Effects of Gamma Irradiation on the Molecular and Physical Detection Properties of Bacillus Spores
Received: 25-Oct-2017 / Accepted Date: 05-Jan-2018 / Published Date: 13-Jan-2018 DOI: 10.4172/2157-2526.1000158
Abstract
Gamma irradiation is a technique for inactivation of biological warfare agents (BWAs). Detection of these irradiated agents by sensor technologies may be affected,; altering sensitivity compared to that of non-irradiated organisms. Here for the first time, we aim to determine if spore morphology, or other physical or chemical properties beyond viability and PCR, is altered by the gamma irradiation process. This study analyzed common detection methods to determine the effects the irradiation process has on Bacillus thuringiensis subspecies kurstaki (Btk) spores compared to nonirradiated spores. Liquid concentrations of 108, 106, and 104 CFU ml-1 of Btk spores and one dry powder sample of 109 CFU g-1 were exposed to varying levels of gamma irradiation. Cell viability studies showed complete inactivation for all concentrations at doses of 5.43 kGy and higher. PCR analysis indicated no loss in sensitivity with increasing doses for both wet and dry spore samples. Visual inspection of the spores through scanning electron microscopy showed a change in morphology as the dosage of irradiation increased. With The inability to distinguish whole spores from cell debris occurred in at the 10.86 kGy samples level. Similarly, fluorescence readings remained stable among all but one dose with only the 10.86 kGy sample showing an increase in fluorescence. For PCR based detection methods, no bias towards irradiated versus non-irradiated spore samples exist while optical detection technologies are likely affected by the physical changes to spore morphology and rupturing. Results from this study demonstrate that gamma irradiation of Bacillus spores causes damage to the organism which may make them unrecognizable alter how they to are perceived by detection technologies. When using irradiated analyte to evaluate performance of detection technologies, the results may be impacted by the irradiation process.
Keywords: Bacillus; Gamma irradiation; Detection; Physical properties; Fluorescence; Spore
Introduction
The intentional use of microbes to kill or harm individuals has been a tactic to overtake populations of people since the beginning of modern civilization regardless of the understanding, or lack thereof, of microbiology. Using biological agents during warfare date back to, and likely further than 1346 when the Mongols laid sedge on Caffa, Crimean Peninsula by catapulting bodies of plague victims over the city walls. In 1763, the British distributed blankets of smallpox patients to Native Americans, Confederate soldiers sold clothing from yellow fever and smallpox patients to Union troops in 1863, and most recently in 2001 weaponized Bacillus anthracis was distributed through the United States Postal Service [1]. The use of chemical and biological weapons during the First and Second World Wars was a major offensive tactic to demoralize, injure and kill combatants on the battlefield. As the threat for the use of these weapons became more prevalent in use and the potential for deployment more sophisticated, the need for detection capabilities of airborne threats also increased. As a result, the first chemical and biological detectors were produced and used in theatre starting in the early 1940s. The anthrax attack of 2001 highlighted that Biological Warfare Agents were no longer limited to the battlefield but could also be employed as a terroristic tactic in civilian populations and during peacetime, thus heightening the need for reliable and autonomous detectors both within and outside of the battlefield. In response to the 2001 anthrax attacks, a plethora of biological triggers, collectors, and identifiers have been evaluated for their effectiveness and since have been captured in market surveys [2]. The evaluation of such technologies is time consuming, inherently dangerous, and often cost prohibitive at breadboard and low developmental stages if live virulent biological material is used for early assessment. For those reasons, either live non-pathogenic near-neighbor organisms or inactivated threat agents are routinely used for biological detector testing.
There are a multitude of methodologies for inactivating microorganisms; heat, chemical treatment, ultra violet light, and gamma irradiation to name a few [3-5]. A common method of inactivation of BWAs used for research is exposure to gamma irradiation at levels which render the organism inactivated and unable to be cultured. Despite the significant amount of data demonstrating the effectiveness of gamma irradiation on the inactivation of BWAs, little data exist which characterizes any changes in physical or chemical properties and how these changes may affect the organism’s response to biological sensing technologies beyond viability and PCR sensitivity [4,6,7]. A practice to ensure that organisms that are inactivated through gamma irradiation are, in fact, completely inactivated is to expose the organisms to levels of irradiation that significantly exceed minimum thresholds for rendering BWAs unculturable. However, it is unknown how overexposure to gamma irradiation can affect the ability for detection technologies to detect and identify the over-exposed organisms. This uncertainty could lead to inaccurate conclusions as to the detection sensitivity of the technologies being evaluated. Although a number of recent papers have reported on the ability to perform PCR on irradiated material, no information is known regarding changes in morphology of individual spores and if those changes could affect recognition by detection sensors.
Autonomous biological detection systems are typically comprised of two or three components; trigger, collector, and identifier or simply a collector and identifier. Triggers are sophisticated systems that continuously monitor the air for changes in particle size, particle number, and emittance spectra of particulates in response to an excitation wavelength. The interrogation is performed by analyzing the emittance spectrum of particulates as they pass through specific wavelengths. In the case of biological material, much of the florescence is due to NADH, NADPH, flavins and amino acids – molecules which typically are found in abundance in the interior of the cell [8,9]. The three amino acids that are fluorophores, tryptophan, tyrosine, and phenylalanine, have excitation and emission maxima of 255, 275, and 280 nm and 282, 303, and 348 nm respectively. In the case of NADH and NADPH, the excitation and emission maxima are at 340 and 450 nm respectively, and flavin compounds have an excitation and emission maxima at approximately 450 and 520 nm respectively [10]. Other components of the atmosphere such as smoke, dust, pollens, and chemical compounds would likely not fit the spectral/size/number profile that a BWA cloud would present, thus not alarm a trigger. Collectors are less discriminatory than triggers and pull in ambient air without interrogation of the environment. Whether or not a trigger initiates collection to begin, or if the collector is running constantly as an independent unit, any particle that is in the portion of air being pulled into the collector is deposited onto or into the collection medium until it is extracted for analysis. Once extracted from the collection medium, the sample then undergoes identification analysis, mainly PCR [2]. A number of recent reports have demonstrated that Bacillus anthracis spores can be inactivated by gamma irradiation between 15 kGy and 25 kGy depending on the concentration of spores while the integrity of the genomic material remains stable enough to be amplified using qPCR [3,4,11]. A 2015 shipment of inadvertently live Bacillus anthracis spores, thought to have been inactivated by gamma irradiation and sent from Dugway Proving Ground, Utah to 194 laboratories across the United States and nine foreign countries, highlighted the need for further investigation into inactivation of spores through gamma irradiation [12]. A possible solution included the potential for exposing biological agents to doses of gamma radiation beyond what has been identified for inactivation [3,4,6,11]. However it is unknown what effect dosages of gamma radiation beyond that previously reported as sufficient for inactivation of spores will have on the sensitivity of detection technologies with respect to bacterial properties such as fluorescence response and changes to spore morphology. This study investigates the effect gamma irradiation has on PCR, fluorescence response, and spore structure as a function of liquid or dry preparation, sample concentration, and exposure to varying doses of gamma irradiation. For the reasons mentioned above, primarily cost and health hazard, that make virulent Bacillus anthracis less desirable to use as a test material for early stages of technology assessment as well as its common use as a surrogate for Bacillus anthracis, we have chosen to perform our study using the nonvirulent near neighbor Bacillus thuringiensis subspecies kurstaki as our microorganism of choice for this study.
Materials and Methods
Large-scale production of B. thuringiensis subsp. kurstaki strains for dissemination
Wild-type (WT) B. thuringiensis subsp. kurstaki (ATCC 33679) and barcoded B. thuringiensis subsp. kurstaki spores derived from ATCC 33679 (2) were grown in NZ-Amine A growth medium consisting of the following (in grams per liter unless other units noted): 10.0 glucose, 5.0 casein peptone type S, 1.0 yeast extract, 4.0 K2HPO4, 3.0 KH2PO4, 0.134 CaCl2 • 2H2O, 0.02 FeSO4 • 7H2O, 0.05 MgSO4 • 7H2O, 0.023 MnSO4 • H2O, and 0.02 ZnSO4 • 7H2O, and 0.05 ml antifoam 204 (Sigma- Aldrich) per liter of culture volume. All medium ingredients were autoclaved together with the exception of glucose and metals. Glucose filtered through 0.8/0.2-_m filters was aseptically added to the sterile media. The two metal solutions (one containing CaCl2• 2H2O, FeSO4 • H2O, MgSO4 • H2O, and MnSO4 • H2O and the other containing the ZnSO4 • 7H2O) were separately prepared, filtered sterilized, and added to prevent precipitation of CaSO4. One-liter seed cultures in a 4-liter flask were prepared by inoculating with two vials of frozen stock and incubating (Innova 4300 shaking incubator; New Brunswick Scientific) at 30°C and 200 rpm until the optical density at 600 nm (Genesys 20 spectrophotometer; ThermoSpectronic) reached approximately 0.4 optical density units. The 1-liter seed cultures were aseptically transferred by a presterilized 2-liter transfer bottle into the 100-liter fermentor (IF 150; New Brunswick Scientific) containing the presterilized NZ-Amine A medium. The operating conditions for the 100-liter fermentor were controlled at 240 rpm, an airflow of 1 vvm (i.e., air volume per liquid volume per minute), 30°C, and pH 7.0 (with 3 M H3PO4 and 3 M NaOH). When the percentage of spores (as estimated by phase-contrast microscopy [microscope model BX51; Olympus]) exceeded 95%, the spore suspension was heat-shocked for 1 h at 70°C. After cooling down to 30°C, the spore suspension was diafiltrated (0.2-_mPellicon TFF filters; Millipore) to approximately a 20-liter retentate volume, washed five times with deionized water, and concentrated to a final volume of approximately 10 liters. The washed and concentrated 10-liter spore suspension was spray dried (Niro) and subsequently milled (Sturtevant) with Aerosol 202 (Evonik). The spraydried spore preparation was characterized in terms of particle size distribution (using a TSI, Inc., aerodynamic particle sizer [APS 3321]), moisture content (Karl Fischer C20 moisture analyzer; Mettler-Toledo), and viability (serial dilution and plate counting). The final dried and milled products contained 1.1_1011 (WT B. thuringiensis subsp. kurstaki) and 2.6_1011 (barcoded B. thuringiensis subsp. kurstaki) CFU/g. Analysis of a representative preparation using a Petroff-Hauser chamber revealed that approximately 90% of total spores were viable prior to dissemination.
Liquid sample preparation
Liquid spore samples were prepared by adding five grams of dried and milled spores to 100 ml of sterile water followed by 25 seconds of vortexing to create the stock solution. Three subsequent working suspensions were made by adding 1 ml of the preceding suspension to 99 ml of sterile water. Cell enumeration to establish the stock concentration of spores in was performed by serial dilution on Trypticase Soy Agar plates in triplicate and incubated overnight at 37°C. Working stock concentrations from plate counting were 1.8 × 108 CFU ml-1, 1.6 × 106 CFU ml-1, and 6.7 × 104 CFU ml-1 respectively. All stocks were kept at 4°C when not in use.
Irradiation and viability of samples
One ml of each liquid spore suspension (one per concentration) and 0.1 gram (109 CFU g-1) of powder sample were each placed in separate 1.5 ml microcentrifuge tubes for irradiation at the following levels: 0, 1.5, 2.0, 2.5, 4.15, 5.43, 8.2, and 10.86 kGy. Samples were Irradiated with a JL Shepard 484R-2 cobalt (60Co) irradiator (San Fernando, CA) for 23, 31, 39, 64, 84, 127, and 169 minutes respectively. Viability of spores post irradiation was performed by six 10-fold serial dilutions of 100 μl of irradiated suspension onto TSA plates in triplicate. The plates were incubated at 37°C overnight and viable colonies were counted by hand. The serial dilution in which 30 - 200 CFUs per plate were observed was the serial dilution used to calculate viability.
Real time polymerase chain reaction
Samples were analyzed for their ability to be identified via PCR by using the ABI 7900HT Fast Real-Time PCR System (Thermo Fisher Scientific Inc., Waltham, MA). Bacillus thuringiensis kurstaki Target 1 FastBlock Master Mix (catalog number PCR-BTK-1FB-K) was obtained from the Defense Biological Product Assurance Office (DBPAO) for Btk plasmid detection. The master mix utilizes a 6-carboxyfluorescein (FAM) fluorescent reporter at the 5’ end of the probe, a tetramethylrhodamine (TAMRA) quencher at the 3’ end, and Platinum® Taq DNA polymerase. Samples were tested by combining 5 μl of spore sample with 15 μl of master mix and cycling in two stages. Stage 1 consisted of 2-minute incubation at 50°C followed by a 20 second hold at 95°C. Stage 2 consisted of 1 second at 95°C followed by 20 seconds at 60°C, repeated for 45 cycles.
Fluorescence methods
Five hundred microliters of the 108 CFU ml-1 iIrradiated samples and controls were vortexed in an additional 5 ml of sterile water and allowed to settle overnight. One ml of supernatant was transferred to 3 ml of water and vortexed. Approximately 2 ml of the diluted sample was added to a 10 mm × 10 mm quartz cuvette for measurement on the JASCO FP-8300 Fluorescence Spectrometer (Gross-Umstadt, Germany) utilizing a Xenon light source. Samples were exposed to an excitation wavelength of 290 nm and the measurement range of 295- 500 nm was recorded at intervals of 0.5 nm with a scan speed 500 nm min-1. 5 ml of sterile water was used as the control and subtracted from the fluorescent signature of the test samples.
SEM imaging
Preparations of spores for visualization were generated by drying 25 μl of spore suspension onto SEM stubs in a biosafety cabinet overnight and were visualized with a JEOL JCM-5700 (Peabody, MA) scanning electron microscope under ambient conditions.
Results
Viability post-irradiation
Inactivation of microbial pathogens through high doses of gamma irradiation has been demonstrated many times and in some cases, detrimental effects on detection and identification have been reported [6]. While it is essential to understand what level of gamma irradiation is essential to render all organism in a sample unculturable, the level must not affect the properties that the detection and identification technologies are interrogating. Plating aliquots of irradiated Btk spore samples revealed that the ability to culture colonies was dependent upon the dosage of irradiation the samples received; effectively zero culturable colonies grew at doses of 5.43 kGy or higher (Figure 1). However, at the 5.43 kGy dose and above, on some plates a small number (≤ 3) of colonies were observed on one or two plates of a respective replicate. Graphically, these are shown as zero due to their the unreliability to in the ability reproduce these results through additional plating and statistical insignificance [13].
Effects of irradiation on PCR sensitivity
To determine the effect of irradiation on the integrity of the genomic material used for detection and identification, real time PCR was performed at each irradiation level for each sample preparation and concentration. Our results found that increased exposure to radiation at the levels used in this study resulted in a < 3 cycle difference (less than 1 log difference in template concentration) in qPCR threshold cycle (Ct) values as compared to non-irradiated control samples of identical concentration and preparation; thus not significantly affecting the ability to amplify and identify the desired sequence target (Figure 2). The Ct values for the powder and liquid 108 CFU ml-1 samples were 19.9 and 19.5, respectively. The Ct values for the 106 CFU ml-1 samples and 104 CFU ml-1 samples were higher at 25.6 and 32.0, respectively, and consistent across all irradiation levels at their respective concentrations.
Figure 2: PCR analysis. PCR analysis of Btk spore preparations at 104 cfu/ml, 106 cfu/ml, 108 CFU ml-1, and dry powder at a concentration of 109 CFU/gram (0.1 g was suspended in liquid to make 1 mL for this chart) exposed to various doses of radiation. Irradiated and non-irradiated control spore preparations were amplified in duplicate and averaged to determine plotted threshold cycle (Ct) values. PCR data plotted as the Ct value versus radiation exposure in kilogray (kGy).
Spore morphology
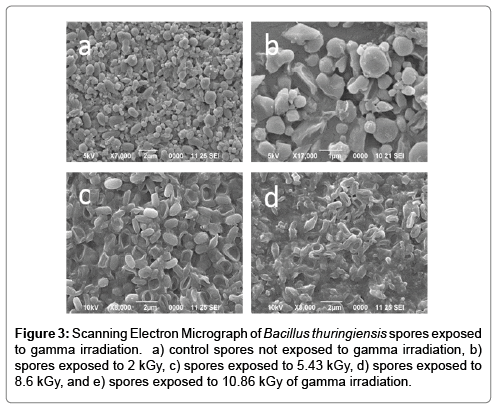
BWA detectors use background conditions as a baseline for “seeing” a potential pathogen. Particle number, particle size, light scattering and fluorescence are some of the conditions which triggers use to sense a potential threat. We viewed individual spores under SEM to determine if gamma irradiation has an effect on spore morphology as particle shape and size could affect the optical properties of the threat agent. We report that exposure to irradiation had a significant impact on the morphology of individual Bacillus thuringiensis spores at high levels. Spores which were not exposed to gamma irradiation had a very typical rod-like shape with a smooth surface (Figure 3). As the spores are exposed to increasingly more irradiation, they begin to wrinkle and look like deflated footballs, Brady balls, and eventually rupture displaying a donut-like appearance. At 10.86 kGy a mixture of the burst spores and spore debris was observed, with no intact spores present.
Effect on fluorescence
Biological particles have a distinctive composition that, when excited by specific wavelengths, will respond by emitting a unique spectra. Emission from the specific excitation wavelengths allow for the pathogens to be distinguished from non-threats and detected by trigger systems as a potential hazard. To determine if gamma irradiation altered the spore in a way which would result in an abnormal response to commonly used excitation wavelengths, we recorded the fluorescence of each sample from 295-500 nm when exposed to an excitation wavelength of 290 nm, close to the excitation wavelength of phenylalanine, a known fluorophore (Figure 4). The non-irradiated control and all samples other than the 10.86 kGy sample resulted in similar spectra patterns and relative intensities. The 10.86 kGy sample had the highest intensity value, peaking at 2.7 Arbitrary Units (AU), while control 1.5, 2.5, and 5.43 kGy samples all peaked at less than 2 AU.
Discussion and Conclusion
Irradiation of Bacillus thuringiensis spores at a level almost twice what is needed to render inactivated had very little effect, if any at all, on the stability of the DNA as evidenced by the ability to identify specific targeted genomic material through qPCR. The consistent Ct values within each respective sample preparation, in addition to the visualization of DNA extractions on agarose gels (Supplementary Figure 1) and electropherogram comparison (Supplementary Figure 2), did not show any apparent degradation or breakage of DNA. These results are not unexpected as tThe complete genome of Btk is 6.4Mbs and the target in our study was only ~80bp; thus the likelihood of a single breakage within the sequence of interest is very minute [14]. These results differ from recent papers which investigated both DNA fragmentation as a result of gamma irradiated damage and qPCR of Bacillus anthracis. However, the dosage of irradiation given to those samples was 10-fold higher than the dosage needed in our study to render the material unculturable [3,6]. The focus of our study was to identify low irradiation levels that can inactivate the pathogen and investigate the subsequent effects on detection. Thus we did not expose our samples to the higher dosage used in Bacillus anthracis studies. However we do believe that if we were to expose our samples to the same levels of irradiation, we would expect to see similar fragmentation results due to the organisms used in both studies being belonging in the same genus. The dry powder sample (concentration of 108 CFU ml-1 after 0.1g [109 CFU g-1] of power was re suspended in 1 ml of water), had nearly identical Ct values at each irradiation level although the ability to culture viable colonies declined at a much greater rate as the level of irradiation increased. In fact, no colonies were observed from the powder sample when exposed at a dose of 4.15 kGy while the liquid sample of the same concentration and irradiation dose resulted in 5 logs of CFUs. A potential reason for this result is that there is significantly less available gamma absorbing water in the powder sample; the only available water is inside the spore and even then, there is very little present. A greater proportion of the energy during irradiation is absorbed by the internal contents of the spores as opposed to the additional external water in the liquid suspensions. Unfortunately, the data presented here do not provide direct evidence towards the specific mechanism for the inactivation of the spore: a chemical reaction involving gamma-induced reactive oxygen species degradation or a thermal reaction. It is widely known that DNA is sensitive to cleavage in the presence of free radicals and that hydroxyl radicals are products in the gamma radiolysis of water [15,16]. However, it is unclear in our study as to the exact mechanism of gamma irradiation in vivo but presumably the production of free radicals may have a detrimental effect on DNA and even possibly protein stability. Longer periods of exposure to irradiation resulted in a “deflated”, deformed, and ultimately ruptured spore morphology in individual spores (Figure 3). The 10.86 kGy samples had a significantly higher peak in spectral emittance (Figure 4) likely due to the breaking of the spore coat and leakage of the internal components including fluorophores of the spore into the supernatant. This observation is consistent with recent reports which demonstrate that both heat and plasma treatment lead to similar spore morphologies that we observed in this study [17,18]. Kim et al. investigated fluorescence as an effect of inactivation treatment and also found a correlation between spore morphology and spectral deviation from the control samples [17]. Molva and Baysal demonstrated that the OD260 and OD280 ratios and spore morphology also changed as heat treatment increased [18]. During the process of thermo cycling, spores were are subjected to high temperatures resulting in the breaking of spore coat and release of genomic material. It would seem logical that non irradiated spores exposed to thermo cycling would be indistinguishable from irradiated samples with similarly broken spore coat and internal components released into the supernatant provided levels of irradiation were not high enough to cause fragmentation. Gamma irradiated spores are potentially a suitable simulant for BWA identification detection technologies that rely solely on PCR for identification. Despite receiving a high enough dosage of gamma radiation energy to rupture the spore coat and thus releasing the contents of the spore into the surrounding liquid, there is very little or no effect on the sensitivity of PCR compared to that of control samples which were not irradiated. Visualization of DNA on agarose gels (Supplementary Figure 1) confirms that there is no noticeable degradation of genomic material at exposures up to 10.86 kGy. However, our study did identify that there were a few colonies at 5.43 kGy and greater that were still culturable. This could be problematic for studies which utilize pathogenic material as some resistant colonies have the potential to be infectious at the doses reported here and should likely not be used outside of an appropriate bio safety level laboratory unless irradiated at significantly higher doses. Additionally, high dosages of gamma irradiation may have a significant impact on detection technologies, specifically those which use optical approaches to detect airborne BWAs. Investigating the different ways spores may be affected by gamma irradiation has demonstrated that irradiated material can be an appropriate simulant for live pathogenic material given the that physical and optical recognition properties are not necessary for testing or research as inactivation levels and PCR sensitivity are not affected.
Acknowledgements
The authors would like to thank Matthew Browe of the CBR Filtration Branch at Edgewood Chemical and Biological Center for performing the fluorescence measurements in this study. The authors would also like to thank Tiffany Sutton and Stacey Broomall for suggestions in experimental setup and comments that improved the manuscript. Funding which enabled this project to occur was provided by the United States Department of Homeland Security, Science and Technology Division.
Conflict of Interest
The authors certify that there is no conflict of interests, financial or other, in the subject matter or materials discussed in this manuscript.
References
- Frischknecht F (2003) The history of biological warfare. Human experimentation, modern nightmares and lone madmen in the twentieth century. EMBO reports 4: S47-52.
- Dang JL, Heroux K, Kearney J, Arasteh A, Gostomski M, et al. (2001) Bacillus spore inactivation methods affect detection assays. Appl Environ Microbiol 67: 3665-3670.
- Dauphin LA, Newton BR, Rasmussen MV, Meyer RF, Bowen MD (2008)  Gamma irradiation can be used to inactivate Bacillus anthracis spores without compromising the sensitivity of diagnostic assays. Appl Environ Microbiol 74: 4427-4433.
- Moeller R, Raguse M, Reitz G, Okayasu R, Li Z, et al.(2014) Resistance of Bacillus subtilis spore DNA to lethal ionizing radiation damage relies primarily on spore core components and DNA repair, with minor effects of oxygen radical detoxification. Appl Environ Microbiol 80: 104-109.
- Broomall SM, Ait Ichou M, Krepps MD, Johnsky LA, Karavis MA, et al. (2015) Whole-genome sequencing in microbial forensic analysis of gamma-irradiated microbial materials. Appl Environ Microbiol 82: 596-607.
- Fiester SE, Helfinstine SL, Redfearn JC, Uribe RM, Woolverton CJ (2012) Electron beam irradiation dose dependently damages the bacillus spore coat and spore membrane. Int J Microbiol 2012: 579-593.
- Hill SC, Pan YL, Williamson C, Santarpia JL, Hill HH (2013) Fluorescence of bioaerosols: mathematical model including primary fluorescing and absorbing molecules in bacteria. Opt Express 21: 22285-22313.
- Lakowicz J (1983) Principles of fluorescence spectroscopy, Plenum Press, New York.pp: 111-150.
- Pan YL (2015) Detection and characterization of biological and other organic-carbon aerosol particles in atmosphere using fluorescence. J Quant Spectrosc Radiat Transf 150: 12-35.
- Horne T, Turner GC, Willis AT (1959) Inactivation of spores of Bacillus anthracis by gamma-radiation. Nature 183: 475-476.
- Ostrowski MPA (2015) Individual and institutional accountability for the shipment of viable bacillus anthracis from dugway proving ground.AR 15-6 Investigation Report. pp:1-157.
- Goldman E, Green LH (2008) Practical Handbook of Microbiology (2nd edn.). CRC Press, Florida.
- Day M, Ibrahim M, Dyer D, Bulla L (2014) Genome sequence of Bacillus thuringiensis subsp. kurstaki Strain HD-1. Genome Announc 2: e00613-e00614.
- Balasubramanian B, Pogozelski WK, Tullius TD (1998) DNA strand breaking by the hydroxyl radical is governed by the accessible surface areas of the hydrogen atoms of the DNA backbone. Proc Natl Acad Sci U S A 95: 9738-9743.
- LaVerne JA (2000) OH radicals and oxidizing products in the gamma radiolysis of water. Radiat Res 153: 196-200.
- Kim JY, Lee IH, Kim D, Kim SH, Kwon YW (2016) Effects of reactive oxygen species on the biological, structural, and optical properties of Cordyceps pruinosa spores. Royal Society of Chemistry 6: 30699-30709.
- Molva C, Baysal AH (2015) Analysis of alicyclobacillus acidoterrestris spores from different sporulation media subjected to wet-heat. Microbiol Res 6: 6219.
Citation: Edmonds JM, Love CE, Harvey T (2018) Effects of Gamma Irradiation on the Molecular and Physical Detection Properties of Bacillus Spores. J Bioterror Biodef 9: 158. DOI: 10.4172/2157-2526.1000158
Copyright: © 2018 Edmonds JM. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 10185
- [From(publication date): 0-2018 - Dec 19, 2025]
- Breakdown by view type
- HTML page views: 9168
- PDF downloads: 1017