on the microbial dehydrogenase were determined by incubating the partially purified enzyme with the effectors for 30 min at 4°C. Also, the effects of temperature and pH were assessed by varying the temperature and pH ranges from 10°C to 60°C and 2.0 to 8.0, respectively.
Results: The specific activities of the enzyme from the microorganisms were 7.10, 7.73, 6.47, 6.26, 9.66 and 10.58 mg Formazan/mg cell dry wt/h, respectively. Calcium ion, Mg2+, ethanol and butanol significantly increased (p<0.05) dehydrogenase activities in all the microorganisms studied while Zn2+, Fe2+ and EDTA decreased the activities.
Conclusion: These activators can be harnessed within the recorded optimum temperature and pH ranges to enhance microbial growth; which is essential for the degradation of domestic waste thereby promoting cleaner environment.
Keywords
Dehydrogenase activities; Effectors; Domestic waste
Introduction
The quality of life on the Earth is linked inextricably to the overall quality of the environment. Wastes were traditionally disposed of in landfills in the past. This traditional mode of waste disposal was publically unacceptable due to the increasing conversion of scarce agricultural lands to dump sites. Over time, new technologies for waste disposal that use high-temperature incineration and chemical decomposition (e.g., base-catalyzed de-chlorination, UV oxidation) evolved. These technologies have been very effective at degrading a wide range of contaminants; yet, they have several setbacks. They are complex, expensive, and also lack public acceptance. The associated deficiencies in these methods have resulted to a paradigm shift towards harnessing bioremediation as a better alternative.
Literature Review
The process of bioremediation mainly relies on microorganisms which enzymatically utilize the pollutants as substrates and convert them to nonhazardous or less-hazardous products [1]. Bioremediation can be effective only where environmental conditions permit microbial growth and activity. Its application often involves the manipulation of environmental parameters to allow microbial growth and degradation to proceed at a faster rate by providing convenient environmental conditions [1].
The process of bioremediation is very slow and only few species of bacteria and fungi have proven abilities to degrade pollutants. Many strains are known to be effective as bioremediation agents but only under laboratory conditions [2]. The limitation of bacterial growth is under the influence of pH, temperature, oxygen, soil structure, moisture and appropriate level of nutrients, poor bioavailability of contaminants, and presence of other toxic compounds. Both bacteria and fungi rely on the participation of different intracellular and extracellular enzymes for the degradation of recalcitrant lignin and organo-pollutants [2]. Dehydrogenase is an important intracellular microbial enzyme that is tightly linked to microbial oxido-reduction process and as such, serves as a potent bioindicator for the overall soil microbial activities [3]. Microbial dehydrogenase (EC 1.1.1.) is an intracellular enzyme that occurs in all viable microbial cells. This enzyme functions as a measurement of the metabolic state of soil microorganisms [4]. Dehydrogenase activity (DHA) is one of the most adequate, important and most sensitive bioindicators, relating to soil fertility [3]. Its activity depends on the same factors which influence microbial abundance and activities [4]. Dehydrogenases play significant roles in the biological oxidation of soil organic matter (OM) by transferring hydrogen from organic substrates to inorganic acceptors [5]. Many specific dehydrogenases transfer electrons from the substrate to an electron carrier; what carrier is used depends on the reaction taking place. Common electron acceptors used by this enzyme are NAD+, FAD, and NADP+. These electron carriers are reduced in this process and considered oxidizers of the substrate. Electron carriers are coenzymes that are often referred to as "redox cofactors [3,6]. DHA serves as an indicator of the microbiological redox-system and can be considered a sensitive and adequate measure of microbial oxidative activity in soil. It has been reported that active dehydrogenases can utilize both O2 and other compounds as terminal electron acceptors, although anaerobic microorganisms produce most dehydrogenases [7]. Therefore, DHA reflects metabolic ability of the organism and its activity is considered to be proportional to the biomass of the microorganisms in the environment [8].
Temperature and pH affect the activities of microbes, but some organisms thrive in relatively high and relatively low values of temperature and pH which invariably affect the activities of most extracellular enzymes like the dehydrogenases [9]. Fungi can grow in relatively high temperature ranges of 42°C-45°C; this temperature range denature and hence affect enzyme activities. Most enzymes are sensitive to pH, have specific pH range and optimum pH of activity. The pH can stop enzyme activity by denaturating (altering) the three-dimensional shape of the enzyme, by breaking ionic and hydrogen bonds; with most enzymes functioning between pH of 6 and 8 [9].
In our modern society, environmental pollution is a serious threat that impedes adequate utilization of arable lands for agricultural and economic purposes. The activities of microorganisms that promote plant growth can be altered by high concentrations of certain pollutants. Heavy metal pollution, for instance, not only cause adverse effects on parameters relating to plant quality and yield but also causes changes in the size, composition and activity of the microbial community such as organic matter mineralization [10]. Some metal ions increase the activities of certain microbes both in small and high concentrations. Metalloproteins and metal ion-activators like K+, Fe2+, Fe3+, Cu2+, Co2+, Zn2+, Mn2+, Mg2+, Ca2+, and Mo3+ [11], bind to certain enzymes for effective catalysis. Ethylene Diamine Tetra Acetic Acid (EDTA) has been known as a chelating agent, reported to inhibit a range of metallopeptidases, and the method of inhibition is believed to occur through the chelation of the metal ions required for catalytic activities.
This study is aimed at determining the optimum conditions for maximum activities of dehydrogenases from different microbial sources in domestic waste dump and also to expose them to selected effectors.
Materials and Methods
Microbial isolation
Soil samples were collected from three domestic waste dumpsites using sterile containers. The soil samples were mixed, cultured in potato dextrose agar and nutrient agar for fungal and bacterial growth respectively. Growths were recorded in each growth medium. Microorganisms of interest were identified by their unique characteristic growth pattern, shape and colour. The isolated S. aureus, E. coli, P. aeruginosa, and C. albicans, T. Mentagrophytes, F. oxysoprium were sub-cultured in nutrient and potato dextrose agars respectively. The bacterial strains were grown to mild exponential phase (for 24 h) at 30°C under static conditions.
Screening for dehydrogenase activity
On a colony of each bacterial and fungal isolates growing on nutrient and potato dextrose agars respectively, 1 drop of 1:1 mixture of aqueous solution of 0.4% w/v 2,3,5-triphenyl tetrazolium chloride (TTC) and glucose (2% w/v) were added. The plates were incubated at room temperature for 10 min. Production of red colored formazan indicated the presence of dehydrogenase.
Methods of enzyme isolation
Cells were harvested by centrifugation at 4000 rpm for 10 min, washed three times with 50 mm potassium phosphate buffer (pH 7.5), and suspended in the same buffer containing 2 mm EDTA and 1 mm Dithiothreitol (DTT). Cells were ruptured using osmotic shock by Nossal and Heppel [12]. The cells were suspended in 20% sucrose buffer then separated by centrifugation at 4000 rpm for 10 min. The resulting paste was dispersed in acetone at 4°C. Cellular debris and unbroken cells were removed by centrifugation at 4000 rpm for 45 min at 4°C. The supernatant obtained constituted the crude microbial extracts (soluble enzyme fraction) for each microbe.
Purification of enzyme
The enzyme was partially purified from the crude microbial extracts in four steps: ammonium sulfate precipitation, dialysis, sephadex G200 gel filtration chromatography and DEAE-cellulose column chromatography. All the steps were performed at 4°C.
Ammonium sulphate precipitation
The protein sample was allowed to thaw to determine total volume, and was centrifuged at 3000 rpm for 30 min. This was transferred into a beaker containing a stir bar and was placed on a magnetic stirrer. While the sample was being stirred, solid ammonium sulfate crystals were added to bring the final concentration to 60% saturation. (The volume of ammonium sulphate used was equal to the volume of the sample. Adding the ammonium sulphate very slowly ensured that local concentration around the site of addition did not exceed the desired salt concentration). Once total volume of ammonium sulphate was added, the beaker was kept at 4°C for 6 hours. The solution was transferred to centrifuge tube and the precipitate centrifuged at 10,000 rpm for 10 min. The supernatant was carefully concentrated to 80% using ammonium sulphate, and was then stirred. After 60 min, it was centrifuged at 12,000 rpm. The supernatant was discarded and the pellet or precipitate collected and re-suspended in the minimal volume of 0.002 mM phosphate buffer pH 7.2. The DHA at this level was checked by TTC reduction. The protein was further purified using dialysis.
Dialysis
The membrane was placed in 1% sucrose solution for 30 min and the cell free crude extract that was re-suspended in minimum volume of 0.02 M phosphate buffer solution (pH 7.2) was introduced into the dialysis bag. The solution was dialyzed against 500 ml of the same buffer for 12 hours at 4°C in a refrigerator to remove the excess salt with one change after every 4 hours. Thus, an enzyme preparation concentrated against sucrose was obtained. This was followed by centrifugation of the resulting solution at 12,000 rpm at 4°C and the supernatant was tested for DHA.
Sephadex G-200 method
This technique was used to further purify the enzyme extract. This was carried out using the method of Hasan et al. [13]. The enzyme extract was equilibrated with 0.01 M sodium phosphate buffer (pH 6.5) and the slurry was allowed to swell overnight at room temperature. Sodium azide (0.02%) was added to prevent microbial growth. This was then loaded into a 1.6 × 40 cm column, equilibrated with 0.25 M sodium phosphate buffer and fractionated through the Sephadex G-200 column. Five milliliters (5 ml) of the enzyme preparation was applied carefully to the top of the gel and allowed to elute through the gel in the column. The buffer was added without disturbing the gel surface and the reservoir. Elution was carried out with the buffer at a flow rate of 20 ml/h. Fifty (50) fractions (5 ml each) were collected and absorbance was read at 280 nm using spectrophotometer. Protein and DHA in the fractions were estimated. The eluted active enzyme fractions were pooled and used further purification.
Ion-exchange chromatography
The enzyme was further purified using DEAE-cellulose column chromatography. The method used was as described by Yannis [14]. DEAE-cellulose was suspended in 8 vol. of Tris-buffer containing 50 mM of NaCl and kept overnight for equilibration. The column was carefully packed equilibrated with 8 vol. of same buffer containing 0.25 mM of NaCl. Then 3 ml of the partially purified enzyme was diluted to 15 ml and loaded into the column and was washed with appropriate 100 ml of the equilibration buffer. The protein was eluted with 0.25M buffer pH 7.2 and NaCl gradient 0.1-1 M, was passed through the exchanger at a flow rate of 30 ml/hr. Then 3 ml fractions were collected. Protein content was traced in accordance to absorbency at 280 nm. The elute was collected as a single fraction and tested for DHG activity.
Calculation
The % yield, purification fold and specific activities were calculated using the following formulae:
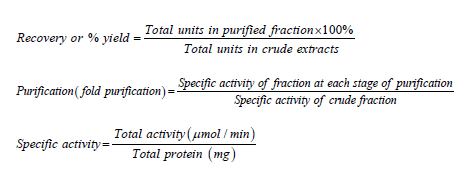
Total protein=Protein concentration × volume of extract.
Impacts of effectors
The impacts of effectors, EDTA, ethanol and butanol on the activities of the purified dehydrogenases were determined. The enzyme was incubated with the effectors for 30 min at 4°C and the activities were estimated with spectrophotometer at 340 nm using TTC as substrate. The percentage effect was calculated thus:
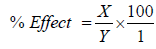
X=Enzyme activity without effector-Enzyme activity with effector; Y=Enzyme activity without effector.
Effects of metal ions
The effects of various metal ions like Zn2+, Ca2+, Fe2+, and Mg2+, on the activity of the DHG were studied. The enzyme was incubated with 5 mM of each metal ion for 30 min at 4°C and the activities were measured with the aid of a spectrophotometer at 340 nm using TTC as the substrate. Percentage effect of metal ion was calculated thus:
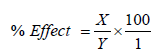
X=Enzyme activity without effector-Enzyme activity with effector; Y=Enzyme activity without effector.
Effect of pH
Enzyme activity was determined at varying pH values of the reaction medium, ranging from pH 2.0-8.0 at 34°C.
Effect of temperature
Enzyme activity was determined at a temperature range of 10°C-60°C.
Statistical Analysis
Data obtained were analyzed using one-way Analysis of Variance (ANOVA). Values were considered significantly different at p<0.05.
Discussion
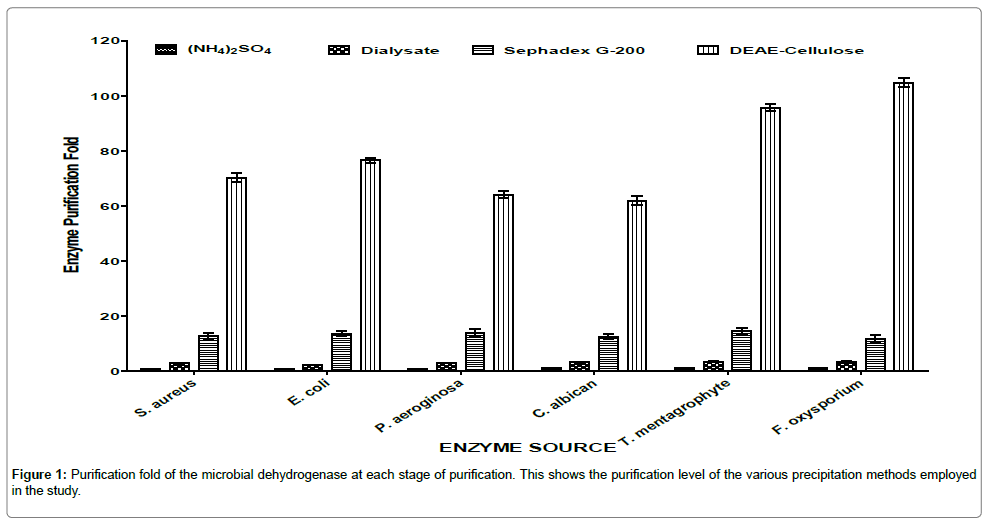
The purification procedures applied in the study present convenient ways to purify dehydrogenase (DHG) enzymes from the various microorganisms. Figure 1 summarized the degree of purification achieved on dehydrogenases from the six microorganisms. At every level of purification there was increase in purification fold.
The highest purification fold and specific activities (Figure 2) of the dehydrogenases in the corresponding microorganisms were recorded at the DEAE-cellulose column purification. The increase in specific activity at each stage of purification hence confirmed the report of Iheme [15] which said that specific activity of the successive fractions containing the enzyme of interest should be greater than the preceding fractions until complete purification is achieved.
The dehydrogenase activity (DHA) varied among the bacterial strains. The Gram-positive Staphylococcus sp. had low dehydrogenase activity than the Gram-negative Pseudomonas sp. This was in agreement with the report of Nweke et al. [16] but at variance with the observations of Nwaogu et al. [17] and Nwaogu et al. [18] in which the Gram-positive organisms had higher DHA than the Gram-negative organisms. These variations may be due to differences in bacterial physiology, including cell wall components or DHG systems, since different microorganisms have been reported to have different dehydrogenase systems [19]. From the study, fungi strains (T. mentagrophyte and F. oxysporium) had the highest DHA.
Many metal ions alter the activities of enzymes by combining with them in a way that influences the binding of the substrates. These metal ions can be called effectors either being activators or inhibitors [20]. The effects of metal ions (like Ca+, Mg2+, Fe2+ and Zn2+) on the DHA activities were shown in Tables 1 and 2. According to the results, Ca2+ increased the specific activities of the partially purified DHG from all the studied microorganisms. Table 3 showed the percentage activation and inhibition of the metal ions and the inhibitors. Calcium ion (Ca2+) generally activated DHA in all the studied organisms. Calcium ion (Ca2+) is an integral part of DHG (metalloprotein) and its activating effect has been reported by Prakash et al. [20]. Magnesium ion caused increase in the percentage activities in all the studied organisms but for E. coli, P. aeroginosa and T. mentagrophytes. This is at variance with the report of Prakash et al. [20]. Iron and zinc ions on the other hand decreased the enzyme activities in all the studied microorganisms. This result was also at variance with the report of Sagar and Krutika [21], but corroborated with the findings of Sonia and Saksham [10].
| Microorganism | Specific activity (μmol/min/mg) | |||
|---|---|---|---|---|
| (NH4)2SO4 | Dialysate | Sephadex G-200 | DEAE-cellulose | |
| S. aureus | 0.09 ± 0.01 a | 0.29 ± 0.05b | 1.28 ± 0.12 c | 7.10 ± 0.09 d |
| E. coli | 0.09 ± 0.02 a | 0.23 ± 0.05 b | 1.38 ± 0.10 c | 7.73 ± 0.10 d |
| P. aeroginosa | 0.10 ± 0.04 a | 0.31 ± 0.08 b | 1.41 ± 0.13 c | 6.47 ± 0.13 d |
| C. albican | 0.11 ± 0.03 a | 0.33 ± 0.09 b | 1.27 ± 0.08 c | 6.26 ± 0.11 d |
| T. mentagrophyte | 0.12 ± 0.03 a | 0.35 ± 0.07 b | 1.48 ± 0.12 c | 9.66 ± 0.13 d |
| F. oxysporium | 0.12 ± 0.04 a | 0.36 ± 0.10 b | 1.19 ± 0.12 c | 10.58 ± 0.16 d |
Table 1: Specific activity of partially purified microbial dehydrogenase.
| Enzyme Source | Specific activity | Microbial dehydrogenase activity (U/L) | ||||||
|---|---|---|---|---|---|---|---|---|
| CaCl2 | FeSO4 | ZnCl2 | MgCl2 | EDTA | Ethanol | Butanol | ||
| S. aureus | 4.96 ± 0.10 h | 8.93 ± 0.21a | 2.99 ± 0.09b | 2.49 ± 0.09c | 5.94 ± 0.10d | 1.23 ± 0.05e | 9.41 ± 0.15f | 8.34 ± 0.13g |
| E. coli | 6.58 ± 0.14 a | 6.74 ± 0.12 a | 2.57 ± 0.11b | 2.33 ± 0.13b | 4.78 ± 0.17c | 1.16 ± 0.06d | 7.93 ± 0.18e | 7.04 ± 0.15f |
| P. aeroginosa | 4.20 ± 0.10 c | 5.99 ± 0.10 a | 2.01 ± 0.09b | 1.90 ± 0.10b | 4.03 ± 0.14c | 0.96 ± 0.07d | 7.32 ± 0.13e | 6.86 ± 0.18 |
| C. albican | 4.47 ± 0.09 h | 8.31 ± 0.14 a | 3.07 ± 0.10b | 3.03 ± 0.14b | 6.11 ± 0.15c | 1.53 ± 0.09d | 9.22 ± 0.17 e | 7.55 ± 0.14f |
| T. mentagrophyte | 7.31 ± 0.15 h | 9.41 ± 0.17 a | 3.86 ± 0.12b | 3.58 ± 0.14b | 6.46 ± 0.15c | 1.84 ± 0.10d | 10.39 ± 0.13e | 8.04 ± 0.13f |
| F. oxysporium | 5.56 ± 0.10 h | 10.34 ± 0.15a | 3.94 ± 0.17b | 3.71 ± 0.15b | 7.92 ± 0.12c | 1.93 ± 0.10d | 11.34 ± 0.18e | 9.23 ± 0.15f |
Table 2: Effect of metal ions and inhibitors on microbial dehydrogenase activities.
The enzyme activity decreased on treatment with EDTA, whereas butanol and ethanol increased the activities. It is well known that some enzymes contain Ca2+ as their essential component and are often inhibited by chelating reagent like EDTA; presumably because of its chelating properties [20]. These results thus showed that DHG is among the various enzymes that are not resistant to this chelating agent. A similar trend with respect to concentration-dependence was observed in case of EDTA on amylase. Kiran and Chandra [22] and Femi-Ola and Olowe [23] reported that EDTA inhibited amylase activity; suggesting that EDTA acted by chelating Ca2+.
Dehydrogenase was also found to be sensitive to butanol and ethanol. A significant stimulation on the enzyme specific activity was observed with ethanol. The result showed that ethanol may have been utilized by the enzyme as substrate and; ethanol at low concentration had been shown to promote the growth of microorganisms. This finding was in line with the report of Seitz et al. [24] who reported that ethanol activated human gastric dehydrogenase. Butanol on the other hand, increased DHA activities as recorded in all the studied organisms. The percentage activations of DHA activities by ethanol and butanol as shown in Table 3 were higher in F. oxysporium, C. albican, S. aureus, and C. albican respectively, when compared to the activities in other studied microorganisms.
| Enzyme Source | Effects (%) | ||||||
|---|---|---|---|---|---|---|---|
| CaCl2 | FeSO4 | ZnCl2 | MgCl2 | EDTA | Ethanol | Butanol | |
| S. aureus | 80% | -40% | -50% | 20% | -75% | 90% | 68% |
| E. coli | 2% | -61% | -65% | -27% | -82% | 21% | 7% |
| P. aeroginosa | 43% | -52% | -55% | -4% | -77% | 74% | 63% |
| C. albican | 86% | -31% | -32% | 37% | -66% | 106% | 69% |
| T. mentagrophyte | 29% | -47% | -51% | -12% | -75% | 42% | 10% |
| F. oxysporium | 86% | -29% | -33% | 42% | -65% | 104% | 66% |
Table 3: Percentage effects of metal ions and inhibitors on microbial dehydrogenase activities.
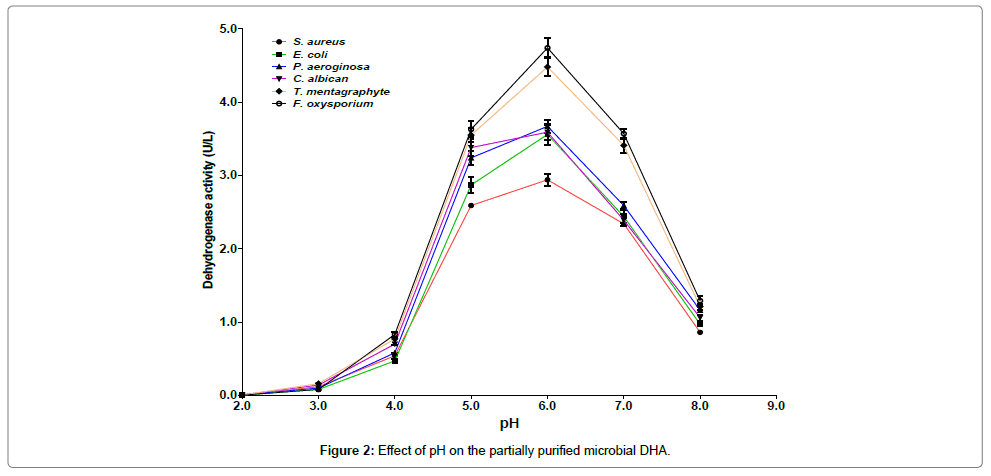
As shown in Figure 2, the maximum DHA activity was recorded for F. oxysporium at pH 6.0 and the lowest was observed in S. aureus at pH 2.0. At pH 6.0, 7.0 and 8.0 the DHA activity from S. aureus was lower than those from P. aeruginosa, T. mentagraphytes and F. oxysporium.
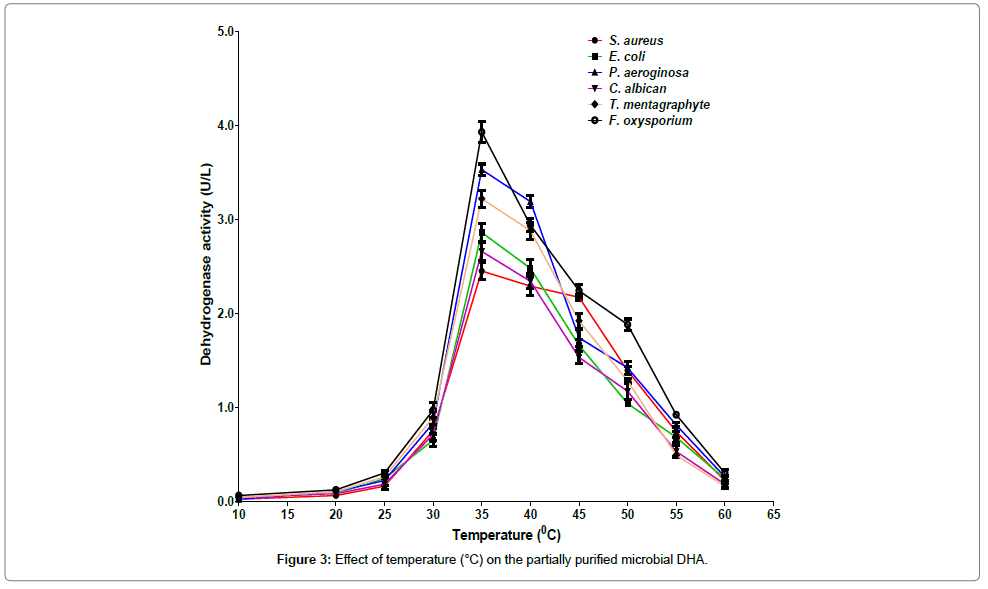
The maximum activities were recorded at 35°C while minimum activities were recorded at the temperature of 10°C (Figure 3). Among the organisms studied, the highest DHA activity was recorded for F. oxysporium and the lowest activity was observed for P. aeruginosa at the optimum temperature of 35°C. At 35°C, the DHA activities from S. aureus, E. coli and C. albicans were lower than those from P. aeruginosa, T. mentagraphytes, and F. oxysporium. The result obtained in this study is in contrast with that of Dickinson and Monger [25] who reported that the optimum temperature and pH for microbial DHA were 25°C and pH 7.5, respectively.
Conclusion
The effects of metal ions and inhibitors on the partially purified dehydrogenases from microorganisms were assessed. The optimum pH and temperature were also determined at 6.0 and 35°C, respectively. The activity was severely inhibited by EDTA but butanol and ethanol had marked activating effects. There was increase in activity of the enzyme on Ca2+ treatment, whereas Mg2+ had a moderate increase but activities were decreased by both Zn2+ and Fe2+. Hence, in the presence of these activators, optimum temperature and pH, the activity of microbial dehydrogenase can be enhanced for degradation of domestic wastes dump sites and by extension, remediation of the soil for industrial and agricultural purposes.
Declaration of Interest
The authors hereby declare that no conflict of interest exists.
References
- Vidali M (2001) Bioremediation: An overview. Pure and Applied Chemistry 73: 1163-1172.
- Hammel E, Cadisch G, Giller K (1997) Fungal degradation of waste.In: Driven by Nature Litter Quality and Decomposition,pp: 33-45.
- Subhani A, Changyong H, Zhengmiao Y, Min L,El-ghamry A (2001) Impact of Soil Environment and Agronomic Practices on Microbial/Dehydrogenase Enzyme Activity in Soil: A Review. Pakistan Journal of Biological Sciences 4: 333-338.
- Wolinska J, Stepniewska B (2012) Oxidoreductases extracellularly secreted by microbes. Journal of Molecular Biology 8:38-58.
- Zhang SB, Wu ZL (2010) Identification of amino acid residues responsible for increased pH of microbial dehydrogenase. Journal of Bioresources and Technology 102:2093-2096.
- Voet S, Donald F (2006) Fundamental of biochemistry life at the molecular level. John Wiley and Sons, New York, USA, pp: 105-114.
- Brzezi�?ska M, St�?pniewski W, St�?pniewska Z, Przywara G (2001) Effect of Oxygen Deficiency on Soil Dehydrogenase Activity in a Pot Experiment with Triticale CV Jago Vegetation. International Agrophysics 15: 145-149.
- Salazar S, Sanchez L, Alvarez J, Valverde A, Galindo P, et al. (2011) Correlation Among Soil Enzyme Activities Under Different Forest System Management Practices. Ecological Engineering 37: 1123-1131.
- Watts C, Sun L, Mochly R (2010) Mitochondrial aldehyde dehydrogenase. Biochemistry Journal 45: 227.
- Sonia S, Saksham G (2014) Heavy metal impact on soil microbial biomass, soil dehydrogenase activity and soil respiration rate. IntJAdv Res Biol Sci 1:29-34.
- Pfeiffer J (1954) Enzymes, the physics and chemistry of life. Wiley, New York, pp: 171-173.
- Nancy GN, Leon AH (1966) The Release of Enzymes by Osmotic Shock from Escherichia coli in Exponential Phase. J Biol Chem 241:3055-3062.
- Hassan C, Fredlier A, Neri H, Reed R (2013) Sephadex G200. Method of enzyme purification to increase specificity, pp: 98-105.
- Yannis S (2014) Purification and Characterization of lactate dehydrogenase, An undergraduate biochemistry laboratory experiment. Advances in Biochemistry 2:14-23.
- Iheme CI (2013) Enzyme isolation, purification and characterization. In: Laboratory Experiments in Biochemistry (A Students’ Manual).In: Ojiako AO (ed.), FUTO Press Ltd., Owerri, Nigeria, pp: 300-333.
- Nweke CO, Okolo JC, Nwanyanwu CE, Alisi CS (2006) Response of planktonic bacteria of New Calabar River to zinc stress. Afr J Biotechnol. 5: 653-658.
- Nwogu LA, Alisi CS, Ibegbulem CO, Igwe CU (2007) Phytochemical and antimicrobial activity of ethanolic extract of Landolphia owariensis leaf. Afr J Biotechnol 6: 890-893.
- Nwogu LA, Alisi CS, Igwe CU, Ujowundu CO (2008) A comparative study of the antimicrobial properties of the ethanolic extracts of Landolphiaowariensis leaf and root. Afr J Biotechnol 7: 368-372.
- Praveen K,Tarafdar JC (2003) 2,3,5-Triphenyltetrazolium chloride (TTC) and Electron Acceptor of Culturable Cell Bacteria, Fungi and Antinomycetes. Biol Fert Soil 38: 186-189.
- Prakash O, Nivedita J, Rajesh K (2011) Effect of metal ions, EDTA and Sulfhydryl Reagents on soyebean amylase activity. Asian Journal of Biochemistry6:282-290.
- Sagar T, Krutika M (2015) Purification and characterization of allinase produced by curiavidus nectar and its application for generation of cytotoxic agent. Saudi Journal of Biological Science 3: 432.
- Kiran KK, Chandra TS (2008) Production of surfactant and detergent-stable, halophilic, and alkalitolerant alpha-amylase by a moderately halophilic Bacillus sp strain TSCVKK Appl Microbiol. Biotechnol 77:1023-1031.
- Femi-Ola TO, Olowe BM (2011) Characterization of alpha amylase from Bacillus subtilis BS5 Isolated from amitermes evuncifer silvestri. Research Journal of Microbiology 6: 140-146.
- Seitz HK, Egerer G, Simanowski UA, Waldherr R, Eckey R, et al. (1993) Human gastric alcohol dehydrogenase activity: effect of age, sex, and alcoholism. Gut 34:1433-1437.
- Dickson K, Monger B (1973) Effects of temperature and pH on dehydrogenase activity extracted from microbial sources.Biores Tech 4:34-67.
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 7886
- [From(publication date):
July-2017 - Aug 19, 2025] - Breakdown by view type
- HTML page views : 6757
- PDF downloads : 1129