Research Article Open Access
Studies on the Mitochondrial Genomics in Salmo trutta caspius Population in Three Rivers of Caspian Sea
Rezaei A1*, Akhshabi S1 and Jamalzadeh HR21Department of Genetics–School of Basic Science, Islamic Azad University, Tonekabon Branch, Iran
2Department of Fisheries Sciences and Marine Biology-School of Basic Science, Tonekabon Branch, Islamic Azad University, Tonekabon, Iran
- *Corresponding Author:
- Rezaei A
Department of Genetics School of Basic Science
Islamic Azad University, Tonekabon Branch, Iran
Tel: +98 21 4486 5179
E-mail: a.rezaei@tonekaboniau.ac.ir
Received Date: November 24, 2016; Accepted Date: December 28, 2016; Published Date: January 13, 2017
Citation: Rezaei A, Akhshabi S, Jamalzadeh HR (2017) Studies on the Mitochondrial Genomics in Salmo trutta caspius Population in Three Rivers of Caspian Sea. J Fisheries Livest Prod 5: 213 doi: 10.4172/2332-2608.1000213
Copyright: © 2017 Rezaei A, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Fisheries & Livestock Production
Abstract
Mitochondrial DNA is suitable for phylogenetic studies. Hence in this study mitochondrial DNA genes in Salmo trutta caspius were sequenced and deposited in Genbank. Six sequences of S.t. caspius mitochondrial ATPase 6 gene with accession nos. LC011387.1, LC154842- LC154847 and the full length of the mitochondrial genome in S.t. caspius (accession no. LC011387.1) were deposited in Genbank. Sequences were aligned between S.t. caspius and Salmo trutta by BLAST program. Although results showed a high homology between both sequences, but single nucleotide variations between ND genes in S.t. caspius and Salmo trutta were observed. S.t. caspius, S. trutta, and Salmo salar sequences were converted into a circular map in an online system named Organellar Genome DRAW (http://ogdraw. mpimp-golm.mpg.de). Maps created by OGDRAW were different for S.t. caspius, S. trutta and S. salar. Moreover, Maximum Parsimony method was conducted for evolutionary analysis using MEGA 6.0 program tree between S.t. caspius and other salmonids. The results showed that S.t. caspius, Salmo trutta, Salmo trutta fario and S. trutta are covered in a group. On the other hand, results of evolutionary analysis using Tajima's test were conducted in MEGA 6.0 program for three sequencing including, S.t. caspius, S.t. trutta specimen voucher Nor 00, and S.t. fario used as an outgroup for equality of evolutionary rate. Comparison of partial mitochondrial sequence in Iranian S.t. caspius population was performed between three regions [Tonekabon (Cheshmekileh Roud), Ramsar (Safa Roud) and Talesh (Nav roud)]. The results showed that except for one single nucleotide mutation in the partial sequence of mitochondria of S.t. caspius [Ramsar (Safa Roud)], there has been 100% homology between them. In conclusion, the mitochondrial genome homology of S.t. caspius and other salmonids were high, though some SNPs were observed between S.t. caspius and other salmonid species.
Keywords
Salmo trutta caspius; Mitochondrial genomics; Salmo trutta; Phylogenetic studies
Introduction
The Caspian salmon, S.t. caspius, is an anadromous form and one of the endemic subspecies of fish in Caspian basin, living at the western and southern coasts of the Caspian Sea. S.t. caspius populations are at risk of extinction and it was listed as a threatened species by the IUCN Red List of Threatened Species. Moreover, S.t. caspius populations are important for livestock economics and aquaculture industry. Salmon is a popular food in Asia pacific region, Europe, and America and salmonid species are considered a main food in Asia [1]. Today’s evidence shows that S.t. caspius species are very rare in the world and are very sensitive to water pollution of seas and rivers. The adult female salmons also should travel to fresh water for spawning, so the status of phylogeny will also be affected by their traveling from sea to river for reproductive behaviour. Hence, S.t. caspius population is an important species for studies on molecular markers. Evolutionary history of Salmo taxa, such as brown trout, Salmo salar, and Salmo trutta populations, has been studied [2,3]. In addition, most of the populations examined were genetically highly divergent, indicating that they may represent distinct and potentially locally adapted gene pools [4]. Many molecular markers were used for evolution studies, such as microsatellites, RFLP (Restriction Fragments Length Polymorphism) and AFLP (Amplified Fragments Length Polymorphism), RAPD (Random Amplification of Polymorphic DNA), partial and full mitochondrial genome such as cytochrome b, NADH 1 and D-Loop genes [5] and phylogeography [6,7] for studying the effects of stocking, conservation and management of populations and species [8-11], and resolving taxonomy confusions [6,11-21]. mtDNA is necessary for phylogenetic studies because complete mitochondrial genome (mtDNA) is inherited for maternal traits whereas paternal traits are inherited by nucleus genome [5]. mtDNA is also inherited maternally without intermolecular recombination and it has a higher mutation rate [22], which is one of the important parameters for phylogenetic and phylogeographic research [6,7,11,23-26].
The aim of the current research was to study the phylogenetic of S.t. caspius in Iranian populations. According to Berg [25], Salmo trutta contains six subspecies: S.t. trutta L., Salmo trutta labrax Pallas, S.t. caspius Kessler, Salmo trutta oxianus Kessler, Salmo trutta aralensis Berg and Salmo trutta ezenami Berg [25]. Additionally, Kottelat & Freyhof [2] referred to different populations of Caspian trout as Salmo trutta (northern Caspian basin), Salmo ciscaucasicus Dorofeeva [27] western Caspian basin), and Salmo caspius Kessler 1877 (southern Caspian basin). Berg and other researchers only used phenotypic pattern and geographical studies, but here we will use both phenotypic and genotypic characteristics of salmonids and will compare them within and between salmonids.
This study tried to answer current questions, a) is there any variation in mtDNA genome of S.t. caspius in different regions of Iran? Is there any variation between mtDNA genome of S.t. caspius and other populations of Salmonids? How is the evolution of S.t. caspius and its relationship with other salmonids, drawn by phylogenetic tree methods?
Materials and Methods
Salmon samples
Samples of salmon were collected from Cheshame Kileh (city of Tonekabon), Safa roud (city of Ramsar), and Nav roud (city of Talesh), Iran. The samples of salmonids that were caught in fall of 2015 were three years old. Fish were anesthetized and after tagging, the left side was photographed and the right pectoral fin was clipped, tagged like the whole fish, and fixed in 96% ethanol.
DNA extraction, PCR amplification, and sequencing
Total genomic DNA was extracted from muscle tissue (10 mg) taken from fresh specimens. Tissue was digested by incubating with proteinase K/SDS solution at 37° for two hours. DNA extraction was carried out following phenol-chloroform and ethanol precipitation protocol. The quality of DNA extracted was checked on 1% agarose gel. Double-stranded PCR amplification were carried out in 50 μl reaction buffer including, 1x standard Taq buffer (10 mM Tris-HCl, 50 mM KCl, pH 8.3), 3.5 mM MgCl2, 200 μM dNTPs, 0.4 μM of each primer, 1 U Taq DNA polymerase (Fermentas, Inc.) and 2 μl DNA template (100 ng/μl). The primer pairs used included degenerate primers known to amplify mitochondrial DNA in fish species as well as primers constructed from Atlantic salmon (S. salar) sequence produced for this study (Milly Sin Yan So, 2001, Thesis). PCR amplification was performed as follows: 94° for 2 min followed by 40 cycles of 94° for 15 sec, 60° for 30 sec and a final extension at 70° for 10 min. 10 μl of the amplified PCR product was analyzed by electrophoresis on 1% agarose gel in TBE buffer, stained with ethidium bromide and visualized under UV light. Purified PCR products were sequenced with next-generation sequencing (Sequencing Facility, India).
Primer development
Thirty-three pairs of mitochondrial-specific primers were designed using a template generated from the consensus sequence of different complete salmonid mitochondrial sequences such as S. salar (NC_00861) and Salmo trutta (JQ390057). Conserved regions were selected to place overlapping forward and reverse primers. Primer design was carried out using Primer 3 online program.
Sequence analysis
10 ng of purified PCR products for every 100 bp of the PCR product and 1 μl of primer (10 pmol stock) were used for sequencing machine.
Gen-Bank deposition
Six sequences of S.t. caspius mitochondrial ATPase 6 gene deposited in Genbank with accession nos. LC154842- LC154847 and the full length of the mitochondrial genome of S.t. caspius was deposited in Genbank with accession no. LC011387.1
Bioinformatics programs for analysis
A multiple sequence alignment of fourteen salmon species’ mtDNA sequences was generated with MEGA program (Version 6.0) and NCBI Network system. The degree of divergence and the number of transition and transversion were also investigated. The location of sites that were variable amongst thirteen salmonid species sequences was recorded to study the relative rate of evolution at different regions of the mtDNA genome of salmon.
Results
Genomic DNA organization
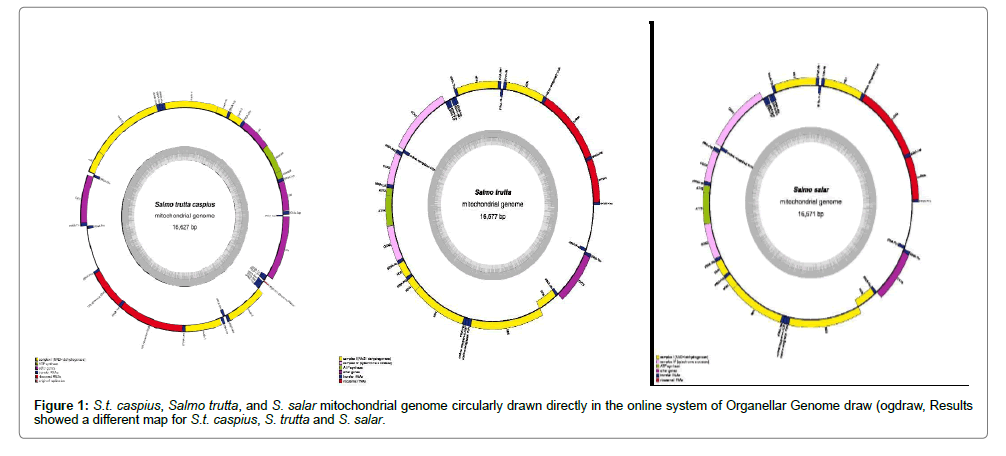
Figure 1 shows that few short spacer regions are present between some genes, but they are mostly one to two bp in size. The longest spacer is 14 bp located between ND1 and tRNA-isoleucine. The overlapping arrangements in three sets of genes, ATPase 8/ATPase6, ND4L/ND4 and ND5/ND6, were 10, 7 and 4 bp, respectively. Furthermore, other overlappings were found in the 16671-16674 bp region. The Salmo trutta mitochondrial genome sequenced is 16679 bp in length. Small single or double base pair insertions in D-Loop and 1 bp in the 16S ribosomal RNA gene demonstrated that the control region is the most flexible region to accept indel mutations.
Nucleotide compositions
The nucleotide composition of the coding strand in S.t. caspius mitochondrial genome illustrated a bias against the use of guanine nucleotide with the range of 14.1 to 22.5% for G, 14.5 to 31.6% for T, 22.3 to 35% for C and 24.1 to 36.9% for A (Table 1). The average distribution of the bases for Salmo trutta was 14.8 to 29.5% for T, 22.4 to 34.4% for C, 22.7 to 32.3 % for A, and 28.3% for G (Table 1).
| Salmotruttacaspius | Salmotrutta | Salmosalar | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide | T | C | A | G | T | C | A | G | T | C | A | G |
| D-loop | 31.6 | 22.3 | 31.2 | 14.1 | 31.4 | 22.4 | 31.7 | 14.4 | 31.7 | 22.2 | 31.5 | 14.7 |
| 12SrRNA | 19.2 | 27.8 | 29.7 | 22.6 | 19.5 | 27.3 | 29.5 | 22.3 | 19.8 | 27.9 | 30.5 | 22.5 |
| 16SrRNA | 19.8 | 25.6 | 32.5 | 22.5 | 19.4 | 25.4 | 32.3 | 22.4 | 19.5 | 25.8 | 32.6 | 22.3 |
| ND1 | 26.8 | 31.2 | 26.9 | 15.4 | 26.8 | 32.4 | 26.9 | 14.3 | 27.5 | 31.6 | 25.9 | 15.3 |
| ND2 | 25.1 | 34.2 | 26.9 | 13.9 | 25.6 | 35.3 | 26.3 | 13.6 | 26.5 | 33.7 | 28.4 | 11 |
| ND3 | 26.9 | 33.0 | 24.1 | 16.0 | 27.4 | 34.3 | 22.7 | 15.3 | 27.5 | 34.6 | 23.4 | 14.8 |
| ND4 | 27.7 | 30.6 | 27.2 | 14.0 | 27.4 | 30.2 | 27.6 | 13.3 | 28.4 | 30.5 | 27.7 | 14.8 |
| ND4L | 24.2 | 34.3 | 24.9 | 16.5 | 24.6 | 34.3 | 24.8 | 16.3 | 26.6 | 32.4 | 24.7 | 16.8 |
| ND5 | 27.5 | 30.6 | 28.4 | 13.6 | 27.5 | 30.4 | 28.3 | 13.7 | 27.7 | 30.6 | 29.8 | 12.7 |
| ND6 | 14.5 | 35.0 | 36.9 | 13.6 | 14.8 | 34.5 | 37.3 | 13.7 | 15.6 | 36.7 | 38.6 | 12.9 |
| COX1 | 28.1 | 28.1 | 27.8 | 16.1 | 19.5 | 27.3 | 29.5 | 22.3 | 30.5 | 27.5 | 24.8 | 17.4 |
| COX2 | 29.1 | 31.5 | 24.0 | 15.3 | 29.5 | 28.7 | 24.6 | 13.3 | 28.5 | 27.8 | 27.7 | 16.5 |
| COX3 | 29.5 | 29.1 | 24.6 | 16.3 | 29.4 | 29.5 | 24.3 | 16.9 | 28.8 | 29.6 | 26.3 | 15.7 |
| ATPase8 | 25.0 | 32.7 | 29.8 | 12.5 | 25.4 | 32.4 | 29.8 | 12.3 | 26.8 | 31.7 | 29.8 | 12.6 |
| ATPase6 | 27.5 | 34.5 | 25.1 | 12.5 | 28.4 | 34.4 | 25.5 | 12.8 | 28.4 | 33.5 | 26.8 | 11.9 |
| Cytb | 29.2 | 31.5 | 24.5 | 15.7 | 29.3 | 31.3 | 24.7 | 15.4 | 29.6 | 30.8 | 24.0 | 15.9 |
Table 1: The nucleotide composition of the coding strand of mitochondrial genome compared between S.t. caspius, S. trutta, and S. salar illustrated a bias against the use of four DNA nucleotides (A-T-C-G) with the range of 14.1 to 36.9.
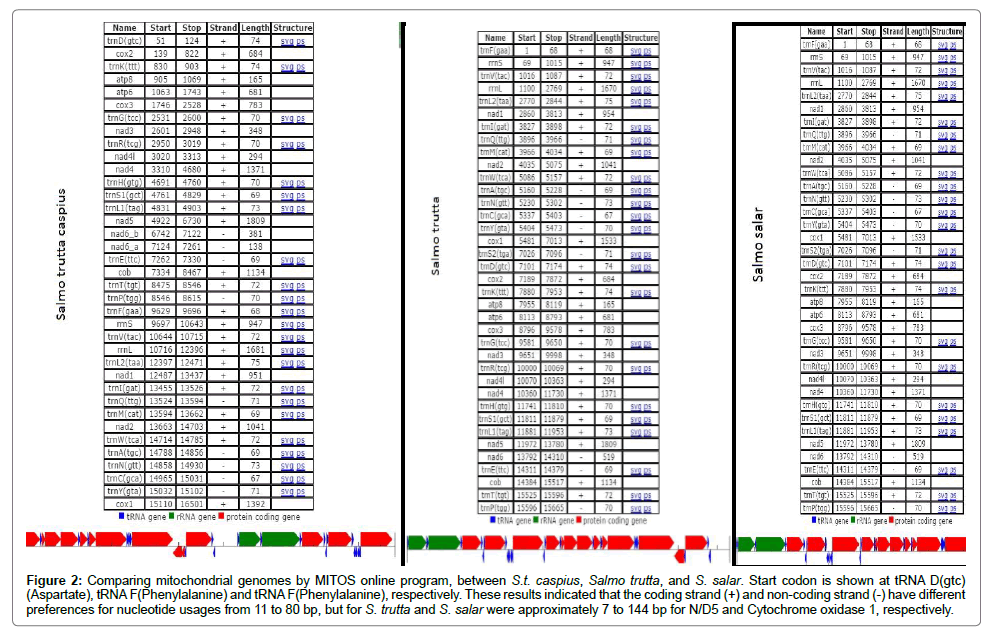
The Salmo mitochondrial genome contains one non-coding control region (D-Loop), two genes for ribosomal RNA, 22 genes encoding tRNAs and thirteen protein-coding genes (Figure 2). The encoded proteins include seven subunits of the NADH dehydrogenase, cytochrome b, cytochrome c oxidase complex (COX), ATPase 6 and 8 in three species, S.t. caspius, Salmo trutta and S. salar and were directly turned into a circular map in an online system named Organellar Genome DRAW (OGDRAW). OGDRAW is a specialized tool to convert genetic information stored in GenBank entries to graphical maps. This application is specifically optimized and adapted for the construction of high-quality, complete circular maps of organellar genomes like the mitochondrial genome. The maps drawn were different for S.t. caspius, S. trutta and S. salar (Figure 2).
Figure 2: Comparing mitochondrial genomes by MITOS online program, between S.t. caspius, Salmo trutta, and S. salar. Start codon is shown at tRNA D(gtc) (Aspartate), tRNA F(Phenylalanine) and tRNA F(Phenylalanine), respectively. These results indicated that the coding strand (+) and non-coding strand (-) have different preferences for nucleotide usages from 11 to 80 bp, but for S. trutta and S. salar were approximately 7 to 144 bp for N/D5 and Cytochrome oxidase 1, respectively.
In Figure 3 shows the comparison of mitochondrial genomes created by the online program, MITOS between S.t. caspius, S. trutta and S. salar. Start codons were at tRNA D(gtc)(Aspartate), tRNA F(Phenylalanine) and tRNA F (Phenylalanine), respectively. These results indicate that coding strand (+) and non-coding strand (-) were variable from 11 to 80 bp; but in case of S. trutta and S. salar, it was approximately from 7 to 144 bp for N/D5 and Cytochrome oxidase 1, respectively.
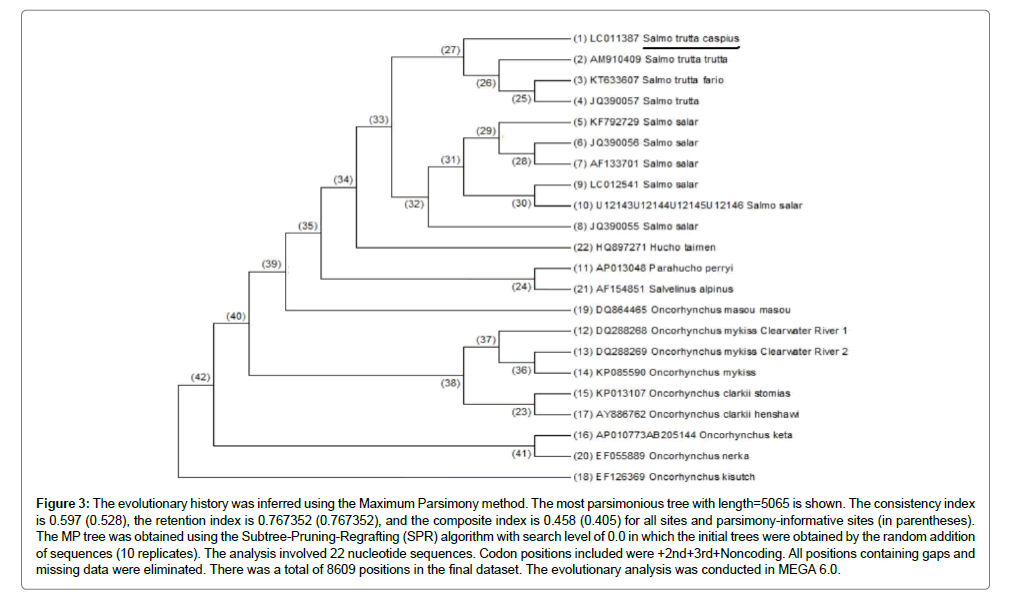
Figure 3: The evolutionary history was inferred using the Maximum Parsimony method. The most parsimonious tree with length=5065 is shown. The consistency index is 0.597 (0.528), the retention index is 0.767352 (0.767352), and the composite index is 0.458 (0.405) for all sites and parsimony-informative sites (in parentheses). The MP tree was obtained using the Subtree-Pruning-Regrafting (SPR) algorithm with search level of 0.0 in which the initial trees were obtained by the random addition of sequences (10 replicates). The analysis involved 22 nucleotide sequences. Codon positions included were +2nd+3rd+Noncoding. All positions containing gaps and missing data were eliminated. There was a total of 8609 positions in the final dataset. The evolutionary analysis was conducted in MEGA 6.0.
On the other hand, results showed that the mitochondrial genome is a closed circle, so there really is no "correct" beginning or ending, but by convention, most of the entries for fish mitochondrial genome begin near the tRNA-Phenylalanine gene just before the tRNA-Val and tRNA-Leu genes. In S.t. caspius, the sequence begins near the tRNAAsp gene just before the Cytochrome Oxidase subunit II gene (Figure 3).
Evolution of S.t. caspius and other salmonids
In Figure 3 shows Maximum Parsimony method for evolutionary analysis of tree conducted by MEGA 6.0 program between S.t. caspius and other salmonids. The most parsimonious tree with the length of 5065 is shown. This tree is drawn as a phylogram, the length of a given branch is scaled to the amount of character change. All species were recovered in 100% of the bootstrap replicates. Strict consensus of four trees is resulting from maximum parsimony analysis of 25 salmonid species (right). The results showed that S.t. caspius, S.t. trutta, S.t. fario and S. trutta are covered in a group (Evolutionary analysis was conducted in MEGA 6.0).
On the other hand, results for Tajima’s test were conducted in MEGA 6.0 program for three sequencing including, S.t. caspius mitochondrial DNA complete genome, S.t. trutta complete mitochondrial genome specimen voucher Nor 00, and S.t. fario mitochondrial complete genome was used as an outgroup for equality of evolutionary rate. The χ2 test statistic was 12339.01 (P=0.00; df=2; p<0.05). A p-value less than 0.05 is often used to reject the null hypothesis of equal rates between lineages. The analysis involved 3 nucleotide sequences. Codon positions included were 1st+2nd+3rd+Noncoding. All positions containing gaps and missing data were eliminated. There was a total of 16626 positions in the final dataset (Figure 4).
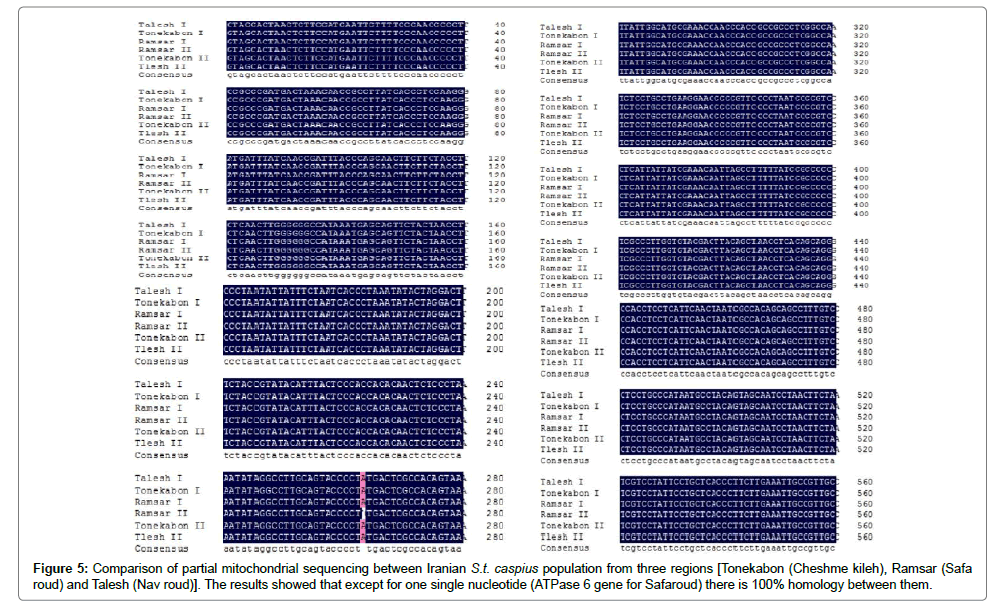
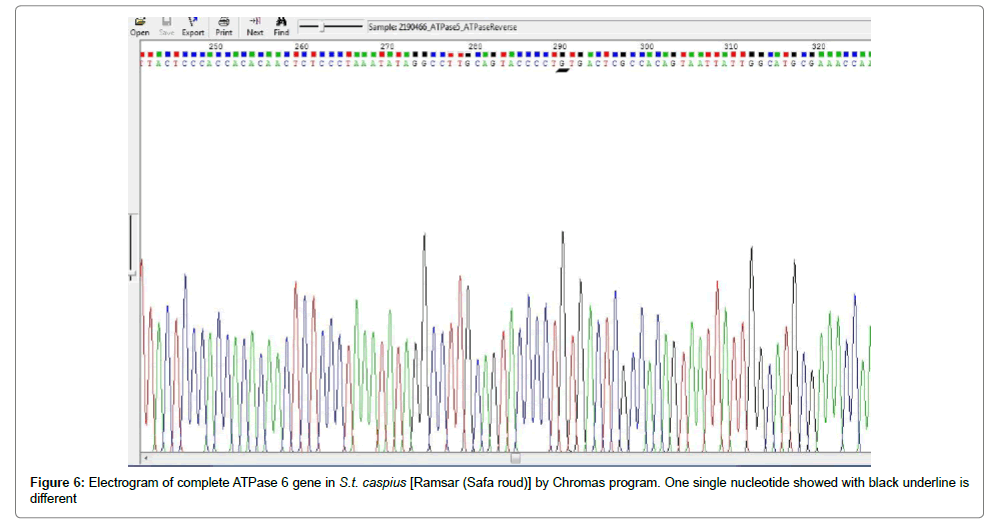
In Figure 5 shows the comparison of partial mitochondrial sequencing in Iranian S.t. caspius population between three regions [Tonekabon (Cheshme kileh), Ramsar (Safa Roud) and Talesh (Nav roud)]. The results showed that except for one single nucleotide mutation in ATPase 6 gene in S.t. caspius caught from Safa roud, there is 100% homology between them. Moreover, the result of electrogram by Chromas program (Version 2.40) showed one single nucleotide mutation which is underlined with black (Figure 6).
Figure 5: Comparison of partial mitochondrial sequencing between Iranian S.t. caspius population from three regions [Tonekabon (Cheshme kileh), Ramsar (Safa roud) and Talesh (Nav roud)]. The results showed that except for one single nucleotide (ATPase 6 gene for Safaroud) there is 100% homology between them.
Discussion
Why we compared S.t. caspius with S. trutta in present study?
S.t. caspius is the endemic of Caspian Sea and it cannot be found in other areas. According to Berg’s theory, S.t. caspius and Atlantic salmon (S. salar) can be originated from S. trutta because S. trutta migrated from the Atlantic Ocean to the White Sea and then derived to S.t. caspius in the Caspian Sea [25,26]. Here by our analysis, the evolutionary path of S.t caspius were confirmed in Figure 5. The substitutions per site between sequences were studied between S.t. caspius and other salmonids by Maximum Composite Likelihood model. Although the rate of substitutions between S.t. caspius, S.t. trutta, S.t. fario and S. trutta were very low (0.0); but we also observed a high rate of substituents for Hucho taimen.
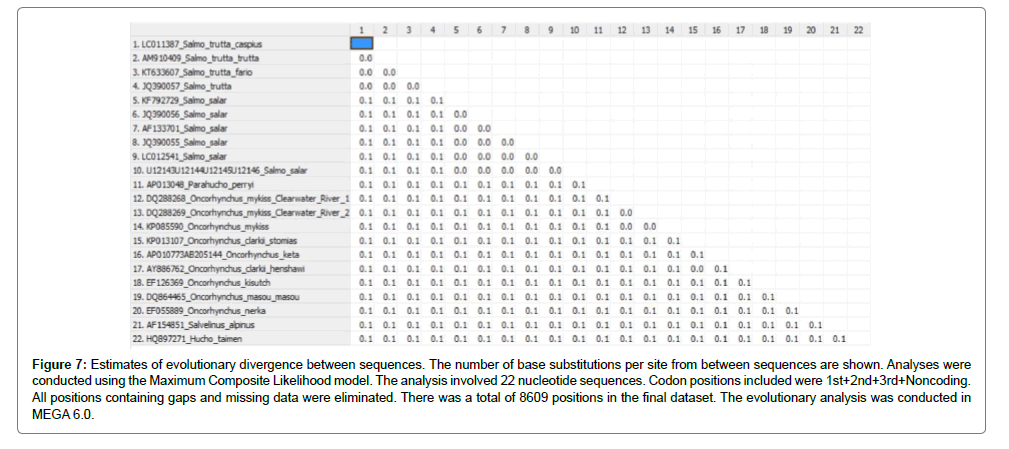
In addition, the status of evolution based on mitochondrial genomics could be explained by S.t. caspius > S. trutta > S. salar > Onchorhynchus mykiss > Hucho taimen, approximately (Figure 7).
Figure 7: Estimates of evolutionary divergence between sequences. The number of base substitutions per site from between sequences are shown. Analyses were conducted using the Maximum Composite Likelihood model. The analysis involved 22 nucleotide sequences. Codon positions included were 1st+2nd+3rd+Noncoding. All positions containing gaps and missing data were eliminated. There was a total of 8609 positions in the final dataset. The evolutionary analysis was conducted in MEGA 6.0.
S.t. caspius population is native of Caspian Sea and have been mentioned on the Red List as a threatened species. This is a very rare anadromous subspecies in the Caspian Sea territory. The Caspian lake trout reaches 50 kg and lives up to 10 years. This lake trout once spawned in the tributaries of Volga and Ural Rivers is considered to be a subspecies of the migratory Kumzha lake trout. This fish has not been seen in the wild for more than 15 years. It was once more widely distributed in the southwestern Caspian Basin. Mitochondrial genome was selected for this study, because at least in mammals, it is evolving more rapidly than nuclear genome [28]. Mitochondrial genome was used for phylogenetic analysis in three brown trout from the Oden State Fish Hatchery, the Muskegon, and Rogue Rivers, and Lake Michigan by using PCR-RFLP techniques which revealed two haplotypes within the Gilchrist strain, two within the Wild Rose strain, and reproduced with permission of the copyright owner [29]. An additional haplotype was found in three Lake Michigan trout which were determined to be Seeforellen based on Michigan DNR stocking records [29]. The current study also confirmed Berg's theory. Our results were analyzed by different options of MEGA software like Maximum Parsimony analysis of taxa and Pairwise Distance. The results showed a high homology between S.t. caspius, S.t. trutta, S.t. fario and S. trutta. It confirmed a suitable relationship between S.t. caspius and other salmonids that cited here. The results are shown in Figures 3-5. Further analysis of the MEGA gene showed sequence similarity at mtDNA genome between salmonid species. A phylogenetic tree containing S.t. caspius and other salmonids like S.t. trutta, S.t. fario, S. trutta, S. salar and etc. was constructed to further investigate their genetic relatedness (Figure 5). This tree grouped all salmonids in accordance with their conventional phylogeny; the genus Salmonid formed one branch (S.t. caspius, S.t. trutta, S.t. fario and S. trutta) and two genus of Salmo (S. salar; different accession nos.) and oncorhynchus (different accession nos.) formed another branch (Figure 3).
The low variability recorded between S.t. caspius (within a population of S.t. caspius) may be the result acquired from substantial interpopulation diversity in this species (Figure 3). For example, S.t. caspius may migrate from salt water (Caspian Sea) to spawn in fresh water (the rivers connected to it). Moreover genetic variability between the populations of salmonids like S.t. caspius, S.t. trutta, S.t. fario and O. mykiss found in the current study may be related to life history characteristics of the particular populations that are connected geographically. Similar studies have done by Bernatchez [4], Wills [30], Tiano [29] and Apostolidis, et al. [31] on the brown trout populations by partial mitochondrial genomic such as ND1, ND5/6 genes. They proposed that brown trout populations had limited spawning and nursery conditions in their environment compared to other populations which could lead to low recruitment. Atypical recommitment resulting in low effective population sizes or repeated bottlenecks could lead to a loss of genetic variability. However, isolation combined with a population bottleneck and genetic drift could also result in significant genetic differentiation among brown trout populations. In current study a variety between S.t. caspius, O. mykiss, and S. salar populations were found that are related to spawning and life history (Figures 3-5). Furthermore, phylogenetic trees would be expected to display conservation as it is a protein coding region by mitochondrial genes group that revealed by Nilsson, et al. [32], Cornell and Ward [33]. Relatively constant rates of mutation were observed among different salmonid species by these genes too [34].
The amount of sequence variation between S.t. caspius and other salmonids
In this study, we selected full length of mitochondrial genomics in S.t. caspius for making it possible to examine the relationship among S.t caspius, S.t. trutta, S. salar, and S.t. fario. For getting good quality, the specimens were analyzed by MEGA gene program (Version 6.0). In this study, twenty samples were selected from S.t. caspius. After sequencing the mitochondrial genome, they were analyzed within and between sequences, and no variation between them were found, so one sequence from examined samples was selected for comparing the populations.
Mitochondrial genomics is widely accepted as an essential tool for determining relationships among closely related species [35]. Furthermore here the high rate of nucleotide substitution between S. trutta and S.t. caspius is displayed in Figures 1 and 2 which show the graphical mapping of mitochondrial genomics in S.t. caspius and S. trutta. Results showed that mitochondrial genome in its circular form started in tRNA D(gtc)(Aspartate) and cytochrome 3 gene in S.t. caspius, but for S. trutta and S. salar it starts from tRNA F(Phenylalanine) and 12 srRNA gene.
These results are very exciting because homology was high but the results of mitochondrial genomics were different between them. Substitution of nucleotide was also compared between S.t. caspius, S. trutta and S. salar. Results showed that nucleotide variation (A-T-C-G) was 14.9 to 36.9 between them, however, approximately 98% homology observed between them (Table 2).
| Configuration | Count |
|---|---|
| Identical Sites in all three sequences | 4251 |
| Divergent Sites in all three sequences | 11 |
| Unique differences in sequence A | 12351 |
| Unique differences in sequence B | 4 |
| Unique differences in sequence C | 9 |
Table 2: The equality of evolutionary rate between sequences A (S.t. caspius mitochondrial DNA complete genome) and B (S.t. trutta complete mitochondrial genome specimen voucher nor 00), with sequence C (S.t. fario mitochondrion complete genome) used as an outgroup in Tajima's relative rate test.
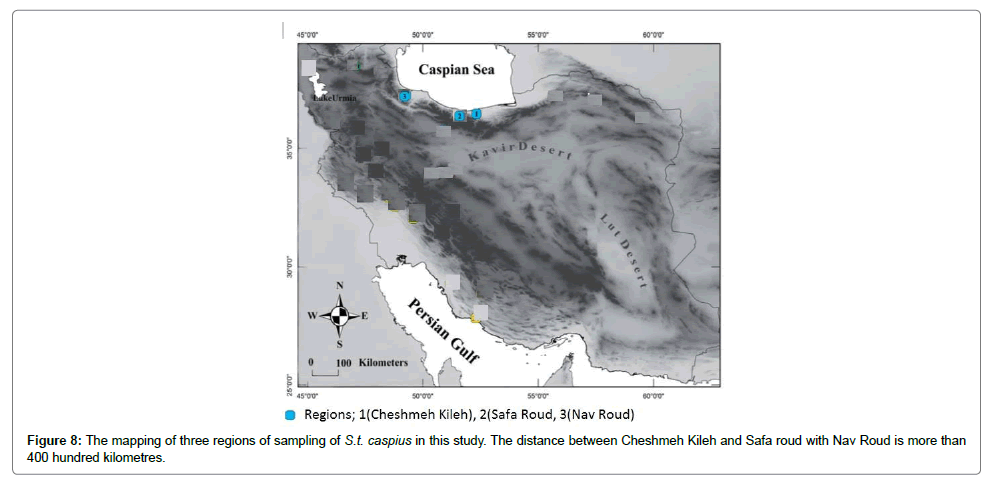
In Figures 5 and 6 compare S.t. caspius populations in three regions of Iran for phylogenetic status and our results showed that except for one single nucleotide mutation, there is a high homology between them. Three inland Rivers (Cheshme Kileh in Tonekabon, Safa roud in Ramsar and Nav roud in Talesh) populations are isolated by geographical chains stretching from the south (Tonekabon) to the south-west of rivers connected to Caspian Sea in Iran (Talesh) (Figure 8). These lakes are isolated from the southern Caspian basin in the northern part of the country, and their resident and migratory forms of S. trutta have been reported by Abdoli [28]. S.t. caspius usually inhabits the Cheshme Kileh in Tonekabon, Safa roud and Nav roud Lake basin, however to date it is unknown why this habitat is suitable for living of salmons and it may be related to a series of changes in behavioural, physiological and morphological parameters for migration to river from Sea water. The distance between rivers of three regions that were selected for this study are different; the distance between Cheshme Kileh and Safa Roud is around forty kilometres while between Cheshme kileh and Safa roud with Nav roud it is about 400 km. However, according to our study, distance is not the reason for differentiation in S.t. caspius. This is because the phylogenetic status of S.t. caspius in several regions including populations inhabiting Iranian inland basins and the Persian Gulf basin (Turkey) shows a good similarity with North African populations that was previously unknown [15].
In Figure 6 found one single nucleotide mutation in S.t. caspius originated from Safa roud (Ramsar) while showed high homology (100%) between two other origins of S.t. caspius (Nav roud and Cheshme Kileh). Segherloo used the complete mtDNA control region to compare the Iranian populations to one another; whereas for comparing the results for these populations with other published haplotypes, c. 995 bp of this region was used. A total of 129 specimens from six populations were used in sequencing, and seven haplotypes including HKa (Karaj River), HH1, HH2 (Haraz River), HBa (Babolrud River), HCo (the most frequent haplotype in all the studied rivers except for Karaj River), HM1 and HM2 (Mardagh River) have shown genetic diversity between them, while in this study we didn’t found any diversity.
In conclusion, mitochondrial genomics is suitable for phylogenetic studies, but for getting perfect results we should be using nuclear genomics. When nuclear DNA is used, the results show very clearly the different basins while mtDNA is not capable of that. Apostolidis, et al. [36] found no mtDNA diversity between populations that showed the highest values of nuclear heterozygosity amongst them. mtDNA is transmitted by females (maternal traits) while nuclear DNA is inherited from both female and male (parental traits), so here we have restricted our research to mtDNA but nuclear DNA is widely used for diversity studies. Our results showed no diversity between Iranian populations while Segherloo et al. found diversity [37-45].
On the other hand, Berg [25,26], have reported that salmonid species such as S.t. caspius are originated from S. trutta, and in fact S. trutta has branched to S.t. caspius and S.t. fario in the Caspian Sea. Here we investigated to see if that theory is right and S.t. caspius may be originated from S. trutta. Interestingly, because Berg's theory just confirmed geographical and phenotypical salmonids but here we used mitochondrial genome for studying the phylogenetic of salmonids. The rate of accuracy with mtDNA is more than phenotypical studies. Also, our studies confirmed Berg’s theories [25,26].
Acknowledgments
This work was financially supported by the Research Council of Islamic Azad University Tonekabon Branch, Iran.
References
- William SD, Ben FK, Steven JMJ, Patricia I, Rodrigo V, et al. (2010) Sequencing the genome of the Atlantic salmon (Salmo salar). Gen Bio 11: 403.
- Kottelat M, Freyhof J (2007) Handbook of European freshwater fishes. Publications Kottelat, Cornol, Switzerland p: 646.
- Simonovi�? P, Mari�? S, Nikoli�? V (2007) Trout Salmo spp. complex in Serbia and adjacent regions of the western Balkans: reconstruction of evolutionary history from external morphology. J F Biology 70: 359-380.
- Apostolidis AP, Stoumboudi MTH, Kalogianni E, Cote G, Bernatchez L (2011) Genetic divergence among native trout Salmo trutta populations from southern Balkans based on mitochondrial DNA and microsatellite variation. J Fish Biology 79: 1950-1960.
- Zhang DX, Hewitt GM (2003) Nuclear DNA analyses in genetic studies of populations: practice, problems and prospects. Mol Ecology 12: 563-584.
- Bernatchez L (2001) The evolutionary history of brown trout (Salmo trutta L.) inferred from phylogeographic, nested clade, and mismatch analyses of mitochondrial DNA variation. Evol 55: 351-379.
- Corteys M, García-Marín JL (2002) Evidence for phylogeographically informative sequence variation in the mitochondrial control region of Atlantic brown trout. J Fish Biol 60: 1058-1063.
- Apostolidis AP, Madeira MJ, Hansen MM, Machordom A (2008) Genetic structure and demographic history of brown trout (Salmo trutta L.) populations from the southern Balkans. Fre Biology 53: 1555-1566.
- Vera M, Sanz N, Hansen MM, Almodovar A, GarciaMarin JL (2010b) Population and family structure of brown trout, Salmo trutta, in a Mediterranean stream. M Freshwater Research 61: 676-685.
- Kohout J (2012) Effects of stocking on the genetic structure of brown trout, Salmo trutta, in Central Europe inferred from mitochondrial and nuclear DNA markers. TJ FA Sciences 19: 252-263.
- Kohout J, Šedivá A, Apostolou A, Stefanov T, Mari�? S, et al. (2013) Genetic diversity and phylogenetic origin of brown trout Salmo trutta populations in eastern Balkans. BS Zoology 68:1229-1237.
- Mrdak D, Nikoli�? V, Toši�? A, Simonovi�? PM (2012) Molecular and ecological features of the soft-muzzled trout Salmo obtusirostris (Heckel, 1852) in the Zeta river, Montenegro. Bio Sec Zoology 67: 222-233.
- Lee CE (2000) Global phylogeography of a cryptic copepod species complex and reproductive isolation between genetically proximate populations. Evol 54: 2014-2027.
- McGlashan DJ, Hughes JM (2000) Reconciling patterns of genetic variation with stream structure, earth history and biology in the Australian freshwater fish Crate rocephalus stercusmuscarum (Atherinidae). Mol Ecol 9: 1737-1751.
- Jarman SN, Elliott NG, Nicol S, McMinn A (2002) Genetic differentiation in the Antarcticcoastal krill Euphausia crystallorophias. Her 882: 80-28.
- Weiss S, Maria S, Snoj A (2011) Regional structure despite limited mtDNA sequence diversity found in the endangered Huchen, Huchohucho Hyd 658: 103-110.
- Rezaei A, Akhshabi SH (2011) Evolutionary Genetic Analysis of the cytochrome b gene variation in the Salmo trutta fario with other Salmons. Egypt Acad J Biolog Sci 3: 65-71.
- Rezaei A, Akhshabi SH, Jamalzadeh HR (2011) Sequencing and sequence analysis of Growth Hormone type 1 (GH1) gene homologue in the s.t.caspius. Adv S Biol 3: 111-122.
- Rezaei A. (2012) Studies of Cytochrome b nucleotide sequence variation in the Salmo trutta fario. Egypt Acad J Bio Sci 3: 125-131.
- Rezaei A, Akhshabi SH, Jamalzadeh HR (2012) The relation between growth hormone (GH) gene and Cytochrome b gene in three salmon types. Egyp Acad J Biolog Sci 3: 133-140.
- Rezaei A, Akhshabi SH (2012) Studies of cytochrome b protein modelling sequence in the Salmo trutta fario and comparing with other Salmonids. Egypt Acad J Biolog Sci 4: 1-8
- Avise JC (2000) Phylogeography: the History and Formation of Species. Harvard University Press, Cambridge.
- Mari�? S, Sušnik S, Simonovi�? P, Snoj A (2006) Phylogeographic study of brown trout from Serbia, based on mitochondrial DNA control region analysis. G Sel Evolution 38: 411-430.
- Vera M, Cortey M, Sanz N, Garcia-Marin JL (2010a) Maintenance of an endemic lineage of brown trout (Salmo trutta) within the Duero river basin. J Zool Syst Evol Res 48: 181-187
- Berg LS (1948) Freshwater fishes of the USSR and adjacent countries. Zoological Institute Akademy Nauk Moscow, USSR.
- Berg WJ, Ferris SD (1984) Restriction endonuclease analysis of Salmonid mitochondrial DNA. Can J Fish Aquat Sci 41:1041-1047.
- Dorofeeva EA (1967) Comparative morphological principles of taxonomy of East European salmons. Voprosy Ikhtiologii 7: 3-17.
- Anderson SMHL, Debruijn AR, Coulson IC, Eperon F, Sanger, et al. (1982) Complete sequence of bovine mitochondrial DNA: Conserved features of the mammalian mitochondrial genome. J Mol Biol 156: 683-717.
- Tiano T, Willis CM, Noble AA, Luttenton MR, Nikitin AG (2005) Genetic identification of hatchery stocks of brown trout (Salmo trutta) using mitochondrial DNA polymorphism. Master’s thesis. Grand Valley State University
- Wills TC (2006) Comparative abundance, survival, and growth of one wild and two domestic brown trout strains stocked in Michigan rivers. North American J Fisheries Management 26: 535-544.
- Apostolidis AP, Triantaphyllidis C, Kouvatsi A, Economidis PS (1997) Mitochondrial DNA sequence variation and phylogeography among Salmo trutta L. (Greek brown trout) populations. Mol Ecology 69: 531-542.
- Nilsson J, Gross R, Asplund T, Dove O, Jansson H, et al. (2002) Matrilinear phylogeography of Atlantic salmon (Salmo salar L.) in Europe and postglacial colonization of the Baltic Sea area. Mol Eco pp: 1089-102.
- Cornell PS, Ward RH (2000) Mitochondrial genes and mammalian phylogenies: increasing the reliability of branch length estimation. S Mol Bio Evol 17: 224-234.
- Doiron S, Bernatchez L, Blier PU (2002) A comparative mitogenomic analysis of the potential adaptive value of Arctic Charr mtDNA introgression in Brook Charr populations. S Mol Bio Evol 19: 1902-1909.
- Dorit RL (1986) Molecular and morphological variation in Lake Victoria Haplochromine cichlids (Perciformes: Cichlidae). Harvard University, Boston, MA.
- Apostolidis AP, Karakousis Y, Triantaphyllidis C (1996b) Genetic differentiation and phylogenetic relationships among Greek Salmo trutta L. (brown trout) populations as revealed by RFLP analysis of PCR amplified mitochondrial DNA segments. Her 77: 608-618.
- Abdoli A (2000) The inland water fishes of Iran. Iranian museum of nature and wildlife: 378.
- Bernatchez L, Guyomard R, Bonhomme F (1992) DNA sequence variation of the mitochondrial control region among morphologically and geographically remote European brown trout Salmo trutta populations. J Mol Ecology 1:161-173.
- Elliot JM (1994) Quantitative Ecology and the Brown trout. Oxford University Press p: 286.
- Rezaei A (2015a) Phylogenetic analysis of NADH dehydrogenase subunit 1(NADH1) gene in Salmo trutta caspius. J Ent Zoo Stu 3: 185-193.
- Rezaei A (2015b) Studies on protein structure of cytochrome c oxidase subunit II gene in s.t.caspius. J Adv Zool 36: 1-10.
- Hashemzadeh SI, Farahmand H, Abdoli A, Bernatchez L, Primmer CR, et al. (2012) Phylogenetic status of brown trout Salmo trutta populations in five rivers from the southern Caspian Sea and two inland lake basins, Iran: a morphogenetic approach. J F Biology 81: 1479-1500.
- IUCN (2012) IUCN Red List of Threatened Species. Version 2012.1. IUCN 2012. IUCN Red List of Threatened Species. Downloaded in June 2012.
- Tamura K, Stecher G, Peterson D, Filipski A, Kumar S, et al. (2013) Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30: 2725-2729.
- Bernatchez L, Osinov A (1995) Genetic diversity of trout (genus salmo) from its most eastern native range based on mitochondrial DNA and nuclear gene variation. Mol Ecol 4: 285-297.
Relevant Topics
- Acoustic Survey
- Animal Husbandry
- Aquaculture Developement
- Bioacoustics
- Biological Diversity
- Dropline
- Fisheries
- Fisheries Management
- Fishing Vessel
- Gillnet
- Jigging
- Livestock Nutrition
- Livestock Production
- Marine
- Marine Fish
- Maritime Policy
- Pelagic Fish
- Poultry
- Sustainable fishery
- Sustainable Fishing
- Trawling
Recommended Journals
Article Tools
Article Usage
- Total views: 4334
- [From(publication date):
March-2017 - Aug 31, 2025] - Breakdown by view type
- HTML page views : 3342
- PDF downloads : 992