Our Group organises 3000+ Global Conferenceseries Events every year across USA, Europe & Asia with support from 1000 more scientific Societies and Publishes 700+ Open Access Journals which contains over 50000 eminent personalities, reputed scientists as editorial board members.
Open Access Journals gaining more Readers and Citations
700 Journals and 15,000,000 Readers Each Journal is getting 25,000+ Readers
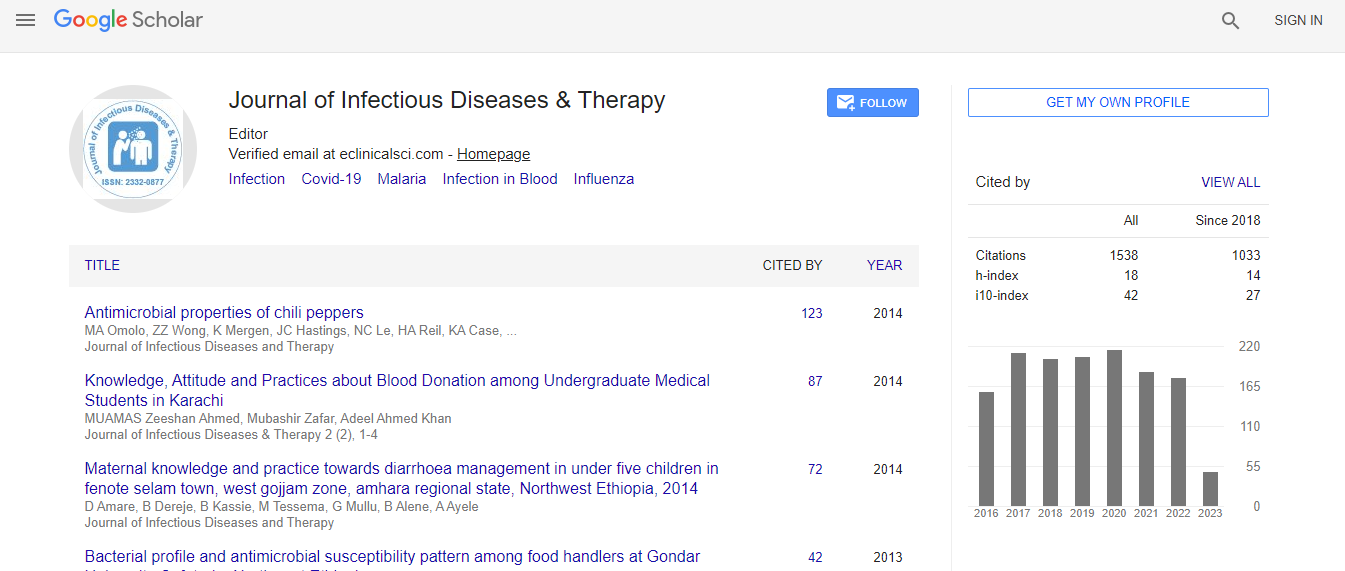
Google Scholar citation report
Citations : 1529
Journal of Infectious Diseases & Therapy received 1529 citations as per Google Scholar report
Indexed In
- Index Copernicus
- Google Scholar
- Open J Gate
- RefSeek
- Hamdard University
- EBSCO A-Z
- OCLC- WorldCat
- Publons
- Euro Pub
- ICMJE
Useful Links
Recommended Journals
Related Subjects
Share This Page
Multiple regulatory small RNAs control virulence in enteropathogenic Escherichia coli
Joint Event on 2nd World Congress on Infectious Diseases & International Conference on Pediatric Care & Pediatric Infectious Diseases
Shantanu Bhatt, Marisa Egan, Valerie Jenkins, Thomas Buerkert, Jasmine Ramirez, Christian Xander, Jamie Palmer and Elizabeth Storm
Saint Joseph��?s University, USA
ScientificTracks Abstracts: J Infect Dis Ther
Abstract
Enteropathogenic Escherichia coli (EPEC) is responsible for considerable disease and death amongst infants in developing countries. EPEC belongs to the attaching and effacing (A/E) family of bacterial pathogens, which are aptly named because they adhere intimately to intestinal cells and destroy cellular microvilli to form characteristic pathomorphological A/E lesions, which lead to diarrhea and dehydration. The ability to form A/E lesions resides within the virulence module locus of enterocyte effacement (LEE), which encodes a type III secretion system (T3SS). To date, over fifty non-LEE encoded regulators have been identified in EPEC. The vast majority of these regulators affect the expression of the LEE-encoded transcription factors, Ler, GrlR and GrlA. Intriguingly, every regulator of the LEE that has been identified to date in EPEC is a proteinaceous factor. Thus far, not a single regulatory small RNA (sRNA) has been implicated in its virulence. We set out to identify and characterize sRNA regulators of the LEE in EPEC. Our preliminary data suggest that Hfq globally represses gene expression from all the LEE-encoded genes including the grlRA operon. Because Hfq and Hfq-dependent sRNAs typically target the 5â�?�? region of the first gene in an operon, we constructed a reporter E. coli strain in which only the 5â�?�? UTR and 45 nucleotides of the grlR ORF were fused to a chromosomal N-terminally truncated lacZ gene driven by the heterologous ParaBAD promoter (ParaBAD-grlR-lacZ). Inactivation of hfq resulted in elevated �?²-galactosidase activity from the grlRâ�?�?-â�?�?lacZ fusion suggesting that the cloned 5â�?�? region of grlR was sufficient to elicit Hfq-dependent repression. These results also suggest that one or more Hfq-dependent sRNAs, conserved between EPEC and E. coli, regulate grlRA. To identify these sRNAs, each of the 27 conserved Hfq-dependent sRNAs was individually overproduced in the grlR-lacZ reporter strain. Three sRNAs-MgrR, RyhB and McaS reproducibly repressed the grlR-lacZ fusion. Using IntaRNA we aligned each of the 3 sRNAs to the cloned 5â�?�? region of grlR. Bioinformatic analysis revealed that MgrR exhibited the most extensive and contiguous region of complementarity (~10 bp) with the grlR leader region. The predicted base-pairing region in MgrR was substituted with a scrambled oligonucleotide sequence that lacks complementarity to grlR. Predictably, mutation of the base-pairing region abolished the ability of MgrR to pair to and repress the grlR-lacZ fusion, thereby providing genetic evidence for direct base-pairing between grlR and MgrR. Subsequent experiments revealed that MgrR binds to the same region on the grlRA transcript as CsrA and counteracts its stimulatory effect. Meanwhile, RyhB appeared to form a relatively shorter (~6 bp) duplex with the ribosome-binding site of grlR. An oligonucleotide substitution in the base-pairing region of RyhB also prevented the sRNA from repressing the grlR-lacZ fusion, suggesting that RyhB, like MgrR, base pairs to the 5â�?�? region of grlR. In contrast to MgrR and RyhB, McaS did not possess any regions of complementarity to grlR and presumably exerts its effect by sequestering CsrA. In summary, our results provide the first piece of evidence to implicate multiple Hfq-dependent sRNAs in controlling the LEE-encoded virulence of EPEC. Future studies are aimed at elucidating the molecular mechanism by which MgrR, RyhB and McaS regulate the LEE and the ensuing A/E lesion formation.Biography
Shantanu Bhatt is affiliated to Saint Joseph’s University, USA.
Email: sbhatt@sju.edu

 Spanish
Spanish  Chinese
Chinese  Russian
Russian  German
German  French
French  Japanese
Japanese  Portuguese
Portuguese  Hindi
Hindi