Our Group organises 3000+ Global Conferenceseries Events every year across USA, Europe & Asia with support from 1000 more scientific Societies and Publishes 700+ Open Access Journals which contains over 50000 eminent personalities, reputed scientists as editorial board members.
Open Access Journals gaining more Readers and Citations
700 Journals and 15,000,000 Readers Each Journal is getting 25,000+ Readers
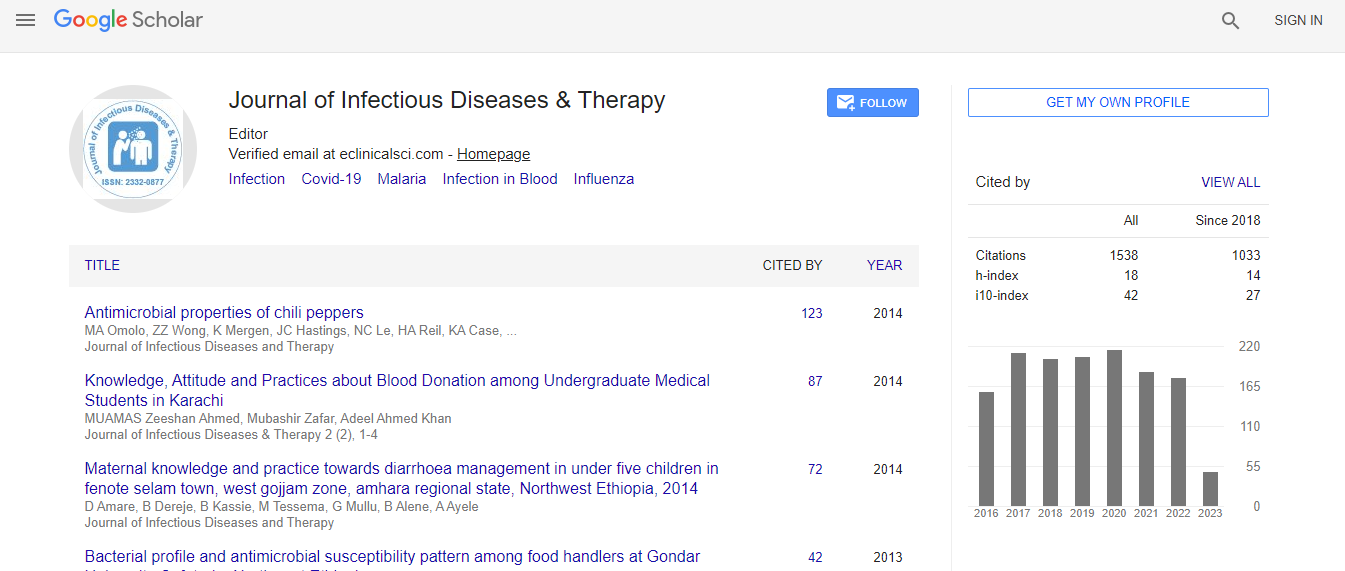
Google Scholar citation report
Citations : 1529
Journal of Infectious Diseases & Therapy received 1529 citations as per Google Scholar report
Indexed In
- Index Copernicus
- Google Scholar
- Open J Gate
- RefSeek
- Hamdard University
- EBSCO A-Z
- OCLC- WorldCat
- Publons
- Euro Pub
- ICMJE
Useful Links
Recommended Journals
Related Subjects
Share This Page
Investigation of the role of the sRNA RyhB in regulating the locus of enterocyte effacement pathogenicity Island in enteropathogenic Escherichia coli
Joint Event on 2nd World Congress on Infectious Diseases & International Conference on Pediatric Care & Pediatric Infectious Diseases
Marisa Egan, Jasmine Ramirez, Christian Xander and Shantanu Bhatt
Saint Joseph��?s University, USA
ScientificTracks Abstracts: J Infect Dis Ther
Abstract
Enteropathogenic Escherichia coli, commonly known as EPEC is a diarrheal pathogen that infects infants in developing countries. The bacterium belongs to the attaching and effacing (A/E) morphotype of pathogenic E. coli, since it infects infants by directly binding to intestinal epithelial cells and destroying the microvilli. The virulence of EPEC is attributed to its major pathogenicity island: The locus of enterocyte effacement (LEE). Treatment of EPEC infections is particularly challenging, because currently there are no vaccines against this bacterium. The problem is only exacerbated by the emergence of multi-drug resistant strains of EPEC. Thus, understanding the regulatory pathways that govern the LEE is critical towards the development of effective measures to combat EPEC infections. The LEE is responsive to a myriad of environmental cues with the majority of them targeting three LEEencoded transcription factors: Ler, GrlR, and GrlA. Whereas transcriptional regulation of the LEE has been widely characterized, post transcriptional regulation including regulation by trans-encoded regulatory small RNAs (sRNAs), remains understudied. Most sRNAs exert their effects by directly base-pairing to their target mRNAs to influence the translation and/or stability of the target mRNA. A subset of these sRNAs requires Hfq, a chaperone protein that assists in the finding and base-pairing of sRNAs to their target mRNAs. One such sRNA is RyhB. Preliminary results suggest that Hfq and RyhB core press the grlRA mRNA that encodes GrlR and GrlA. To better understand the mechanism of action of RyhB on the grlRA mRNA, we preformed in silico alignment analysis. By using IntaRNA, we identified regions of complementarity between RyhB and the ribosomal binding site, in the 5��? untranslated region (UTR), of the upstream gene grlR in the grlRA mRNA. In order to confirm this prediction of direct base-pairing between RyhB and grlRA, we constructed a polynucleotide mutation in the seed region of RyhB; this mutation completely abolished the ability of the mutant RyhB to base pair to and represses the grlR-lacZ fusion. Thus, collectively, our results suggest that RyhB represses the LEE by directing base-pairing to the leader segment of the grlRA mRNA and preventing the expression of both GrlR and GrlA. Future studies are aimed at further understanding the RyhB-mediated regulation of the LEE by genetic, biochemical and phenotypic assays to with the goal of developing efficacious and potent pharmacological targets against EPEC.Biography
Marisa Egan is affiliated to Saint Joseph’s University, USA.
Email: sbhatt@sju.edu

 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi