Research Article Open Access
Stability Analysis of Rice Root QTL-NILs and Pyramids for Root Morphology and Grain Yield
| Grace Sharon Arul Selvi1, Farhad Kahani2 and Shailaja Hittalmani3* | |
| 1Department of Genetics and Plant Breeding, University of Agricultural Sciences, Bangalore-65, India | |
| 2Marker Assisted Selection Laboratory, Genetics and Plant Breeding, University of Agricultural Sciences, GKVK, Bangalore 560065, India | |
| 3University Head, Genetics and Plant Breeding, University of Agricultural Sciences, GKVK, Bangalore-560065, India | |
| Corresponding Author : | Shailaja Hittalmani Professor and University Head Genetics and Plant Breeding University of Agricultural Sciences GKVK, Bangalore 560065, India Tel: 91-8023624967 E-mail: shailajah_maslab@rediffmail.com |
| Received October 02, 2015; Accepted October 26, 2015; Published October 31, 2015 | |
| Citation: Selvi GSA, Kahani F, Hittalmani S (2015) Stability Analysis of Rice Root QTL-NILs and Pyramids for Root Morphology and Grain Yield. J Rice Res 3:153. doi:10.4172/2375-4338.1000153 | |
| Copyright: © 2015 Selvi GSA, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
| Related article at Pubmed, Scholar Google | |
Visit for more related articles at Rice Research: Open Access
Abstract
Cultivation of rice in the rain fed conditions is threatened by frequent spells of water deficits and limits the productivity to a greater extent. Root system plays a major role in uptake of water and they contribute to drought tolerance in a major way. In this study, Root QTL were pyramided and evaluated under aerobic and drought conditions and the stable genotypes were identified. Two QTL and three QTL pyramid lines for roots were developed and evaluated under drought, aerobic and in different locations to study the performance. While qRT26-9 with 2 QTL pyramid performed better with respect to the root traits, qRT16-1+7 and qRT17-1+7 performed better for shoot morphology over the various growth water regimes. Among the pyramids, qRT11-7 × qRT18-1+7-17 recorded increased performance for plant height and seed yield while qRT11-7 × qRT18-1+7-32 recorded increased performance for total biomass and maximum root length. qRT24-9 × qRT11-7-32 recorded increased performance for root traits only across environments. Lines with high means and average stability were identified as suitable across growth niches, while those with low stability and high means were identified as suitable for growth under poor environments and for specific locations.
| Keywords |
| Rice; Aerobic; Stability; Root QTLs; Environments |
| Introduction |
| Rice (Oryza sativa L), the second most important cereal of the world is traditionally grown under submerged anaerobic conditions. However, this cultivation is now foraying into the less traditional rain fed uplands and marginal lands with mounting pressure on land availability. This coupled with changes in the climate make cultivation in these delicate ecosystems rather intricate. Therefore, the development of genotypes that consistently perform under conditions of climate change with less moisture availability is a viable option. The constancy or preferably increase in yield potential under climate change scenario is fundamental for food sustainability in the near future, given the expected population growth projections. Cultivation in the rain fed uplands is threatened by frequent spells of water deficits being a major limiting factor directly affecting grain yields during reproductive phases. Several mechanisms that determine drought tolerance and or resistance have been outlined, of which manipulation of the root system to maintain the water status of a crop under conditions of increasing water deficits has been the choice breeding strategy for drought. Several QTLs governing root traits across populations have been identified in rice. Root studies have become very important now that there are several ways to study them [1-17]. |
| Pyramiding of genes is conducted to develop a genotype that expresses the said genes genes appropriately, such that the phenotype is enhanced. It has been used extensively in major gene controlled rice blast, rice blight and against insect pests such as diamond back moth (Cao et. al., 2002). Pyramids enhance the phenotype effectively and can be used to analyse the effect of QTLs upon each other as they offer a common background for the QTLs to interact. Subsequently QTL pyramiding was attempted by several researchers. Consistent and quality performance of developed genotypes is always desired as it increases the longevity of the genotypes. In breeding exercises, stable and high performance of developed varieties in target growth environments or across different environments and or seasons is an important attribute. Stability of the lines is measured as a non-significant deviation from its regression coefficient and is stated with reference to its mean. Lines with high means and average stability can be identified to suit in most environments [18-30]. |
| Materials and Methods |
| Plant material |
| A set of twenty-nine near-isogenic lines with Root QTL introgressions of IR64 (indica, high yielding) with QTL introgressions from Azucena (japonica, drought tolerant) controlling root morphology (QTL Introgressed Lines (QILs)) developed by [31] and fine mapped by [16] was used for the study. These QILs were used in a pairwise crossing programme to develop 2 and 3 QTL pyramids. The QILs, the generated pyramids along with parents: IR64, Azucena and checks: Budda and Moroberekan were evaluated in RCBD design with 2 replications over the various growth regimes (Table 1) in 2011-2012 (Tables 2-4). |
| Phenotypic observations |
| Five plants with QTL pyramids were selected at random in each entry for recording observations. The average of these five plants was used for the statistical analysis. The individual plants were observed for plant height (cm) from the base to the tip of the panicle at harvest days to 50% flowering i.e., first flowering in 50 per cent of the plants, number of tillers per plant, number of panicles per plant, panicle length (cm) from collar to the tip, number of filled grains per panicle, number of chaffy grains per panicle, panicle weight (g) total grain weight per plant (g) root length (cm) from the crown to the tip of the longest roots, root thickness (mm), root number at 15cm root depth, total biomass (g) and test weight (g) was observed. |
| Statistical analyses |
| The data that was generated was subject to a series of statistical analyses to elucidate the relative effects of the presence of various QTLs in the rice genotypes and are presented here below: |
| Two way Analysis of variance: The data obtained was subjected to two way analysis of variance using the method outlined by [32] for each character in order to ascertain existence of genotype x environment interaction. If interaction was found to be significant, then the data was further subjected to stability analysis. |
| Stability analysis: The stability model proposed by [33] was adopted to analyze the data over the studied environments. The model considers three parameters: the mean (M), the regression co-efficient (bi) which is the regression of the mean of environmental index and deviation for regression (S2di), which is a measure of genotype -environment interaction of an unpredictable type. |
| The model involves the estimation of three stability parameters: mean (μi), regression co-efficient (bi) and deviation from regression (S2di), which are defined by the following mathematical formula. |
| Yij=μi + βiIj + δij |
| Where, |
| Yij : mean of the ith genotype in the jth environment. |
| μi : mean of the ith genotype over environments. |
| βi : regression co-efficient that measures the response of ith genotype to varying environment. |
| δij : deviation from regression of the ith genotype in the jth environment and |
| Ij : environmental index obtained by subtracting the grand mean of the ith genotype from the mean of all genotypes in the jth environment. |
| Stability parameters |
| The mean (μi), the regression co-efficient (bi) and mean square deviation from linear regression line (S2di) are the three stability parameters proposed by [33] in their stability model. The three parameters are computed using the following formulae: |
| Mean: |
| Regression co-efficient |
| Deviation from regression co-efficient |
| Where, |
| n : number of environments |
| Where, |
| n : number of environments |
| v : number of genotypes with |
| The total variation is partitioned into genotypes, environment, environment (linear), genotype x environment (linear), pooled deviation and pooled error. |
| F test |
| a. To test the significance of differences among the genotypic means, the ‘F’ test followed was: |
| Where, MS1 : mean sum of squares of varieties |
| MS3 : mean sum of squares of pooled deviation |
| b. To test individual from linear regression, the formula is as follows, |
| Where, n: number of environments |
| c. To test the hybrids/ varieties which do not differ for their regression on the environmental index, the appropriate test was, |
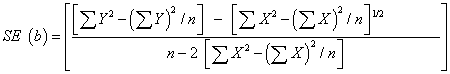 |
| Where, |
| X : environmental index |
| n : number of environments |
| A joint consideration of the three parameters such as |
| 1. The mean performance of the genotype over the environments (x) |
| 2. The regression co-efficient (b) |
| 3. The deviation from linear regression (S2d) is used to define stability of a genotype. |
| The estimate of deviations from regression (S2d) suggests that the degree of reliance that should be put to linear regression in interpretation of the data. If these values are significantly deviating from zero, the expected phenotype cannot be predicted satisfactorily. When, deviations (S2d) are not significant, the conclusion may be drawn by the joint consideration of mean, yield and regression co-efficient (b) values as given below (Table 1). |
| While interpreting the results, S2di is first looked into. A nonsignificant deviation from S2di=0, then stability is interpreted based on bi and mean values. If bi=1, a genotype is considered to possess average stability i.e., same performance in all the growth conditions. If bi is more than unity, then the genotype is said to have less than average stability i.e., good performance under favorable environments. If bi is less than unity, then the genotype is said to possess above average stability i.e., good performance under poor environments. Thus, genotypes possessing unit regression coefficient and non-significant deviation from regression were considered ideal, widely adapted and stable genotypes. |
| Results |
| Environment- wise analysis of variance indicated a significant mean sum of squares for the QTL-NILs and the generated pyramids for most characters studied. Combined Analysis of Variance for the pyramids qRT24-9 x qRT11-7 (data not shown) indicated significant variance for genotype as well as for genotype x environment [33] model for stability analysis was applied as the genotype x environment component of variance was found significant. The performance of genotypes in different environments for five selected characters based on two-way analysis of variance and Bartlett’s test are elaborated below (Table 5). |
| Plant height |
| The varying environmental indices indicated that there was a significant difference for plant height across environments and across genotypic entries. qRT6-2 x qRT19-1+7 was the shortest (47.74cm), while qRT6-2 x qRT11-7 was the tallest (62.77 cm). The bi and S2di were found to be non-significant for all the genotypes studied. |
| Seed yield per plant |
| As indicated by the environmental indices and the environmental means, seed yield per plant showed significant differences across environments. qRT11-7 x qRT6-2 recorded the highest seed yield (9.95 g), while qRT11-7 x qRT18-1+7 recorded the least seed yields (6.92 g). The bi and S2di were found to be non-significant for all the genotypes studied. |
| Total Biomass per plant |
| The varying environmental indices indicated that there were significant differences for total biomass across environments. qRT6-2 x qRT19-1+7 was the lightest (35.82 g), while qRT20-1+7 x qRT18- 1+7 was the heaviest (50.92 g). The bi and S2di were found to be nonsignificant for all the genotypes studied. |
| Maximum root length |
| As indicated by the environmental indices and the environmental means (11.86 to 18.63), maximum root length showed significant differences across genotypes. The QTL-NILs recorded the highest mean maximum root length (18.63 cm), while qRT11-7 x qRT6-2 recorded the least root length (11.86 cm). The bi and S2di were found to be nonsignificant for all the genotypes studied (Table 6). |
| Total number of roots per plant |
| The varying environmental indices indicated that there were significant differences for total number of roots across environments. qRT6-2 x qRT11-7 had the highest mean number of roots (59.13), while qRT11-7 x qRT18-1+7 had the least mean number of roots (52.65). The bi and S2di were found to be non-significant for all the genotypes studied. |
| Discussion |
| Phenotype of an individual is determined by the interaction of the genotype and environment surrounding it, the effects of genotype and environment on phenotype may not always be independent. The phenotypic response to change in environment is not the same for all the genotypes. The interplay in the genetic and non-genetic effects on development is termed as “genotype environment” interaction (Comstock and Moll, 1963) and is of major consequence to the breeder in the process of evolution of improved genotypes. |
| In the present study, twenty nine near isogenic lines of IR64 introgressed with QTL regions on four chromosomes: 1,2,7 and 9 from Azucena, 7 pyramids generated from these lines and the checks: IR64, Azucena, Budda and Moroberekan were grown in three seasons.2011,Season I. 2012 wet and dry seasons. (Season II and Season III). During season 1, they were grown under reproductive stage low moisture stress and well watered conditions for growth, yield and root traits at MRS, Hebbal. During season 2, they were grown under reproductive stage low moisture stress and well watered conditions at Farmer’s field, Pavagada for growth and yield traits. During season 3, the genotypes were grown under reproductive stage low moisture stress and well watered conditions at ZARS, GKVK, under submerged conditions at Farmer’s field, Dodjala and under aerobic non-stress conditions at Farmer’s field, Shettigere for growth, yield and root traits. The genotypes were also grown under vegetative stage low moisture stress and well watered conditions for growth and root traits at MRS, Hebbal. |
| Mean performance of the QTL-NILs and generated pyramids |
| The aerobic, non stress growth condition of Farmer’s field, Shettigere (data not shown) during season 3 was the most conducive environment for plant height in the QTL-NILs, with a mean height of 71.85 cm. Low moisture stress condition of season 2, Farmer’s field, Pavagada was the least conducive for the QTL-NILs. qRT21-1+7 was the tallest genotype across locations. All the generated pyramids recorded maximum plant height under aerobic, non-stress condition at Farmer’s field, Shettigere. The tallest pyramids generated were qRT11-7 xqRT18-1+7-7, qRT24-9 x qRT11-7-12, qRT6-2 x qRT11-7- 15, qRT11-7 x qRT19-1+7-21, qRT20-1+7 x qRT18-1+7-4, qRT11-7 x qRT6-2-1 and qRT6-2 x qRT19-1+7-3. Among the pyramids, qRT11-7 x qRT18-1+7 was significantly taller, while qRT6-2 x qRT19-1+7 was significantly shorter. |
| For grain yield in QTL-NILs, the submerged conditions provided by Farmer’s Field, Dodjala during season 3 was the most conducive, while the least mean yield were recorded under low moisture stress, MRS, Hebbal during season1. A highest mean yield across environments was recorded in qRT11-7. For the pyramids, qRT11-7 x qRT18-1+7, qRT24-9 x qRT11-7, qRT11-7 x qRT19-1+7, qRT11-7 x qRT6-7 and qRT6-2 x qRT19-1+7, the most conducive environment was the aerobic non-stress condition during season 3 at Farmer’s field Shettigere while qRT6-2 x qRT11-7 and qRT20-1+7 x qRT18-1+7 recorded highest mean yields under submerged conditions, season 3, Farmer’s field, Dodjala. The highest yielding pyramids were qRT11-7 x qRT18-1+7-15, qRT24-9 x qRT11-7-1, qRT6-2 x qRT11-7-15, qRT11- 7 x qRT19-1+7-16, qRT20-1+7 x qRT18-1+7-17, qRT11-7 x qRT6-2-1 and qRT6-2 x qRT19-9. |
| The highest mean total biomass was recorded by the QTL-NILs under well watered condition, season 2, Farmer’s field, Pavagada, while the least biomass was recorded under season 1, low moisture stress at MRS, Hebbal. Highest biomass across location was recorded by qRT18- 1+7. For the pyramids qRT11-7 x qRT18-1+7, qRT6-2 x qRT11-7, qRT11-7 x qRT19-1+7 and qRT6-2 x qRT19, the most conducive environment for total biomass production was season 2, Farmer’s field, Pavagada, while for qRT24-9 x qRT11-7, qRT20-1+7 x qRT18-1+7 and qRT11-7 x qRT6-2 yielded highest biomass under aerobic, nonstress conditions during season 3 at Farmers field, Shettigere. Highest biomass were produced by qRT11-7 x qRT18-1+7-4, qRT24-9 x qRT11-7-13, qRT6-2 xqRT11-7-25, qRT11-7 x qRT19-1+7-20, qRT20- 1+7 x qRT18-1+7-8, qRT11-7 x qRT6-2-38 and qRT6-2 x qRT19-30. |
| Mean maximum root length was recorded by the QTL-NILs under low moisture stress, season 1, MRS, Hebbal, with the maximum root length being recorded in qRT26-9. For the pyramids qRT11- 7 x qRT18-1+7, qRT24-9 x qRT11-7, qRT6-2 x qRT11-7, qRT11-7 x qRT19-1+7, and qRT6-2 x qRT19, the most conducive environment for longer root production was low moisture stress condition, season 1, MRS, Hebbal, while qRT20-1+7 x qRT18-1+7 and qRT11-7 x qRT6-2 recorded longest roots under well watered conditions, ZARS, GKVK during season 3. Longest roots were produced by qRT11-7 x qRT18- 1+7-27, qRT24-9 x qRT11-7-32, qRT6-2 xqRT11-7-28, qRT11-7 x qRT19-1+7-10, qRT20-1+7 x qRT18-1+7-33, qRT11-7 x qRT6-2-12 and qRT6-2 x qRT19-24. |
| Highest number of roots were produced by the QTL-NILs under well watered conditions, season 3, ZARS, GKVK, with the highest number of roots being produced by qRT17-1+7. For the pyramid qRT11-7 x qRT18-1+7 the most conducive environment for number of root production was well watered conditions, season 3, ZARS, GKVK, while for qRT24-9 x qRT11-7, qRT11-7 x qRT19-1+7, qRT11-7 x qRT6-2 and qRT6-2 x qRT19, the most conducive environment for number of root production was season 3, aerobic, non-stress condition, Shettigere and for qRT6-2 x qRT11-7 and qRT20-1+7 x qRT18-1+7 the most conducive environment was submerged condition, season 3, Farmer’s field, Dodjala. Highest number of roots were produced by qRT11-7 x qRT18-1+7-14, qRT24-9 x qRT11-7-1, qRT6-2 xqRT11-7- 31, qRT11-7 x qRT19-1+7-6, qRT20-1+7 x qRT18-1+7-8, qRT11-7 x qRT6-2-13 and qRT6-2 x qRT19-4. |
| The aerobic non-stress condition therefore is the most conducive environment to grow the present genotypes. Similar results were obtained by [3,34]. This was opined to be due to aeration of roots leading to efficient utilization of resources [35-38]. |
| Genotype x environment interaction |
| Prior to stability analysis, Bartlett test was done. Based on this test, five characters were selected. Homogeneity in the error variances allowed pooled analysis. |
| The mean sum of squares due to genotypes as well as environments was found to be significant in the two way analysis. Significant GXE interaction was obtained both by two-way analysis and the [32] model. The analysis of variance for stability indicated significant differences among the QTL-NILs as well as between and within the generated pyramids for all the characters. The significant environment (linear) variance indicated considerable additive environmental variance. Variance due to GX E interaction was found to be significant for all the characters indicating differential response of the genotypes in different environments. G X E (linear) was significant for all the characters indicating a contribution of linear portion of GE interaction. The more pronounced linearity of characters indicated that variation among the genotypes could be largely explained by the differences [3,39-41]. |
| Stability parameters |
| Five characters were selected on the basis of homogeneity of error variances and after significance of G X E interactions. Identification of genotypes that perform stably over a range of growth environs would therefore be necessary. Of the many models proposed to this effect, the Eberhart and Russel model was used in the present study. Taller plants were preferred as these could lend to fodder yield in a mixed cropping system. Increase in height coupled with increase in total biomass content could result with increase in number of tillers per plant and flowering mattered for the escapes [42]. Maximum root length as a mechanism of drought tolerance ensures higher crop yields under stress situations. Based on the five characters taken together, among the QTL-NILs, qRT24-9 was found to be best in performance across locations, moisture regimes and seasons. While qRT26-9 performed better with respect to the root traits, qRT16-1+7 and qRT17-1+7 performed better for shoot morphology over the various growth regimes. Among the pyramids, qRT11-7 x qRT18-1+7-17 recorded increased performance for plant height and seed yield while qRT11- 7 x qRT18-1+7-32 recorded increased performance for total biomass and maximum root length. qRT24-9 x qRT11-7-32 recorded increase in performance for root traits only across environments. qRT6-2 x qRT11-7-15 recorded increase in performance for plant height and seed yield across environments qRT20-1+7 x qRT18-1+7-15 was the best in performance considering seed yield, total biomass and number of roots per plant. qRT6-2 xqRT19-1+7-30 recorded best performance for seed yield per plant and total biomass. Since all these genotypes recorded non-significant deviation of the regression coefficient (bi) from 1 and S2di approaching zero, we can conclude that these genotypes have average stability across locations [3,38]. |
| The pyramids of root QTL are very relevant as root morphological parameters are controlled by quantitative genes and these do not act independently. When they are moved into new background the effect and the stability of the QTL in environment play a major role. Both environment specific and pyramids suitable for wider range of environments are useful. Hence root QTL pyramiding is useful for developing genotypes for using in water saving technologies like aerobic cultivation or in case where dry spells prevail and roots help to tide over and minimize the economic loss to the rice growing farmers. |
References |
|
Tables and Figures at a glance
| Table 1 | Table 2 | Table 3 |
| Table 4 | Table 5 | Table 6 |
Relevant Topics
- Basmati Rice
- Drought Tolerence
- Golden Rice
- Leaf Diseases
- Long Grain Rice
- Par Boiled Rice
- Raw Rice
- Rice
- Rice and Aquaculture
- Rice and Nutrition
- Rice Blast
- Rice Bran
- Rice Diseases
- Rice Economics
- Rice Genome
- Rice husk
- Rice production
- Rice research
- Rice Yield
- Sticky Rice
- Stress Resistant Rice
- Unpolished Rice
- White Rice
Recommended Journals
Article Tools
Article Usage
- Total views: 15397
- [From(publication date):
November-2015 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 10564
- PDF downloads : 4833
