Short Communication Open Access
Rc Gene Sequence and Expression Evaluation in a Red-Kernel Rice Genotype
| Barbara Ferrari1*, Alberto Gianinetti2, Franca Finocchiaro2 and Valeria Terzi2 | |
| 1Consiglio per la ricerca in agricoltura e l’analisi dell’economia agraria, Fodder and Dairy Productions Research Centre (CREA-FLC), viale Piacenza 29-26900 Lodi, Italy | |
| 2Consiglio per la ricerca in agricoltura e l’analisi dell’economia agraria, Genomics Research Centre (CREA-GPG), via San Protaso 302-29017 Fiorenzuola D’Arda, Italy | |
| Corresponding Author : | Barbara Ferrari Consiglio per la ricerca in agricolturae l’analisi dell’economia agraria Fodder and Dairy Productions Research Centre (CREA-FLC) viale Piacenza 29-26900 Lodi, Italy Tel: +39 0371 404726 Fax: +39 0371 31853 E-mail: barbara.ferrari@entecra.it |
| Received July 29, 2015; Accepted August 29, 2015; Published September 02, 2015 | |
| Citation: Ferrari B, Gianinetti A, Finocchiaro F, Terzi V (2015) “Rc Gene Sequence and Expression Evaluation in a Red-Kernel Rice Genotype”. J Rice Res 3:146. doi:10.4172/2375-4338.1000145 | |
| Copyright: © 2015 Ferrari B, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
| Related article at Pubmed, Scholar Google | |
Visit for more related articles at Rice Research: Open Access
Abstract
Most cultivated rice is white all over the world; however, pigmented rice varieties could be preferable, as they have the potential to promote human health because containing antioxidative compounds inhibiting/reducing reactive cell-damaging free radicals-even if few in vivo studies have been reported about bioactivity of red rice on humans. Previous studies demonstrated that, generally, dehulled red rice shows a total antioxidant activity more than three times greater than dehulled white rice, due to the presence of proanthocyanidins, which accumulate as colourless compounds during the early steps of maturation and subsequently give the seed its pigmentation after oxidative reactions along maturation and desiccation process. Rc gene, which codifies for a bHLH (basic helix-loop-helix) protein responsible for proanthocyanidins’ accumulation into the inner seed coat below the pericarp, was here studied from the point of view of expression, developing and applying a qRT-PCR (quantitative reverse transcription polymerase chain reaction) assay to all the grain filling phases. Results showed that proantocyanidins are present into the caryopses since the milky stage. This study confirmed the key role of the 14bp deletion in Rc-bHLH in the suppression of rice pigmentation and precisely showed that the maximum Rc gene expression was detected at an early stage of grain-filling, i.e., at the elongation period. Moreover, starting from this specific knowledge of the Rc expression timing, targeted highthroughput experiments (transcriptomics and RNA-seq) could be greatly facilitated for expression analyses in seeds of all pigmented rices and presumably of different small grain cereals.
| Keywords |
| Rc expression; Red rice; Seed development; Antioxidant |
| Introduction |
| Red kernels reflect an ancestral feature of rice, in fact red pericarp is ubiquitous among wild ancestors. However, genotypes with white caryopses have been selected by man several centuries ago (probably because of the more appreciated aspect and the better cooking properties) and, as a consequence, nowadays most cultivated rice is white all over the world. A new interest is anyway arising now on pigmented rice varieties which could be preferable, having the potential to promote human health with their antioxidative compounds inhibiting/reducing reactive cell-damaging free radicals-even if few in vivo studies about bioactivity of pigmented rice have been reported on humans . Chemical studies comparing red rice-here referred to as the rice which presents dehulled caryopses with red coloured integuments and not as weedy rice (not suitable for cultivation)-and white rices have demonstrated that generally dehulled red rice shows a total antioxidant activity more than three times greater than dehulled white rice. Milling and cooking cause significant losses of antioxidants: red rice maintains however higher values, but a full uptake of cooking water by the grains (“risotto”) is better for the nutritional quality, since in this way no soluble antioxidants are lost with the discharged water [1-3]. Red seeds were found to contain proanthocyanidins, which accumulate as colourless compounds during the early steps of embryogenesis, and subsequently they give the seed its chromatic pigmentation after oxidative reactions occurring along maturation and desiccation process. |
| Genetic studies have shown that the Rc gene, which codifies for a regulatory bHLH (basic helix-loop-helix) protein, has a key role in proanthocyanidins accumulation and that the loss of coloration of the common white seed coat is often due to a 14-bp deletion in this gene: Rc gene sequence for O. rufipogon was deposited with GenBank accession number DQ315482. The mutation creates in fact a nonsense codon that blocks the production of proanthocyanidin pigmentation in the pericarp Moreover, Rc gene expression shows a strong tissuespecificity, being activated only in grain-related tissues To gain new information on the modulation of the Rc gene, the timing of its expression was studied in an Italian rice genotype, RNC6, having a red caryopsis that, however, does not shatter at maturity [4-7]. |
| Materials and Methods |
| The RNC6 Rc nucleotide sequence was assessed and its expression was followed in tailored experiments. DNA extraction was performed from RNC6 leaf tissues and the Rc gene was sequenced. Also a white genotype, Perla, and a red genotype, Perla Rosso were sequenced to produce bases alignments. DNA sequences were aligned with BioEdit software, comparing with the published Rc gene of O. rufipogon. |
| The colour of the RNC6 caryopses was also visually analysed after 2% KOH extraction, a useful preliminary test for immature caryopses in which the pigmentation is not yet discernible. The fast KOH test on grains detects the accumulation of proanthocyanidins: red rice caryopses develop a red coloration in 5-10 minutes, while rice’s with white pericarp usually give only a light yellow colour [4,8]. |
| For the expression study, RNC6 kernels were consequently sampled at these following caryopsis-filling stages [9]: |
| • Elongation (stage 1, the caryopsis is elongating to the end of the hull; the endosperm is watery and the integuments are green; partially corresponding to the first part of stage R5) |
| • Expansion (stage 2, the caryopsis has elongated to the end of the hull and is expanding; partially corresponding to the final part of stage R5 to the initial part of stage R6 ) |
| • milky (stage 3, the caryopsis is filled with a white liquid resembling milk; partially corresponding to the middle of stage R6) |
| • Soft dough (stage 4, endosperm is still soft; partially corresponding to the end of stage R6). |
| • Hard dough (stage 5, endosperm is firm; partially corresponding to the end of stage R7). |
| Caryopses were dehulled one by one with scalpel and tweezers and total RNA was extracted according to RNeasy Plant Mini Kit protocol (Qiagen GmBH,). |
| Reverse transcription was followed by qPCR analysis using the following primer pairs, designed exploiting the information stored in public databases TIGR (www.tigr.org) and Genbank/NCBI (www.ncbi.nlm.nih.gov): |
| RISO1 F- GGCATTTTCTCTGCGAGGAGT; RTGGCAGTGCTTGTGTGAAGAGT (amplicon size of 124 bp); |
| RISO2 F- GTGGTAAGAGATGGCTGCTCAA; RCATCTGGCCCTTGCAAATTT (amplicon size of 103 bp); |
| RISO3 F- TGCGACAGCAAGAGATGAAC; RGGTGTTTCTGCTCCCCTGTA (amplicon size of 193 bp); |
| RISO4 F- GCGACAGCAAGAGATGAACA; RGGTGTTTCTGCTCCCCTGTA (amplicon size of 192 bp); |
| A standard RT-PCR amplification profile was used: initial step at 48°C for 30 min, denaturation at 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min; the dissociation stage was at 95°C for 15 sec, and the final annealing at 60°C for 30 sec. A blank control was added to each primers pair assay. |
| Two reference genes were tested for normalization, eEF-1α, eukaryotic elongation factor 1-alpha and UBQ5, ubiquitin 5 (accession numbers, respectively, AK061464 and AK061988). GenEx ver. 4.3 software packages (Multid Analyses AB, Lotsgatan 5A, Goteborg, Sweden) was used for normalizing the RTqPCR data using one reference gene, to minimize the technical variation and obtaining relative quantification. |
| For each gene, expression was calculated relatively to the samples where the gene showed the highest expression, by using this equation: Nrel=2Ct min-Ct being Ct min the threshold cycle for the most expressed gene and Ct the threshold cycle for the gene of interest [10-13]. |
| Results |
| In RNC6 sequence, the Rc-bHLH 14-bp deletion-described by [7&4] in white grain genotypes-was found, but it was accompanied by an additional Guanine deletion 44 bp before the deleted portion. It corresponds to the Rcr mutation, which restores for the Rc gene the reading frame lost in the rc allele by the original 14-bp deletion that gave rise to white rice (Figure 1) the red colouration appears for RNC6 rice and increases during grain filling. Immature grains of pigmented rices are usually green just like the ones from white rices, but they already contain the pigment’s precursors. Proanthocyanidins were present in RNC6 at the milky stage, which means that the regulatory genes for their accumulation have been expressed before. In fact, gene expression is usually more dynamic and intense at the beginning of the grain filling phase [6&14]. |
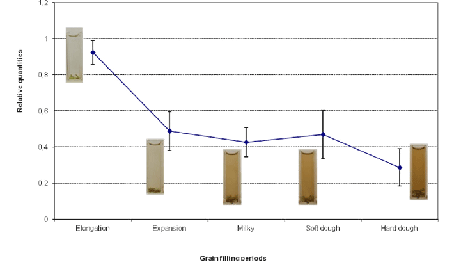
| The amplification plot for eEF1α gene showed the most uniform and stable expression trend in all samples (not shown); so, it was used as endogenous control for normalization of our RTqPCR data. Figure 2 shows that the highest expression for Rc gene was observed during the elongation phase; then, it decreased rapidly in the expansion phase and slowly in the subsequent stages (Figure 2). |
| Discussion |
| In this work we confirmed the key role of the Rc-bHLH 14bp deletion in the suppression of rice pigmentation and we precisely defined in which developmental phase Rc expression is activated. Moreover, starting from this specific knowledge of the Rc expression timing, targeted experiments could be programmed to study environmental effects on the expression levels and therefore on the final content of antioxidants in red rice. Further high-throughput activities (transcript omics and RNA-seq) can be greatly facilitated by the adoption of this proper timing for expression analyses in seeds of all pigmented rice’s and presumably also of different small grain cereals. |
| Acknowledgements |
| This work was supported by “AGRONANOTECH” project of the Ministry of Agriculture and Forestry. We particularly thank Renzo Alberici for the accurate photographs concerning rice caryopses in development and the relevant assistance of all the CREA-GPG technical staff involved in the care of rice materials. |
References |
|
Figures at a glance
 |
 |
| Figure 1 | Figure 2 |
Relevant Topics
- Basmati Rice
- Drought Tolerence
- Golden Rice
- Leaf Diseases
- Long Grain Rice
- Par Boiled Rice
- Raw Rice
- Rice
- Rice and Aquaculture
- Rice and Nutrition
- Rice Blast
- Rice Bran
- Rice Diseases
- Rice Economics
- Rice Genome
- Rice husk
- Rice production
- Rice research
- Rice Yield
- Sticky Rice
- Stress Resistant Rice
- Unpolished Rice
- White Rice
Recommended Journals
Article Tools
Article Usage
- Total views: 16007
- [From(publication date):
October-2015 - Aug 30, 2025] - Breakdown by view type
- HTML page views : 11141
- PDF downloads : 4866
