Research Article Open Access
Common Pitfalls and Novel Opportunities for Predicting Variant Pathogenicity
| Tom van den Bergh1,2*, Bas Vroling2, Remko KP Kuipers1,3, Henk-Jan Joosten1,2 and Gert Vriend3 | |
| 1Laboratory of Microbiology, Wageningen University, Wageningen, The Netherlands | |
| 2Bio-Prodict, Nijmegen, The Netherlands | |
| 3CMBI, Radboud University Medical Centre, Nijmegen, The Netherlands | |
| Corresponding Author : | Tom van den Bergh Laboratory of Microbiology, Wageningen University Wageningen, The Netherlands and Bio-Prodict Nijmegen, The Netherlands Tel: 0031 24 845 7988 E-mail: tvandenbergh@bio-prodict.nl |
| Received January 05, 2016; Accepted January 27, 2016; Published February 03, 2016 | |
| Citation: van den Bergh T, Vroling B, Kuipers RKP, Joosten HJ, Vriend G (2016) Common Pitfalls and Novel Opportunities for Predicting Variant Pathogenicity. Biochem Physiol 5:197. doi:10.4172/2168-9652.1000197 | |
| Copyright: © 2016 van den Bergh T, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Visit for more related articles at Biochemistry & Physiology: Open Access
Abstract
The prediction of missense variant pathogenicity is normally performed using analyses of multiple sequence alignments optionally augmented with analyses of the (predicted) protein structure. The most straightforward way, though, is to search the literature to see whether this variant has already been described. Variant data from homologous proteins are also valuable because mutations in a homologous protein often have similar effects as mutations at the equivalent residues of the protein of interest. Transferring variant data seems trivial but is seriously hampered by the fact that homologous residue positions have different numbers in different species. This problem is even bigger when to proteins have such low sequence identities that they can no longer be aligned based on their sequences only and their structures need to be compared to align them accurately. The protein superfamily analysis software suite 3DM solves these problems, because 3DM is a system that combines high quality structure based multiple sequence alignments in which aligned residues have the same number, with all published mutant and variant data for human and all other species. We have used 3DM to analyze nine human proteins for which many disease-related variants are known. This study reveals that mutation data can be transferred even between very distant homologous proteins. Thus, protein superfamily information systems, such as 3DM, offer a wealth of unused information that can be used in the analysis of human variants.
| Keywords |
| DNA diagnostics; Fabry; Long QT syndrome; Protein superfamily; 3DM |
| Introduction |
| Rapidly evolving gene sequencing technologies have revealed the relation between mutations in genes and the onset of symptoms that can be assigned to corresponding genetic disorders. More than a hundred thousand unique pathogenic variants are available from the Human Gene Mutation Database (HGMD) [1] that are known to cause over ten thousand different monogenic disorders [2] and almost four thousand genes already have been described that are involved in polygenic disorders [3]. Next generation gene sequencing efforts, on the other hand, have revealed that harmless single nucleotide polymorphisms (SNPs) are even more frequent in the human genome. More than ten million DNA variations have been uncovered in the human genome, of which about 4% are located in gene coding regions and about half of those (2%) are non-synonymous SNPs (nsSNPs) that thus result in an amino acid change in the corresponding proteins [4]. In fact, it was estimated by Crawford et al. [4] that on average each gene contains five nsSNPs that are present in more than 5% of the human population. Only a small fraction of these nsSNPs have an effect on the function of the corresponding protein and can be classified as pathogenic. It is evident that gene sequencing is a promising method for diagnosis of genetic disorders, but the frequent occurrence of benign variants drastically hampers routine diagnosis of genetic disorders. |
| Mutation databases, such as HGMD, OMIM [5], and protein specific databases, such as for example the P53 mutation databases [6,7], can be used as reference for previously identified pathogenic variants. In the daily practice of DNA diagnostics one obviously encounters many variants for which these databases have no information available yet. In such cases one normally resorts to literature searches or, if that fails, to any of a series of tools, such as Polyphen-2 [8], SIFT [9], and HOPE [10] that have been developed for the in silico evaluation or prediction of the effects of variants on gene and protein function. However, in many cases variant effects are known for homologous proteins. The concept of evolution is that all homologs share a common ancestor and thus it makes sense that variants on equivalent positions would cause a similar effect (e.g. cause a disease). |
| The first step in transferring mutability data between proteins is to identify the equivalent positions. However, to confidently determine which positions are equivalent is difficult unless the proteins in the alignment are so similar that aligning them becomes trivial. Jordan et al. [11] showed that the publicly available mutant severity prediction methods sometimes produce very poor alignments, and thus questionable predictions. Therefore, protein superfamily information systems that use structural alignments are needed to analyze these proteins and enable data transfer between proteins that are more distantly related but still share a common fold. |
| We have previously described 3DM [12], a system that can generate superfamily systems that fully integrate sequence data with literature data, mutation information, and three-dimensional structures. The 3DM multiple sequence alignments are derived from structure superpositions. This makes them more correct than commonly available alignments, and it allows for larger numbers of sequences to be reliably included in the alignments. 3DM systems contain all available sequence variants (protein- and DNA sequences of all splice variants, with and without leader sequences). We have generated 3DM systems for five proteins that are involved in long QT syndrome (gene names: KCNQ1, KCNH2, SCN5A, SCN1A, KCNJ2) and for four members of the amylase superfamily that are involved in Fabry disease, Schindler- or Kanzaki disease, glycogen storage disease, and cystinuria (gene names: GLA, NAGA, GBE1, SLC3A1, respectively). |
| In this work we show that the mutability of residues can still be transferred even on a superfamily scale where proteins are sequentially very different. The sequence identity between the proteins in our manuscript is as low as 10% and thus these proteins can no longer be aligned by sequence alignment programs. Therefore, protein superfamily information systems that use structural alignments, such as 3DM, are needed to analyze these proteins and transfer data for pathogenicity predictions. Additionally, we show that automated literature mining software can outperform manually curated databases such as HMGD, both in terms of the number of unique mutations extracted, as well as the depth of information per mutation. |
| Method |
| 3DM information systems |
| The 3DM software that generates superfamily information systems is extensively described elsewhere [12,13], and will here only be discussed briefly. A structure based multiple sequence alignment (MSA) forms the backbone of each information system. 3DM uses protein structure data to determine which regions are structurally conserved. These regions are termed core regions and 3DM normally uses only these superfamily core regions to generate the superfamily alignments. All sequences and structures are renumbered so that residues aligned in the MSA get the same number throughout the information system. This enables the transfer of data and knowledge between proteins, and facilitates literature searches for mutations in homologs. |
| Multiple sequence alignments |
| To predict the pathogenicity of non-synonymous variants the quality of alignments is of much greater importance than the completeness of the MSA. Structure based sequence alignment methods, which are used by default in the 3DM systems, tend to produce alignments of higher quality and deeper coverage than classical MSA methods. However, for the long QT related genes, structure information is available for only small parts of the proteins. To enable high-quality predictions, we implemented a method that aligns only those parts of the sequences that can be aligned with great confidence, similar to the way the PROTOMAT algorithm produces what they call BLOCKS: ungapped regions of aligned proteins [14]. For the parts of the long QT related proteins for which structural information is available, structure-based alignments were produced and were subsequently merged with the sequence-based MSAs. |
| Mutation data |
| Mutations were extracted from the literature by the 3DM Mutator module [13]. PubMed was queried for papers containing mutations related to the protein members of the two protein superfamilies here investigated. For the alpha-amylase superfamily Mutator scanned 11,471 full-text papers, whereas mutation-related information for the potassium channel superfamily was extracted from 41,253 full-text articles. In total, this resulted in 5,219 and 65,891 mutations for the alpha-amylase and potassium channel superfamilies, respectively. |
| Results |
| We investigated the transferability of several types of information among members of protein superfamilies, and the power and limitations of automatically extracting mutation data from the literature. |
| Pathogenic variants tend to cluster at equivalent positions |
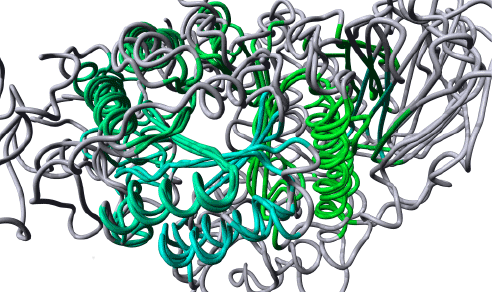
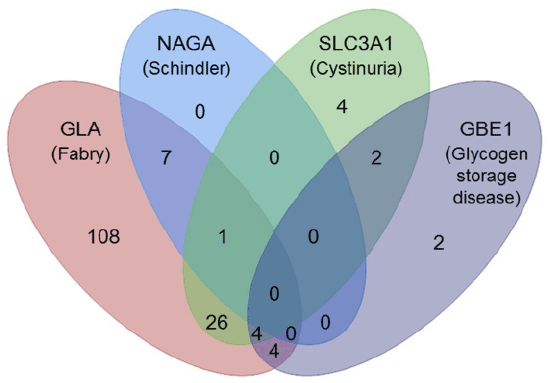
| The basic assumption that allows for the transfer of mutation data between protein family members is that mutations at equivalent positions in homologous proteins tend to result in similar structural and functional effects. Based on this assumption it is to be expected that structurally related proteins have equivalent locations where mutations are well tolerated and locations where mutations are prone to result in detrimental effects. We have tested this hypothesis by analyzing the pathogenic variants that were extracted from the literature by Mutator, in the two protein sets. None of the pathogenic variants are observed with a minor allele frequency of 1% or higher in the ExAC population database [15,16]. The first test set consists of 381 aligned pathogenic variants in four human proteins of the α-amylase superfamily (GLA, NAGA, GBE1, and SLC3A1) that, when mutated, can result in Fabry disease, Schindler- or Kanzaki disease, glycogen storage disease, or cystinuria, respectively. These proteins are sequentially very distantly related to each other. Sequence identities of these proteins range between 10% and 57% as shown in Table 1, which makes it (almost) impossible to align them correctly using sequence based alignment method that are normally used by the standard variant prediction tools (e.g. SIFT or PolyPhen [8,9]), but due to their similar protein structures they can still be aligned correctly (see Figure 1). The 381 pathogenic variants are observed at 158 structurally different positions. Figure 2 shows how often pathogenic variants are observed at corresponding positions in these four proteins. |
| For our second test set, the four potassium channels, structural information is present only for a very small fraction of these proteins, which hampers the transfer of mutation data between these proteins. SCN5A and SCN1A are sequentially closely related and these two can reliably be aligned over nearly the full lengths of their sequences. However, KCNQ1 and KCNH2 can only reliably be aligned at 156 positions that are structurally conserved. This is the transmembrane region of these proteins. No structure data is available for SCN5A and SCN1A and these proteins can reliably be aligned to KCNQ1 and KCNH2 at only 29 of these 156 positions. Due to the absence of structural information and the limited number of mutation data for the four potassium channels, the significance of the overlap of mutation data could only be determined for SCN5A and SCN1A and for the 156 structural conserved positions of KCNQ1 and KCNH2. For these two datasets an even greater overlap is observed of positions that are disease related in both families. Table 1 provides the numerical details of these analyses. |
| Variation in close homologs is indicative of mutation-tolerant positions |
| It is commonly accepted that mutations at conserved positions are likely to be pathogenic, and many MSA-based software packages (e.g. SIFT; [9]) that aim at predicting the significance of mutations for a disease state implicitly use this concept. If an alignment consists only of highly similar sequences, then obviously most positions will be observed as conserved. If in a sequence alignment all sequences are more than 90% sequence identical to the human sequence then obviously the MSA contains only sequences from species that are closely related to homo sapiens, and consequently, any variability observed in this MSA is likely to also be acceptable in the human sequence. To test this hypothesis, we compared the ratio of pathogenic variants at conserved positions with the ratio of pathogenic variants at non-conserved positions. To ensure that this test was statistically meaningful we only used two of the nine human proteins (one from each super-family) for which a large number of different pathogenic variants (>250) are available. Table 2 shows that pathogenic variants are less frequently observed at variable positions in alignments composed of only highly similar sequences. For instance, for 179 of the 420 positions in the alignment of closely related GLAs pathogenic variants (stop codons and deletions excluded) have been reported in the HGMD database. The alignment composed of sequences that are at least 90% identical to GLA contains 387 conserved positions. For 176 (45%) of these positions pathogenic mutations have been described. For the variable positions this is five times less, because for only 3 of the remaining 33 variable positions (9%) pathogenic variants have been described. To determine the significance of the lower frequency of pathogenic mutations at variable positions a p-value was determined, which was <0.01 for all factor values from Table 2. |
| Mutation data extraction: automated mining outperforms manually curated approaches |
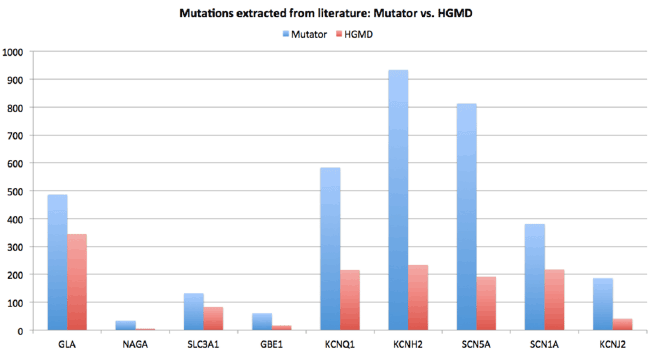
| HGMD [1] is the de facto standard source for mutation information. Like any manually curated database, the high quality of the HGMD comes at the cost of incompleteness. It was shown recently that Mutator is able to extend the HGMD [17]. Figure 3 show that Mutator extracts significantly more mutations from the literature than human experts. This does, of course, not invalidate systems like HGMD because the HGMD also provides details on the effect of a variant. Even though natural language parsing software is developing very rapidly the days that the successor of Mutator will also always correctly extract the effects of those mutations from the literature are not near. In contrast to the HGMD database that stores each unique mutation once, 3DM collects all publications that describe any particular mutation. This can, for example, be two publications that describe the same mutation detected in different patients with different onsets or different symptoms. We have frequently observed that contradicting information is reported for the same mutation in different patients. |
| Discussion |
| We have made a number of interesting observations. First, we find that variants are more likely to be pathogenic if they occur in structurally conserved regions of a protein [13]. Second, we find that it is much less likely that variants are pathogenic if they occur at positions that are variable in alignments composed of only highly similar proteins. From this observation follows that if a close homolog of a human protein has a different residue, it is more likely that other residue types are allowed at the equivalent position of the human protein. Third, we show that there is a large overlap in alignment positions where pathogenic mutations occur even among distantly related human proteins of a superfamily. Therefore, when a missense mutation is pathogenic to its host organism, the chance that a mutation at the equivalent position in a homologous protein (to any residue) is also pathogenic is much higher. These observations show, as was hinted at previously [10], that pathogenicity is much more determined by the location of the mutation in the protein than by the type of amino acid that is introduced. We can conclude that the use of protein superfamily systems can extensively add previously unused data for the investigation of human disease related variants. These revelations can function as very useful predictive features for variant effect prediction models. The availability of a protein superfamily data integration system is valuable for such a model, since it can provide predictive features that otherwise would be missing, such as mutation data for very distant homologs. In fact, these models have been generated for the LQT related genes. We show that the use of superfamily data largely increases the accuracy of variant effect predictions (publication in progress). |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 |
Figures at a glance
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 |
Relevant Topics
- Analytical Biochemistry
- Applied Biochemistry
- Carbohydrate Biochemistry
- Cellular Biochemistry
- Clinical_Biochemistry
- Comparative Biochemistry
- Environmental Biochemistry
- Forensic Biochemistry
- Lipid Biochemistry
- Medical_Biochemistry
- Metabolomics
- Nutritional Biochemistry
- Pesticide Biochemistry
- Process Biochemistry
- Protein_Biochemistry
- Single-Cell Biochemistry
- Soil_Biochemistry
Recommended Journals
- Biosensor Journals
- Cellular Biology Journal
- Journal of Biochemistry and Microbial Toxicology
- Journal of Biochemistry and Cell Biology
- Journal of Biological and Medical Sciences
- Journal of Cell Biology & Immunology
- Journal of Cellular and Molecular Pharmacology
- Journal of Chemical Biology & Therapeutics
- Journal of Phytochemicistry And Biochemistry
Article Tools
Article Usage
- Total views: 12258
- [From(publication date):
March-2016 - Aug 19, 2025] - Breakdown by view type
- HTML page views : 11257
- PDF downloads : 1001
