Commentary Open Access
Computation in Analyzing Inflammation: A General Perspective
Amit Kumar Banerjee*Biology Division, CSIR-Indian Institute of Chemical Technology, Uppal Road, Tarnaka, Hyderabad-500007, Telangana, India
- *Corresponding Author:
- Amit Kumar Banerjee
Biology Division, CSIR-Indian Institute of Chemical Technology
Uppal Road, Tarnaka, Hyderabad-500007, Telangana, India
Tel: +91-4027191361
E-mail: amit_informatics@yahoo.com
Received date: April 15, 2015 Accepted date: April 27, 2015 Published date: April 30, 2015
Citation: Banerjee AK (2015) Computation in Analyzing Inflammation: A General Perspective. Interdiscip J Microinflammation 2:130. doi: 10.4172/2381-8727.1000130
Copyright: © 2015 Banerjee AK, This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at International Journal of Inflammation, Cancer and Integrative Therapy
Introduction
Immunology has grown as a great subject over the years and has evolved as an amalgamation of multiple disciplines and allied sub disciplines as per scientific investigation is concerned. Almost all living being responds towards the stimulation caused by their internal and external surrounding environment in a stepwise manner. Biologically, this is done through a number of cascading physiological and biochemical phenomena. Depending on the specificity, emergency and other important factors, the type of responses alters. Understanding these complex phenomena has been a challenge for the global scientific community since long time.
Immunological reactions claim its own importance due to its association with survival of the individual. Natural process of combating invading pathogens or self-healing after accidents raised the curiosity of the global scientific community to understand the phenomena in detail and boost up the procedure for better healing and cure. Involvement of multiple factors and their complex interactions did not allow us so far to understand the situations completely as it differs greatly from case to case. Considering all the physiological, biochemical, environmental and other factors at a time and understanding their individual impact during the response is a cumbersome task. Requirement of alternative approach is the demand of the hours.
Inflammation
Inflammation, under the light of science could be understood by two means. Information related to the signaling pathways plays pivotal role in invoking inflammatory response and anti-inflammatory pathways involved in diminishing the event of inflammation. In natural cases, the process of inflammation is controlled by innate immunity which follows identification of the microbial pathogens, triggering the defense mechanism against the pathogen through inflammatory cells and subsequent actions depending on the requirement. Any infection, mostly associated with inflammation, ultimately results in different effect such as restoration of damaged tissue architecture, chronic inflammatory response, appended autoimmune response, tissue scarring etc.
Identification and targeting the invading pathogens could be done through various receptors such as Toll like receptors (TLRs), Lectins(C-type), Integrins, cytokines etc. Available literature information revealed that several signaling pathways take part in invoking and regulating the responses related to inflammation such as TIR(Toll-interleukin-1 receptor) signaling cascade, IFN (Interferons) pathway, TAK1(Transforming growth factor-β Activated Kinase-1) pathway, NFκB (Nuclear Factor kappa-light-chain-enhancer of B cells) and MAP Kinase (Mitogen-Activated Protein Kinases) pathway, Caspase pathway [1]. Diminishing the inflammatory responses through prostaglandin, TGFβ and other molecules is done naturally by the innate immune system based on the requirement. Therefore, involvement of many factors, pathways and types of response raised the complexity of the overall process.
Emergence of System Biology
Computational and system biology emerged for such complex biological problems and aided several reliable, timely and logical near truth solutions so far. Evidence of application of such computational, mathematical and statistical approaches is available in ample amount in the literature [2,3]. Mathematical and statistical approaches have proved eminence in simplifying, modeling and understanding several biological phenomena and extract several important interrelations of biomolecules. System biological methodologies upgraded the scenario one step ahead and are being used for understanding various vital parametric interrelations of biological components and biological networks, such as, gene, protein, disease [4,5], microbiome and other important and highly complex networks. Intriguing investigation incorporating experimental and computational methods or system biology are providing solutions to several unresolved queries related to the complex biological phenomena [6,7].
Advancement in both experimental and theoretical techniques and emergence of system biology as a whole enabled us to deep investigate complex biological phenomenon from a vantage point. Quantitative and qualitative analyses are now possible due to the availability of large generated data pool and massive analysis programs. Large problem describing and framing computational platforms such as SBML [8], SBEAMS [9] and visualization and analytical tools such as CYTOSCAPE [10], PAJEK [11] made the feasibility for understanding the negative or positive impact of each component on a whole system. Computation with respect to immune system modeling and inflammation has gained momentum due to the enormous possibility in exploring multiple unanswered questions [12-15]. In the process of understanding immune system, artificial immune system and immune network algorithm was developed which mimics the learning and memorizing capability of natural immune system and is being applied in diverse problems.
Example analysis of the proteins involved in inflammatory responses
It is crucial and present need to understand the protein-protein interactions for inter and intra pathway connectivity. As mentioned earlier, various pathways are involved in either inflammation induction or reduction. Moreover, this scenario may alter from species to species, genus to genus or even between individuals belonging to the same species.
Sophisticated system biological approach allows us to compare and gain insight in such cases and aid in focusing on the relevant problem in consideration. Estimating the involvement of all proteins in any of the inflammation associated pathway is cumbersome through experimentation. Employing statistical association measures it is feasible to narrow down the vast number of associated proteins for a particular pathway and extract most relevant information.
A general analysis was performed considering the major proteins in the pathways involved in invoking inflammation (Interleukin10, Caspase pathway, NFκB and Toll like receptors) and those which play role in diminishing inflammation (Prostaglandin). The ten most relevant proteins which may play role in protein-protein interaction are identified through STRING [16,17] algorithm. In few cases comparison was done between different species to find the resemblances and variations among the associated proteins.
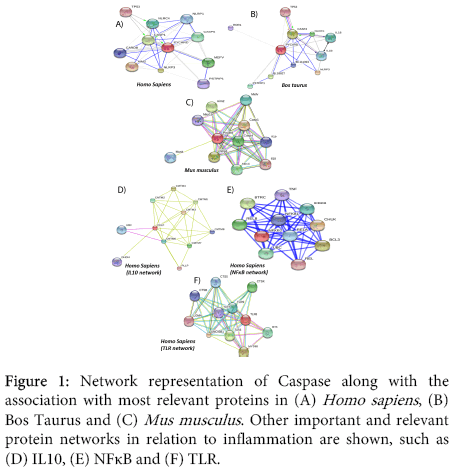
Information on important pathway proteins associated in Caspase pathway (Figures 1A-1C) and other proteins and their respective possible network formation such as IL10, NFκB and Toll Like receptor in Homo sapiens which are related to inflammation were analyzed and visualized (Figure 1). The obtained network represents the complexity in interaction among the associated protein in human.
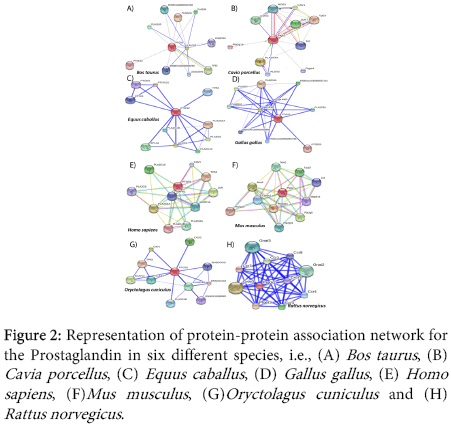
Prostaglandin is a known hormone like lipid compound which has proven role in diminishing inflammation. Exact relevant proteins associated functionally are not known in different organisms. Obtained data represents the higher degree of association of prostaglandins with different proteins in different species (Figure 2). The varying complexity of the network and existence of different type of proteins in each network hints towards the higher degree of complexity. Analysis of individual cases and mathematically modeling certain aspects would be beneficial rather imposing a general mandate for each situation.
Depiction of the proteins associated intensely with Prostaglandin in Homo sapiens, Bos taurus and Mus musculus are represented here (Figure 2).Varying degree of association with diversity in interacting protein molecules is evident from the obtained results. An example of the direction of interrelation of the proteins in Cavia porcellus is provided (Figure 2B), whereas, for Gallus gallus (Figure 2D) and Rattus norvegicus (Figure 2H) representation of the network strength is shown. Thus various modes of representation and visual information yields in better understanding of the interactions.
Knowing inflammation through computation
Inflammations induced through injury are of two major types, intentional, that is due to surgical issues and unintentional or accidental injury. Sudden or induced onset of inflammation includes different signaling pathways leading to the inflammatory response involving numerous factors. In this situation, accounting the cause and effect of the inflammation is important where failing to adopt crucial decision on time may lead to death of the patient.
Estimation of such cause and effect relation for inflammation through traditional medical procedure is cumbersome and so far does not have any standard protocol. Therefore, interdisciplinary approach, especially implementation of predictive measures and assessing their possible outcomes are important [18]. Hence, application of advanced computational methodologies to predict and estimate the relevant issues adhering to inflammation qualitatively and quantitatively is necessary. To compute qualitatively, agent based modeling is predominant in providing reliable solution whereas quantitative calculations are done applying differential equations or partial differential equations.
Gene regulatory networks in inflammation
Studies have been conducted in the recent past on modeling inflammatory events in natural conditions involving multiple signaling pathways. In parallel, computational analysis have been done considering accidental cases where the impact of each component is sudden and ever dynamic. Advanced modeling using experimental datasets and bioinformatics applications helped in construction of gene regulatory network associated with inflammation [19]. This kind of effort may aid in collecting and understanding the overall contribution of the genes in relation to inflammation.
Sudden injury and acute inflammation
Acute inflammation is another important aspect where on spot decision from medical science is vital. Sudden and general burn conditions due to accident are a cause of concern for emergency medicine discipline. Although, depending on the degrees of burning treatments are available but critical cases are requiring more advanced and precise approach. Thermal injury induced models considering interaction of components at different stages of pro-inflammatory conditions and quantitation of the dynamics of NFκB and transcriptional responses has been reported earlier [18]. Certain efforts yielded information in understanding the non-linearity with respect to inflammation, distinguishing effective and unresponsive functionality of the respective responses and impact of application of specific antagonist depending on the circumstances.
Tracking acute inflammatory response
Acute inflammatory response due to trauma or other incidence invokes simultaneously at multiple physiological levels. Experimental analysis to mimic such incidence in animal models had not yielded much information. Mathematical modeling could be a better alternative in such cases where all the impacting variables could be assessed individually or collectively. The incidence of acute inflammation in animals has been computed and represented through mathematical models in the study reported by Vodovotz et al. where experimental information has been used as reference. The outcome received in this study, turned out to be satisfactory [20]. Assessment of dosage variation of lipopolysaccharide, surgical trauma with relevant factors and associated hemorrhagic shock was experimentally and theoretically evaluated and the obtained results showed promises in future reduction of animal use for such studies.
Computing pro and anti-inflammatory signaling and vocal injury
Macrophages play an important role in inflammation reduction. Therefore, understanding the signaling mechanism related to the activity of macrophage is essential. Experimental investigations revealed that several signaling pathways are involved in this process, such as, TNF-α, interleukin-10, cytokines, nuclear factors, etc. Only experimental analysis may not provide the detail information in this regard. Poor expression of proteins in the cell may not yield proper information about the impact of different proteins in the pathway cascade and related feedback mechanism. Through computational modeling, it is possible to understand the up regulation and down regulation of such proteins along with their combined effects. Existing reports in this regard provided valuable outlook on the feedback mechanism and relevant impact for certain factors, such as, TNF-α, IL10, cytokines and others in relation to pro and anti-inflammatory signaling [21]. Modeling and analysis of multiple up regulation and down regulation associated component of inflammatory signals may yield a better and clear perspective about the network components and corresponding interactions.
Vocal injury and its connection to inflammation is a common issue in all population. In silico model has been developed considering the individual patient-specific conditions and healing aspects for acute cases of vocal injury. Agent based modeling (ABM) has been implemented for simulating cause and effect of mediators involved in laryngeal secretions. Computational experimentation related to patient specific conditions showed variation in trajectories generated through computation, hinting towards effective and possible future application of mathematical modeling techniques. Successful application of computational analytical procedures in a specific manner may support the objective of achieving personalized medicine in near future [22].
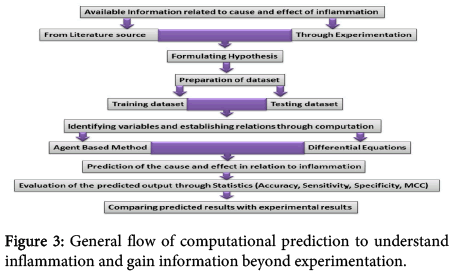
System biology based approaches helped in understanding diseases and other intricate biological events which cannot be known easily through individual traditional experimentation. Expansion of system biology has paved the path towards personalized medicine where individual large scale modeling is possible. Although, to gain further insight regarding inflammation through computation, no standard computational protocol is available so far, yet the strategies to design and analyze the experiments follow some general procedure (Figure 3).
Conclusion
Initiated with estimation of single parameter, mathematical modeling and computational techniques are playing major role in all aspect of scientific discipline including intriguing and dynamic biological problems. Advanced methods along with the support from sophisticated software and hardware are being used in the present days to solve complex problems. Moreover, improvement in bulk data generation methodologies yielded extreme opportunities to theoretical experimentation and allowed us to reach near the real time solutions. Evolution in computational techniques in relation to big data analysis, gigantic data representation and visualization aided ample opportunities to analyze large and dynamic problems collectively. Development of system biology was possible due to this revolution in information technology. Classical computational, statistical or mathematical approaches generally deal with top down approach where surface to deep level analysis is feasible. Era of system biology and personalized medicine altered the efforts and thinking of vast computation and follows a bottom up approach to understand a complex cascade of biological events. In this approach, numerous variables are considered simultaneously and are being treated differently case to case basis, hence supporting the concept of personalized medicine through computation. We are in vicinity to the success related to personalized medicine along with understanding immune system and inflammation applying theoretical techniques. System biological analysis combined with high throughput experimental protocols may aid in understanding each aspect individually within a limited and appropriate time frame.
Acknowledgements
I would like to acknowledge the tremendous efforts of those researchers who are continuing to understand the immune system applying computational approach.
References
- Larsen GL, Henson PM (1983) Mediators of Inflammation. Annu. Rev. Immunol. 1: 335-359.
- Banerjee AK, Ravi V, Murty US, Shanbhag AP, Prasanna VL (2013) Keratin protein property based classification of mammals and non-mammals using machine learning techniques. Comput. Biol. Med. 43(7):889-899.
- Banerjee AK, Ravi V, Murty US, Sengupta N, Karuna B (2013) Application of intelligent techniques for classification of bacteria using protein sequence-derived features. ApplBiochemBiotechnol. 170(6):1263-1281.
- Sonachalam M, Shen J, Huang H, Wu X (2012) Systems biology approach to identify gene network signatures for colorectal cancer. Front Genet.3:80.
- Ideker T, Sharan R (2008) Protein networks in disease. Genome Res. 18: 644-652.
- Murty USN, Amit KB, Neelima A (2009) An In Silico Approach to Cluster CAM Kinase Protein Sequences. J Proteomics Bioinform 2: 97-107.
- Amit KB, Neelima A, Varakantham P, Murty USN (2008) Exploring the Interplay of Sequence and Structural Features in Determiming the Flexibility of AGC Kinase Protein Family : A Bioinformatics Approach. J Proteomics Bioinform 1: 77-89.
- Hucka M, Finney A, Sauro HM, Bolouri H, Doyle JC, Kitano H, Arkin AP et al. (2003) The systems biology markup language (SBML): A medium for representation and exchange of biochemical network models. Bioinformatics 19 (4): 524–531.
- Marzolf B, Deutsch EW, Moss P, Campbell D, Johnson MH, Galitski T (2006) SBEAMS-Microarray: database software supporting genomic expression analyses for systems biology. BMC Bioinformatics 7:286.
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. (2003) Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 13(11): 2498–2504.
- Batagelj V, Mrvar A (2014) Pajek. Encyclopedia of Social Network Analysis and Mining. Springer New York, ISBN: 978-1-4614-6169-2, pp 1245-1256
- Forrest S, Beauchemin C (2007) Computer immunology. Immunol. Rev. 216:176-197.
- Zak DE, Aderem A (2009) Systems biology of innate immunity. Immunol. Rev. 227:264-282.
- Perelson AS (1989) Immune Network Theory. Immunol. Rev. 110:5-36.
- Kirschner DE, Chang ST, Riggs TW, Perry N, Linderman JJ (2007) Toward a multiscale model of antigen presentation in immunity. Immunol. Rev. 216(1):93-118.
- Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43(Database issue):D447-52.
- Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, et al. (2013) STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41(Database issue):D808-15.
- Yang Q, Berthiaume F, and Androulakis IP (2011) A Quantitative Model of Thermal Injury-Induced Acute Inflammation. Math. Biosci. 229(2):135–148.
- Chen BS, Yang SK, Lan CY, Chuang YJ (2008) A systems biology approach to construct the gene regulatory network of systemic inflammation via microarray and databases mining. BMC Med Genomics. 1:46
- Vodovotz Y, Chow CC, Bartels J, Lagoa C, Prince JM, Levy RM, et al.(2006) In silico models of acute inflammation in animals. Shock 26: 235-44.
- Maiti S, Dai W, Alaniz RC, Hahn J, Jayaraman A (2015) Mathematical Modeling of Pro- and Anti-Inflammatory Signaling in Macrophages. Processes 3: 1-18.
- Li NYK, Verdolini K, Clermont G, Mi Q, Rubinstein EN, et al. (2008) A Patient-Specific in silico Model of Inflammation and Healing Tested in Acute Vocal Fold Injury. PLoS ONE 3: e2789.
Relevant Topics
Recommended Journals
- Journal of Lung Cancer Diagnosis & Treatment
- Advances in Cancer Prevention
- Breast Cancer: Current Research
- Cancer Surgery
- Immunology: Current Research
- Current Trend in Gynecologic Oncology
- Journal of Cancer Diagnosis
- Journal of Gastrointestinal Cancer and Stromal Tumors
- Cervical Cancer: Open Access
- Journal of Mucosal Immunology Research
- Journal of Oncology Research and Treatment
- Journal of Orthopedic Oncology
- Journal of Prostate Cancer
- Research and Reviews on Pathogens
Article Tools
Article Usage
- Total views: 16104
- [From(publication date):
July-2016 - Aug 19, 2025] - Breakdown by view type
- HTML page views : 11334
- PDF downloads : 4770