Make the best use of Scientific Research and information from our 700+ peer reviewed, Open Access Journals that operates with the help of 50,000+ Editorial Board Members and esteemed reviewers and 1000+ Scientific associations in Medical, Clinical, Pharmaceutical, Engineering, Technology and Management Fields.
Meet Inspiring Speakers and Experts at our 3000+ Global Conferenceseries Events with over 600+ Conferences, 1200+ Symposiums and 1200+ Workshops on Medical, Pharma, Engineering, Science, Technology and Business
Editorial Open Access
How to Analyze Real Time qPCR Data?
| Mohammad Najafi* | |
| Biochemistry Department, Razi Drug Research Center, Cellular and Molecular Research Center, Tehran University of Medical Sciences, Tehran, Iran | |
| *Corresponding Author : | Dr. Mohammad Najafi Biochemistry Department Razi Drug Research Center Cellular and Molecular Research Center Tehran University of Medical Sciences, Tehran, Iran Tel/Fax: 982188622742 E-mail: nbsmmsbn@ tums.ac.ir |
| Received February 01, 2013; Accepted February 02, 2013; Published February 05, 2013 | |
| Citation: Najafi M (2013) How to Analyze Real Time qPCR Data? Biochem Physiol 2:e114. doi:10.4172/2168-9652.1000e114 | |
| Copyright: © 2013 Najafi M. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Visit for more related articles at Biochemistry & Physiology: Open Access
| Introduction |
| The comparative transcriptome and genome studies are the fields of molecular biology that describe the biological responses and the organism load. These studies are often based on the high throughput techniques and also the copy-limited quantitative (qPCR) and semiquantitative (RT-PCR) PCR methods. In the PCR-based methods, the analytical sensitivity is dependent on the end-point and kinetic analyses so that Real-Time qPCR is known as a high accuracy technique. The Real-Time qPCR is a time-dependent method to determine the primary RNA or DNA copies, based on the signal detection limitations in the exponential phase [1]. |
| Data Analysis |
| The data analyses are commonly performed in “fold change” and “relative expression” procedures [2,3]. |
| The fold change procedure |
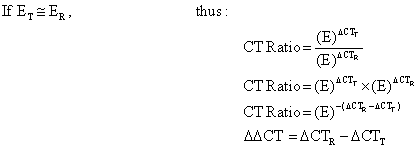
| It shows absolute CT (or Copy) number ratio of target (T) and reference (R) genes in the experiment (Exp) and control (Con) studies without the consideration of statistical tests. Similar to northern blotting technique, the target gene changes (ΔCTT) between the groups (Exp-Con) are normalized to that for the reference gene (ΔCTR) [4]. The procedure needs the standard curves (serial log10-dilution) for the calculations of application efficiency (aE) and copy numbers as presented in the following part: |
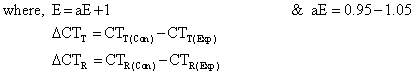 |
 |
| The relative expression procedure |
| In this procedure, the CT (or Copy) numbers of samples are adjusted in the relative mode. Similar to the above procedure, the application efficiencies (aE) (and copy numbers) are calculated using the standard curves and the normalized sample ratios are adjusted to the mean of con-defined group so that they can be compared statistically. |
| The steps are indicated in the following subsections: |
| Determination of CT (Cycle Threshold) number: It is the intersection between threshold line and logarithmic amplification plot and, is used in both procedures. The CT numbers reported at the range of 17-27 are suitable to analyze. |
| CT ratio of target and reference genes: The CT numbers of target genes (CTT) are normalized by these prepared for reference genes (CTR) in all samples (S). |
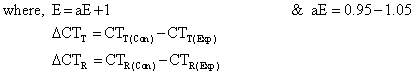 |
 |
| Adjusting CTs: The mean of CTs (mCTs) is calculated in the Con-defined group (or Zero group) and considered to be equal to 1 (mCTs(Con)=1). Other CTs values are adjusted on the basis of mCTs(Con). This approach is able to adjust two variables, ΔCT and E. |
| The relative expression plots (obtained for several groups) are important to show statistical differences. Based on the non parametric distribution of data, the CTs values may be adjusted with the median of CTs(Con) (medCTs(Con)). Although, it usually has not effect on comparative statistical results and the adjusted data distribution but, it can affect the fold reports on the plots. Moreover, the data adjustment with medCTs(Con) may introduce the large deviations around the measures of central tendency (Mean or Median) dependent on the distance between the median and the mean in each group. |
| Statistical tests: The non parametric distribution of the adjusted CTs values is usually evaluated with One-Sample Kolmogorov-Smirnov Test. Furthermore, the data distribution between subgroups can be evaluated using Median Test. In most studies, they are significantly lower than 0.05 so that the non parametric analyses must be performed with Mann Whitney U, Kruskal-Wallis (Independent samples), Wilcoxon and Friedman (Related samples) tests. On the other hand, the data must be evaluated with student t, ANOVA (Independent samples) Pairs, ANCOVA (Related samples) tests [5,6]. |
| In conclusion, the fold changes show absolutely the between-group alterations while the relative changes present statistically betweengroup differences after adjusting the data in the real time qPCR data analyzing procedures. The statistical tests are dependent on the study method and data distribution. |
References
|
Post your comment
Relevant Topics
- Analytical Biochemistry
- Applied Biochemistry
- Carbohydrate Biochemistry
- Cellular Biochemistry
- Clinical_Biochemistry
- Comparative Biochemistry
- Environmental Biochemistry
- Forensic Biochemistry
- Lipid Biochemistry
- Medical_Biochemistry
- Metabolomics
- Nutritional Biochemistry
- Pesticide Biochemistry
- Process Biochemistry
- Protein_Biochemistry
- Single-Cell Biochemistry
- Soil_Biochemistry
Recommended Journals
- Biosensor Journals
- Cellular Biology Journal
- Journal of Biochemistry and Microbial Toxicology
- Journal of Biochemistry and Cell Biology
- Journal of Biological and Medical Sciences
- Journal of Cell Biology & Immunology
- Journal of Cellular and Molecular Pharmacology
- Journal of Chemical Biology & Therapeutics
- Journal of Phytochemicistry And Biochemistry
Article Tools
Article Usage
- Total views: 15835
- [From(publication date):
May-2013 - Jun 14, 2025] - Breakdown by view type
- HTML page views : 11100
- PDF downloads : 4735
