Research Article Open Access
Observations on the Migration of Bacillus Spores Outside a Contaminated Facility During a Decontamination Efficacy Study
Erin E. Silvestri1, Sarah Perkins2, Robert Lordo2, William Kovacik3, Tonya Nichols1, Charlena Yoder Bowling1, Dale Griffin4*and Frank W. Schaefer1
1U.S Environmental Protection Agency, National Homeland Security Research Center, 26 W. Martin Luther King Drive, MS NG16, Cincinnati, OH 45268, USA
2Battelle Memorial Institute, 505 King Avenue, Columbus, OH 43201, USA
3Pegasus Technical Services, Inc., 46 East Hollister St., Cincinnati, OH 45219, USA
4U.S Geological Survey, Coastal and Marine Science Center, 600 4th Street South, St. Petersburg, FL 33701, USA
- *Corresponding Author:
- Dale Griffin
U.S Geological Survey
Coastal and Marine Science Center
600 4th Street South
St. Petersburg, FL 33701, USA
Tel: 850- 274-3566
Fax: 727-502-8001
E-mail: dgriffin@usgs.gov
Received Date: March 24, 2015; Accepted Date: April 29, 2015; Published Date: May 06, 2015
Citation: Silvestri EE, Perkins S, Lordo R, Kovacik W, Nichols T, et al. (2015) Observations on the Migration of Bacillus Spores Outside a Contaminated Facility During a Decontamination Efficacy Study. J Bioterror Biodef 6:135 doi:10.4172/2157-2526.1000135
Copyright: © 2015 Silvestri EE, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Bioterrorism & Biodefense
Abstract
The fate and transport of Bacillus anthracis spores in indoor and outdoor environments is not well understood. The Bio-Response Operational Testing and Evaluation exercise evaluated decontamination technologies in a twostory building experimentally contaminated with Bacillus atrophaeus subspecies globigii spores. The Bio-Response Operational Testing and Evaluation project provided a means to evaluate the potential for the spores dispersed inside the building to migrate to the outside as well as to investigate a new method for processing soils contaminated with Bacillus spores. Duplicate sterile sand samples were placed within the tent covering the building, but outside the building itself, near entrances, exits, and high-traffic areas to assess migration and deposition of newly disseminated spores. The sand samples were utilized during three stages of the decontamination study: before spore dissemination, after spore dissemination, and after decontamination of the building. In addition, two sets of sand samples placed within the building provided positive controls. Results from two different building decontamination approaches were studied. Results were tabulated as presence or absence rather than as a quantitative figure. There was no significant association among positive samples and the location of the samples around the building. There was a significant association between the different stages of each decontamination study and the number of detectable samples. The results of this study demonstrate the potential for spores to migrate out of a contaminated building and the importance of considering migration when assessing the scope of a contamination incident
Keywords
Bacillus anthracis; Bacillus atrophaeus; Spores; Migration; Suspension; Dispersion; Transport
Abbreviations
ATD: Arizona Test Dust; B1-B2: Sample locations inside the building; BOTE: Bio-Response Operational Testing and Evaluation; BROOM®: Building Restoration Operations Optimization Model; CFU: Colony Forming Unit; CT: Cycle Threshold; Decon: Decontamination; Dissem: Dissemination; DNA: Deoxyribonucleic acid; EPA: U.S. Environmental Protection Agency; HVAC: Heating, Ventilation, and Air Conditioning; LOD: Limit of Detection; ND: Non-Detect; NHSRC: National Homeland Security Research Center; PBST: Phosphate Buffered Saline TWEEN-20; PCR: Polymerase Chain Reaction; Qpcr: Quantitative- Polymerase Chain Reaction; TSB: Tryptic Soy Broth; Tween®20: Polyoxyethylene (20) sorbitan monolaurate; USGS: U.S. Geological Survey
Introduction
The potential for an intentional wide-area or indoor release of Bacillus anthracis spores remains a concern, but the fate and transport of B. anthracis spores in indoor and outdoor environments are not well understood. Some studies have examined the possibility of spore transport within ventilation systems and in buildings [1] and transport into a building following an outdoor release [2]. Little research exists regarding the potential for spores to migrate to the outside of a building following an indoor release.
Bacillus species spores have the potential to remain viable in the soil for many years [3-5]. Lasting environmental contamination following a release is a possibility [6], and planning for site characterization and remediation activities should consider both indoor-to-outdoor spore transport and outdoor soil as potential exposure pathways.
The U.S. Department of Homeland Security and the U.S. Environmental Protection Agency (EPA) conducted the Bio-Response Operational Testing and Evaluation (BOTE) project from April to May 2011 to perform field-level facility biological-remediation studies of various decontamination technologies.
During the BOTE exercise, a vacant two-story building at Idaho National Laboratory was experimentally contaminated with Bacillus atrophaeus subsp. globigii spores, which served as a surrogate for virulent B. anthracis spores. The building was covered with a tent to retain any off-spray or fumigant off-gassing from the decontamination agents used during the study and to maintain negative pressure between the interior of the facility and the surrounding areas outside the secondary enclosure. The disinfectants used for this work were pH-amended bleach (liquid application) and chlorine dioxide (fumigant); however, the disinfectants were not the focus of the work reported in this paper and will not be described.
In a preliminary evaluation of soil samples taken from outside the test facility prior to this study, the soil samples were found to be contaminated with B. atrophaeus subsp. globigii spores. The spores might possibly have migrated to the outside of the building and deposited in the soil adjacent to the building during previous exercises held at the facility. Likely routes would include movement through loose seals.
Around doors and windows or via foot traffic of personnel conducting decontamination. The BOTE test bed provided an opportunity to test this hypothesis and evaluate the potential for the B. atrophaeus subsp. globigii spores dispersed inside the building to migrate to the outside. It should be noted that the spore migration study was not a primary goal of the BOTE study, and the study design was limited to the confines of the original BOTE study. The design and subsequent results for the spore migration study are discussed below for the pH-adjusted bleach and chlorine dioxide gas- technology testing rounds. A new method for processing soil samples prior to DNA extraction and further sample analysis was also investigated and is described herein.
Materials and Methods
Bacillus atrophaeus subspecies globigii
The B. atrophaeus subspecies globigii spores (American Type Culture Collection 9372) were used for the BOTE project and have also been used for a number of fumigation and dispersal studies previously [7,8]. The spores were prepared at the Critical Reagents Program Antigen Repository of the Department of Defense. In brief, the preparation of the spores followed the procedure reported by Brown et al. [9] using tryptic soy broth (TSB) supplemented with 3 mg/L MnSO4. When the suspension contained 80 to 90% spores, the suspension was pelleted by centrifugation. Dry spore material was prepared from the pelleted material using a laboratory spray dryer. The dried material was mixed with fumed silica particles. The spore/silica mixture was milled to a uniform particle size. The resulting preparation contained approximately 1011 viable spores/g.
Contamination and Decontamination of the Building
For the BOTE decontamination study, nebulizers were used to disseminate the spores to achieve an approximate concentration of 106 spores/ft2 on the first floor surfaces and 102 spores/ft2 on the second floor surfaces of the test building. Ventilation and air conditioning systems functioned as air handling units for each floor of the facility, and was shut-off two hours after release of spores in the building. Negative air machines controlled airflow within and into the facility. Following contamination, site characterization sampling occurred. Once the extent of contamination was determined, the building was decontaminated. Post-disinfection sampling was conducted to determine the success of the decontamination effort. Selection of Matrix and Preparation of Sample Kits.
Due to the presence of B. atrophaeus subsp. globigii around the test site as noted above and the fact that the spore migration study was conducted as a secondary study to the BOTE study, an uncontaminated matrix was required. Because Bacillus spores are reported to bind to soils composed of humic and calcareous components [10], Pro-Com® silica sand (Cat # 4315024; Ace Hardware Corporation, Oak Brook, IL) was selected as the matrix.
To prepare the samples, 50 g aliquots of sand were placed into aluminum weigh boats and heat-sterilized at 250°C for 10 hours in a quick-dry oven (model 3096, Forma Scientific, Inc., Marietta, OH). Sterility (no observed colony forming units [CFU]) was confirmed by inoculating the sand into TSB tubes and incubating them overnight at 35°C. The TSB tubes were subsequently sub cultured onto tryptic soy agar plates and incubated at 35°C overnight as a final test of sterility. After sterility was confirmed, the 45 g aliquots of sand were transferred aseptically to sterile polystyrene 150 mm plastic Petri dishes (BD Biosciences, San Jose, CA; Falcon Cat # 25373-187), sealed with Parafilm® (Pechiney Plastic Packaging Company, Chicago, IL) and secured with cellophane tape. The sealed Petri plates of sand were bagged in lots of 10 for the exercise.
Placement and Collection of Samples
The prepared sand samples were labeled with bar codes and bagged individually. For each decontamination technology round, there were three sampling times. Pre-dissemination samples were placed the day before dissemination of spores during setup for each round and collected the morning of dissemination to assess spore levels prior to dissemination. The post-dissemination samples and the postdecontamination samples were placed at the same time, immediately after collecting the pre- dissemination samples. Post-dissemination samples were collected following both the dissemination of spores and after the indoor surface sampling efforts were finished to assess spore migration from the building and spore presence following dissemination (approximately 30 hours and 45 hours for VHP and chlorine dioxide rounds respectively). Post-decontamination samples were collected following decontamination of the building and after subsequent indoor surface sampling efforts were completed to assess spore presence amassed throughout the entire round via spore migration from the building.
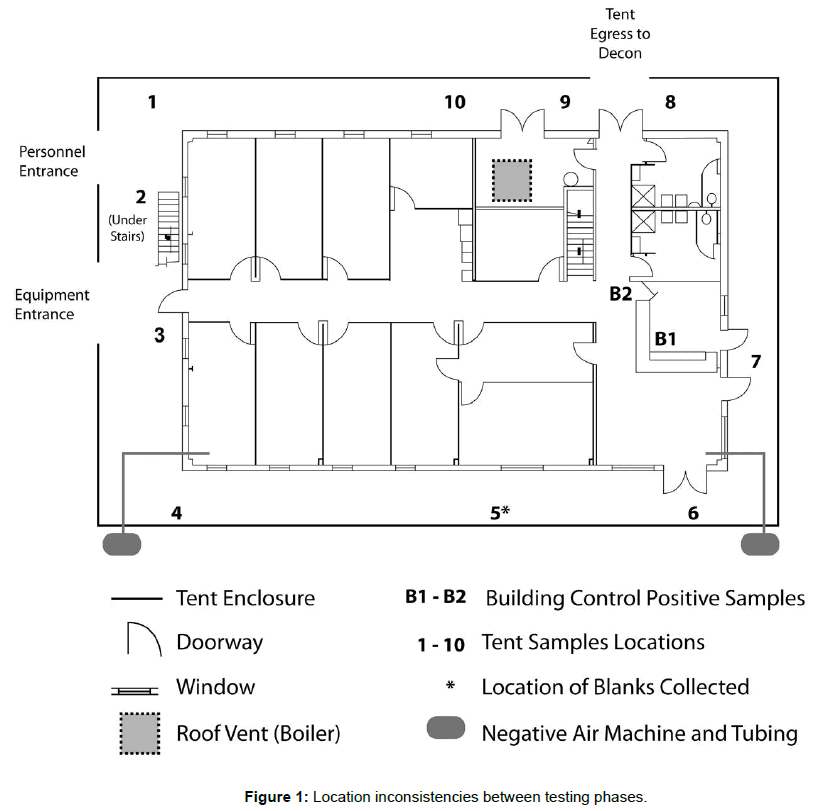
All samples were placed in orange-painted trays to aid visibility and to minimize location inconsistencies between testing phases. Duplicate samples (A and B) were placed around the building between the building wall and the tent, near entrances, exits, and high traffic areas (n=20 for each round; Figure 1). Two sets of duplicate samples were placed inside the building on the first and second floors (not included in Figure 1 as this floor was only utilized for the positive control sample) of the building to serve as positive controls (collected following dissemination) and to determine if the decontamination process interfered with the analytical process (collected following decontamination). Building samples were not collected prior to dissemination. The samples that remained in place during decontamination were intentionally not targeted for decontamination.
The sand samples were placed in an order to avoid walking by the sample tray multiple times. Placement began at tray 1 just inside the entrance to the tent, and moved counterclockwise around the building, placing samples on the tray in sequential order as shown in Figure 1. When placing sand samples inside the building, personnel went to the second floor of the building to place samples at location B2, passed through the airlock and down the stairwell to the first floor to place the samples at location B1. Personnel then exited the tent door to the personnel decontamination area. As personnel positioned the sand samples, they removed the lids of the Petri dishes.
Sand samples were retrieved in the same order in which they were positioned, capped with sterile lids, sealed with Parafilm® and secured with cellophane tape. Each sample was bagged individually.
Tracking labels with barcodes affixed to the individual sand sample dishes were scanned into the Building Restoration Operations Optimization Model (BROOM™, Sandia National Laboratories, Livermore, CA) [11] tool for tracking and time-stamping purposes. Throughout the exercise, samplers also collected site blanks and trip blanks midway through the retrieval process, away from the secondary enclosure entrances at location 5. The purpose of the site blanks was to determine the potential for background contamination of the sampling media at the site. Personnel opened the site blanks on site and then immediately closed, sealed, and re-bagged the site blanks for shipment to the laboratory for analysis. The purpose of the trip blanks was to determine the potential for sample contamination over the course of the entire exercise. Trip blanks were sent to the site but were never opened.
After collection of all samples, the exterior of the bags containing the sand samples was disinfected with Dispatch® bleach wipes (Medline Industries, Mundelein, IL). The bags were then packed and shipped at ambient temperature to the EPA in Cincinnati, OH, via an overnight carrier for analysis. Chain of custody forms generated by the BROOM tool were utilized.
Laboratory analysis
The samples were processed and analyzed for the presence of B. atrophaeus subsp. globigii DNA using a quantitative Polymerase Chain Reaction (qPCR). After the weight of each sand sample was determined and recorded, 45 g of sand was aseptically transferred to a 250 mL centrifuge tube (Corning Cat #430776) using a sterile powder funnel (Fisher, Cat 10-371D). The remaining 5 g was saved for a separate study, not described here. The Petri dish was rinsed with phosphate-buffered saline supplemented with TWEEN ®-20 (PBST, Sigma-Aldrich, St. Louis, MO), which was emptied into a centrifuge bottle. The volume in the centrifuge bottle was adjusted to 125 mL using additional PBST. After vigorous shaking for three minutes, the sample was allowed to settle for five minutes and the supernatant was transferred to a 250 mL Sorvall® centrifuge bottle (Sorvall, Cat #03937). Samples were centrifuged using a Sorvall Evolution® refrigerated centrifuge equipped with a fixed angle rotor (Fisher Scientific, Pittsburg, PA) at 5,900 g for 20 minutes with the brake set to 1 and a temperature of 4ºC to pellet the spores. The supernatant was then aseptically aspirated and discarded. The remaining pellet was resuspended in 25 mL PBST, and the suspension was transferred to a 50 mL conical centrifuge tube (Sorvall, Cat #03072). Suspensions were placed in the Sorvall HS-4 swinging bucket rotor and centrifuged at 5,900 g for 20 minutes with the brake set to 1 and a temperature of 4ºC. Any remaining supernatant was aseptically aspirated and the pellet was saved for DNA extraction (Collaborative unpublished work in our laboratories has indicated percent Bacillus spore recoveries from sandy soil range between 54 and 88%). The entire pellet was used for DNA extraction following the vacuum-based protocol from the PowerSoil® DNA Isolation Kit (MO BIO Laboratories, Inc., Carlsbad, CA).
The extracted DNA was analyzed for the presence of B. atrophaeus subsp. globigii DNA using the qPCR protocol described by Kane et al. [12]. The ABI Prism® 7900HT (Life Technologies Corporation, Carlsbad, CA) was used for qPCR analysis, and samples were analyzed in triplicate for 45 cycles. Each reaction consisted of 2 μL template DNA within a 20 μL PCR. Ninety-six-well plates were used to contain the experimental samples, three no-template controls, three positive controls (B. atrophaeus subsp. globigii DNA) and three negative controls (Escherichia coli DNA). Specificity of qPCR detection was determined by comparing the results to nontarget DNA (Escherichia coli) and positive control B. atrophaeus subsp. globigii DNA. Criteria for acceptance of negative qPCR controls were that all replicate samples be negative. Acceptance of analytical results for positive controls required the observed cycle threshold (Ct) to be within 5% of the prior determined Ct.
Instrument limits of detection (LODs) were determined based upon a concentrated stock solution of purified B. atrophaeus subsp. globigii DNA that was diluted eight-fold. The DNA concentrations ranged from 1.0×105 to 1.0 × 10-2 genomic equivalents. The overall recovery for the sand extraction and analysis method (referred to in this paper as the environmental LOD) was determined with matrix spikes. Blind 45 g sand samples were spiked in triplicate with B. atrophaeus subsp. globigii spore concentrations ranging from 1 to 106 spores/g sand. DNA from the spores collected from each aliquot of spiked sand was extracted by utilizing the same procedure as used for the actual samples Inhibition tests were conducted with the samples collected within the building to determine if residual decontamination chemical used during the BOTE project interfered with the PCR reactions. One sand sample from each floor (1 and 2) and each decontamination treatment round collected post- decontamination was selected for inhibition testing. In addition, one sand extract collected post- dissemination was assessed. Triplicate reactions using extracted template DNA from the original selected sand samples were run alongside triplicate reactions of the sample extract spiked with 10 genomic equivalents of standard B. atrophaeus subsp. globigii spore DNA (internal positive control) to allow for a low but reliable concentration of target DNA to be present within each spiked reaction tube.
Statistical analysis
Categorical data analyses were conducted to assess differences in the distribution of sand sample classifications (detected/non detected) that occurred between the study decontamination-treatment round, the stage of each round, and the location for the samples collected within the secondary enclosure (and outside the building). Statistical analyses to test for significant differences in the proportion of non-detected results between the pH-amended bleach and chlorine dioxide rounds were performed using Fisher’s Exact test [13] in Stata® software (Stata Corp LP, College Station, TX). A logistic regression analysis was fit to the detected/nondetected outcome data from two rounds to assess the extent to which testing round and sampling stage were statistically significant predictors to the proportion of non detected values (model also included a random effect for the sample location). To test the overall trend of each decontamination testing round, Fisher’s Exact test [13] was used to test for significant association between the proportion of non detected results and the sampling stages (pre-dissemination, postdissemination, post-decontamination), to compare the percentages of non detected results between the two disinfection rounds independently for each testing stage, to compare results across both testing rounds and to look at the proportion of non detected results by location. Logistical regression was used to further assess the significance of the samplingstage effect in the model.
Results
Limit of detection and quality control results
No B. atrophaeus subsp. globigii spores were detected in the site blank and trip blank quality assurance samples collected during each round. All qPCR quality assurance results met the acceptance criteria. For the instrument LOD determination, detectable results were attained down to 1.02 genomic equivalents/reaction at an average Ct of 38.29 (SD 0.08; n=3). At levels lower than ~1 genomic equivalent, the instrument registered “undetected.” Therefore, the instrument lower limit of detection was determined to be a Ct value of 38.3, or 1.02 genomic equivalents/reaction. All averaged Ct values greater than 38.3 were considered non-detectable based on the instrument LOD. Analysis of the environmental LOD determined that a minimum of 104 spores/g of sand was required for the average Ct value to be above the instrument LOD. Samples that yielded two or more “undetected” values were classified “non-detected” by the instrument. Detected results for the samples were assigned a degree of detection based upon the averaged Ct value for each sample (Table 1). The analysis code was based upon the instrument LOD (Ct of 38.3) as the lower boundary, and the calculated genomic equivalents/reaction for the cut-off points.
| Average Ct | Description | Code | GEq/ PCR reaction |
|---|---|---|---|
| Undetected or Ct ≥38.3 to <45.0 | Not Detected by the Instrument or Below the LOD | ND | ND |
| ≥36 to <38.3 | Very Weak Detection | 1+ | 1-3 |
| ≥34 to <36 | Weak Detection | 2+ | 3-10 |
| ≥32 <34 | Detection | 3+ | 10-40 |
| ≥30 to <32 | Strong Detection | 4+ | 40-150 |
| <30 | Very Strong Detection | 5+ | >150 |
Table 1: Analysis code descriptions based on mean ct value.
Results from the inhibition testing showed that the Ct value of the samples spiked with the internal positive control decreased when compared to the original sample extracts, or in other words, a previously non detected sample became detectable to the expected spiked concentration. The target DNA averaged a Ct of 33.9 (standard deviation 0.43) (data not shown). Based on these analyses, neither of the decontamination agents caused qPCR inhibition in the sand samples assessed during this study. These results cannot be used to determine if the decontamination agents may have affected the study results due to DNA degradation or other effects.
Sampling results and statistical analysis
In total, 64% (77/120) of the samples collected from the ten locations within the secondary enclosure (but outside the building) were classified as non-detectable by the instrument (Table 2). The lowest Ct value found was a Ct of 30.4. Table 3 shows that the largest proportion of detected results occurred at location 1, near the secondary enclosure personnel entrance. Here, four of the six sampling events led to detectable outcomes, although this difference was not statistically significant. Codified results for co-located sample pairs (A and B) are given in Table 3. Approximately 62% (37/60) of the sample pairs yielded consistent results where sample A results agreed with sample B results in terms of detection or non-detection (Table 3). For sample containers placed inside the building, spores were detected in all but one of the samples collected pre-decontamination (located at B2), and all but one of the post-decontamination samples were classified as non-detectable (B1 location) (Table 3).
| Description | pH-Adjusted Bleach | Chlorine dioxide | Total | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Pre-Dissem | Post-Dissem | Post-Decon | Total Samples | Pre-Dissem | Post-Dissem | Post-Decon | Total Samples | Combined | |
| Undetected by instrument | 12 | 6 | 18 | 36 | 18 | 10 | 13 | 41 | 77 |
| Detected by instrument | 8 | 14 | 2 | 24 | 2 | 10 | 7 | 19 | 43 |
| Total | 20 | 20 | 20 | 60 | 20 | 20 | 20 | 60 | 120 |
Note: Dissem: dissemination and Decon: decontamination
Table 2: Summary of the number of samples with descriptions.
| Location | pH-Adjusted Bleach | Chlorine dioxide | Total Number of Detected Samples Per Location | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Pre-Dissem | Post-Dissem | Post-Decon | Pre-Dissem | Post-Dissem | Post-Decon | ||||||||
| A | B | A | B | A | B | A | B | A | B | A | B | ||
| Outdoor Samples | |||||||||||||
| 1 | 4+ | ND | 2+ | 1+ | ND | 1+ | ND | ND | 1+ | 1+ | 1+ | 2+ | 8 |
| 2 | ND | ND | 1+ | 2+ | ND | ND | ND | ND | 1+ | 1+ | ND | ND | 4 |
| 3 | ND | ND | ND | ND | ND | ND | ND | ND | 1+ | 1+ | ND | ND | 2 |
| 4 | 1+ | 3+ | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND | 2 |
| 5 | 2+ | ND | 1+ | 1+ | ND | 1+ | ND | ND | ND | 1+ | ND | ND | 5 |
| 6 | ND | ND | ND | 1+ | ND | ND | ND | 1+ | ND | ND | 1+ | ND | 3 |
| 7 | 2+ | ND | 1+ | 1+ | ND | ND | 1+ | ND | 1+ | ND | ND | ND | 5 |
| 8 | 1+ | ND | 1+ | 1+ | ND | ND | ND | ND | ND | 1+ | ND | 1+ | 5 |
| 9 | ND | ND | 2+ | 2+ | ND | ND | ND | ND | ND | 1+ | 1+ | 1+ | 5 |
| 10 | 2+ | 1+ | 1+ | ND | ND | ND | ND | ND | ND | ND | ND | 1+ | 4 |
| Indoor Samples (Controls) | |||||||||||||
| B1 | 3+ | 4+ | ND | ND | 3+ | 3+ | 1+ | ND | 5 | ||||
| B2 | 1+ | 1+ | ND | ND | 1+ | ND | ND | ND | 3 | ||||
Note: Dissem: dissemination and Decon: decontamination. No indoor samples were taken pre-dissemination.
ND is instrument nondetection, 1+, 2+, 3+ and 4+ is instrument detection as noted in Table 1.
Table 3: Spore detection by decontamination technology, stage, location, and replicates (A and B).
Of the total samples collected for each round, 60 and 68% of samples in the pH-amended bleach and chlorine dioxide decontamination treatment rounds, respectively, were classified as non-detected results (Table 2), but the association between the percentage of non-detected results and the testing round was not significant (p=0.447). Logistical regression analysis also found no significant effect of testing round on the proportion of non-detected values (p=0.306).
An overall trend was noted (Table 2). As each decontamination testing round progressed, the rate of detection in the sand samples increased following dissemination and decreased following decontamination. Results for the test of overall trend of each decontamination testing round (Table 2) indicated that when the test was performed separately for each round, the difference in the proportion of non-detected samples among the testing stages was significant for both the pH-amended bleach round (p<0.001) and the chlorine dioxide round (p=0.019). The primary contributor to significance in both decontamination rounds was the difference between post-dissemination (stage 2), with a larger number classified as detected, and the other two stages, with a large number of classified as non-detected. When the test was used to compare the percentages of non detected results between the two disinfection rounds independently for each testing stage, the proportion of non detected results was not significantly different between the pHamended bleach pre-dissemination and chlorine dioxide round predissemination (p=0.065). Similar conclusions can be reached for the post-dissemination (p=0.333) and post-decontamination (p=0.127) stages. When the analysis was conducted across both testing rounds, i.e., the data from both decontamination technologies were lumped together for each testing stage, the association among the three stages was found to be highly significant (p=0.001). Again, this outcome was due primarily to the higher detection rates observed in postdissemination compared to the other two stages.
Logistical regression was used to provide further assessment of the significance of the sampling-stage effect in the model. The overall sampling-stage effect was highly significant (p=0.0009). As a result, comparison of the three stages of the two rounds was performed within the model fitting. The p-value of the logistical regression analysis comparing pre-dissemination to post-dissemination was 0.020, whereas the p-value for comparing pre-dissemination to post-decontamination was 0.791, and the p-value for comparing post-dissemination to postdecontamination was 0.001. Because significance was determined here at the 0.05/3=0.0167 level (the three pair-wise comparisons were performed at this significance level to ensure that the overall error rate among all three pairs was no higher than 0.05), only the postdissemination vs. post-decontamination comparison was determined to be significant at an overall 95% confidence level. Thus, the difference between the post-dissemination and the post-decontamination sampling times was the primary contributor to the overall differences among stages. No significant association was observed between the proportion of non-detected results and location (p=0.360).
Discussion
The BOTE project provided a unique opportunity to study migration of spores outside a building following spore dissemination and subsequent building decontamination. Sterile sand samples were placed outside the test building between the building wall and the tent covering the building in strategic locations near the building doorways to determine if spore migration outside the building was occurring. Detected results were noted in 36% of the samples (43/120) and detects were found most often near location 1, indicating the potential for spores to migrate outside the building. Samples were found to be detected significantly more often during the post-dissemination sampling stage compared to samples taken post-decontamination.
Although the results do indicate the potential for spore migration out the building, no statistical conclusions could be reached regarding the actual migration pathway of the B. atrophaeus subsp. globigii spores. One of the contributors to the lack of conclusions regarding the migration pathway in the current study may have been due to the exercise setup. For example, because a tent was placed over the test building to maintain negative pressure, infiltration and exfiltration effects may have been reduced, which in turn may reduce migration from the building. The tent could have interfered with the natural dissemination of spores to environmental areas, causing any escaped spores to become deposited between the exterior building walls and the interior tent walls. The tent also forced samplers to walk through tight spaces around the building, possibly reaerosolizing spores present in the native soils surrounding the building, which were present prior to the start of the exercise. In an actual event, a secondary barrier would not be in place during the release, and thus spores could be carried much greater distances than were studied here. In addition, a roof vent on the building was unintentionally left open (located on the top of the building near sample locations 9 and 10) during the pH-amended bleach phase of the exercise, but the roof vent was sealed just prior to chlorine dioxide decontamination. Spores could have escaped the building via this vent and could have contributed to the detected samples near these locations. During an actual event, any open door, window, or vent would be a point of exit for airborne spores. Unfortunately, there is no way to determine which of these factors may have played the most significant role in determining where detected samples might be found outside the building.
One possible theory of the transport of spores outside the building is disturbance of the spores by sampling and decontamination personnel or movement of the spores through the heating and cooling system. Weis et al. [14] demonstrated the potential for secondary aerosolization of B. anthracis spores from minimal movements, leading to a hypothesis that spores are transported out of the building by physical processes including people, air movement, or electrostatic forces leading to a decrease in spores in the building. However, the actual reason for the decrease in spore detection observed in the samples is still unknown. In a recent study conducted by Van Cuyk et al. [2], B. thuringiensis spores released outside a building migrated into near-by buildings; the highest concentrations were found near the entrances and the heating, ventilation and air conditioning (HVAC) filters. One of the largest outbreaks of inhalational anthrax occurred in 1979 and was attributed to HVAC exhaust filters not being in place while B. anthracis spores were being dried [15].
One factor to consider when drawing conclusions regarding the results of this study is that B. atrophaeus subsp. globigii spores had been used in the test area during previous events, and the normal background level of spores at the site was unknown. During this study, efforts were made to mitigate the influence from contaminated in-situ soil. The sterile sand samples were contained within large orange sampling trays to make personnel aware of their presence and to reduce activity near them. The sterility of the sand samples utilized was checked both before the exercise and with site blanks, which were opened briefly on-site within the contaminated environment. Regardless of these efforts, nothing could be done to prevent in-situ soil spores from being reaerosolized by personnel elsewhere within the tent. Even simple handling of sand dishes during placement and collection or other activities being undertaken during each round such as reset of the facility could have disturbed spores. In addition, carryover from one event to the next may have been possible and may account for some of the detected results during pre-dissemination sampling. The design of this study did not allow for distinguishing whether reaerosolization or carryover contributed more to detected samples. To study spore migration from inside a building to the outdoor soils fully, a controlled setup would be needed to specifically look at the different processes that might cause spores to reaerosolize, cause spores to move through entrances, exits, window seals and vents, and to study sample collection from a more natural soil setting.
Caution should be used when attributing the observed decrease in detected samples following decontamination to the decontamination technologies alone because none of the sand samples received direct decontamination. Decontamination overspray or vapors flowing out into the secondary-enclosure area may have caused the decrease in detected spores post-decontamination, and inhibition testing indicated that the decontamination agents did not interfere with PCR analysis. In a recent study [16], chlorine dioxide gas was investigated for efficacy of decontamination of both Arizona Test Dust (ATD) and topsoil inoculated with B. anthracis and B. subtilis spores at 1 cm and 2 cm depths. Results for topsoil indicated that none of the samples were completely decontaminated at the 1 cm depth and the efficacy of the treatment decreased with the 2 cm depth [16]. ATD was easier to decontaminate than the topsoil. In addition, the study found the B. anthracis log reduction after decontamination treatment to be higher with sterile soils compared to unsterilized soils [16]. Results from the current study could have presented different results if nonsterile sand or different amounts of sand in the dishes had been used.
This study utilized a new method for preparation of the sand samples prior to analysis. The matrix limit of detection for the method of extracting the spores from the sand (104 spores/g of sand) was found to be comparable to the results reported by Ryu et al. [17], who detected 104 CFU /g of B. anthracis spores in soil samples, and Herzog et al. [18], who found the mean matrix LOD for B. anthracis in spiked soils to be 1.2 × 104 CFU/g of soil. However, Kursar et al. [19] reported a much lower limit of detection in soil using the same DNA kit as the current study [19]. With a detection limit of 104 spores/g of sand, detection of the spore concentration disseminated on the first floor (target concentration 106 spores/ft2) of the building was possible but not on the second floor (target concentration 102 spores/ft2). The analysis did not attempt to determine spore viability; because analysis of the samples was qPCR based, it is not known whether the DNA that was detected in the samples came from viable spores or nonviable spores with intact DNA. Improvements to the extraction and analysis method that include a lower matrix limit of detection, additional standard curve data, and viability testing could greatly improve the overall results.
Conclusion
Following an actual release within a building, human activity and airflow can cause the agent of concern to be released into a wide area and potentially affect people outside the building. However, data from this study were not able to determine the quantity of spores that migrated out of the facility. Future studies could help determine the probability and extent of contamination during a decontamination event. Such studies should address the possibility of physical processes within a secondary enclosure that could decrease the presence of spores. In particular, if spores bind to surfaces such as the secondary-enclosure walls, the exterior of the secondary enclosure, or personnel clothing, other matrices of concern during an actual event could be implicated. The LOD for soil matrices is insufficiently sensitive for current research needs, and future research requires a more efficacious method for extracting DNA from soil samples in support of site characterization and clearance sampling. The data collected in this study provide insight on the extent of spore migration that is likely to occur during a buildingbased release-event. Collectively these data should provide valuable guidance for remediation efforts.
Disclaimer and Acknowledgments
The U.S. Environmental Protection Agency (EPA), through its Office of Research and Development, funded and managed the research described herein. Pegasus Technical Services, Inc., a contractor to the EPA (Contract # EP-C-11-006) performed laboratory analysis. Battelle/Chemical, Biological, Radiological, and Nuclear Defense Information and Analysis Center (Contract # SP0700-00-D-3180, Delivery Order 0679, Technical Area Task 886) provided statistical support. Geochemical testing was performed by Midwest Laboratory. The content of this paper has been peer and administratively reviewed and has been approved for publication. Note that approval does not signify that the contents necessarily reflect the views of the EPA. Reference herein to any specific commercial product, process, or service by trade name, trademark, manufacturer, or otherwise does not necessarily constitute or imply its endorsement, recommendation, or favoring by the United States government. The views and opinions expressed herein do not necessarily state or reflect those of the United States government and shall not be used for advertising or product endorsement purposes. We also acknowledge Brian Morris (Pegasus Technical Services) for his contributions to the analysis of the samples and Kathy Nickel for help in initial design of the study.
References
- Sextro RG, Lorenzetti DM, Sohn MD, Thatcher TL (2002) Modeling the spread of anthrax in buildings. Proceedings of the 9th International Conference of Indoor Air Quality and Climate, Monterey, CA, p 506-511.
- Van Cuyk S, Deshpande A, Hollander A, Franco DO, Teclemariam NP, et al. (2012) Transport of Bacillus thuringiensis var. kurstaki from an outdoor release into buildings: pathways of infiltration and a rapid method to identify contaminated buildings. Biosecur Bioterror 10: 215-227.
- Sinclair R, Boone SA, Greenberg D, Keim P, Gerba CP (2008) Persistence of category A select agents in the environment. Appl Environ Microbiol 74: 555-563.
- Lewis JC (1969) Dormancy. Hurst P, Gould GW (ed.) In, The bacterial spore. Academic Press, London, UK.
- Manchee RJ, Broster MG, Melling J, Henstridge RM, Stagg AJ (1981) Bacillus anthracis on Gruinard Island. Nature 294: 254-255.
- Turnbull PC (2008) Anthrax in Humans and Animals. 4th ed. World Health Organization, Geneva.
- DHS (2008) September 2008: Indoor field evaluation of sample collection methods and strategies at Idaho National Laboratory II. NSTD-09-01653. Department of Homeland Security and Joint Program Executive Office for Chemical and Biological Defense: Washington, D.C.
- Amidan BG, Pusipher BA, Matzke BD (2009) Statistical analyses of second indoor bio-release field evaluation study at Idaho National Laboratory. Pacific Northwest National Laboratory and U.S. Department of Energy. PNNL-18932.
- Brown GS1, Betty RG, Brockmann JE, Lucero DA, Souza CA, et al. (2007) Evaluation of a wipe surface sample method for collection of Bacillus spores from nonporous surfaces. Appl Environ Microbiol 73: 706-710.
- Williams G1, Linley E, Nicholas R, Baillie L (2013) The role of the exosporium in the environmental distribution of anthrax. J ApplMicrobiol 114: 396-403.
- Sandia National Laboratories (2006) BROOM (Building Restoration Operations Optimization Model), p. Sandia Copyright 737.731, Beta 1.0 ed. Sandia National Laboratories, Livermore.
- Kane SR1, Létant SE, Murphy GA, Alfaro TM, Krauter PW, et al. (2009) Rapid, high-throughput, culture-based PCR methods to analyze samples for viable spores of Bacillus anthracis and its surrogates. J Microbiol Methods 76: 278-284.
- Fleiss JL, Levin B, Paik MC (1981) Stastistical method for rates and proportions. John Wiley & Sons, Inc., New York, NY.
- Weis CP1, Intrepido AJ, Miller AK, Cowin PG, Durno MA, et al. (2002) Secondary aerosolization of viable Bacillus anthracis spores in a contaminated US Senate Office. JAMA 288: 2853-2858.
- Alibek K, Handelman S (1999) Biohazard: The Chilling True Story of the Largest Covert Biological Weapons Program in the World - Told from the Inside by the Man Who Ran It. Random House, New York, NY.
- Cowan GM (2012) EPA Inactivation of Bacillus anthracis Spores in Soil Matrices with Chlorine Dioxide Gas, U.S. Environment Protection Agency Office of Research and Development. EPA 600/R/12/517.
- Ryu C1, Lee K, Yoo C, Seong WK, Oh HB (2003) Sensitive and rapid quantitative detection of anthrax spores isolated from soil samples by real-time PCR. MicrobiolImmunol 47: 693-699.
- Herzog AB1, McLennan SD, Pandey AK, Gerba CP, Haas CN, et al. (2009) Implications of limits of detection of various methods for Bacillus anthracis in computing risks to human health.Appl Environ Microbiol 75: 6331-6339.
- Kursar D, Pate M, Hubad B, Avbersek J, Logar K, et al. (2012) Detection of Bacillus anthracis in the air, soil, and animal tissue. Acta Veterinaria 62: 77-89.
Relevant Topics
- Anthrax Bioterrorism
- Bio surveilliance
- Biodefense
- Biohazards
- Biological Preparedness
- Biological Warfare
- Biological weapons
- Biorisk
- Bioterrorism
- Bioterrorism Agents
- Biothreat Agents
- Disease surveillance
- Emerging infectious disease
- Epidemiology of Breast Cancer
- Information Security
- Mass Prophylaxis
- Nuclear Terrorism
- Probabilistic risk assessment
- United States biological defense program
- Vaccines
Recommended Journals
Article Tools
Article Usage
- Total views: 14369
- [From(publication date):
May-2015 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 9762
- PDF downloads : 4607