Research Article Open Access
Response of Rice under Salinity Stress: A Review Update
Bhaswati Ghosh, Nasim Ali Md* and Saikat GantaitFaculty Centre for Integrated Rural Development and Management, School of Agriculture and Rural Development, Ramakrishna Mission Vivekananda University, Ramakrishna Mission Ashrama, Narendrapur, Kolkata 700103, India
- Corresponding Author:
- Nasim Ali Md
Faculty Centre for Integrated Rural Development and Management
School of Agriculture and Rural Development
Ramakrishna Mission Vivekananda University
Ramakrishna Mission Ashrama, Narendrapur, Kolkata700103, India
Tel: +91-9831941312
E-mail: nasimali2007@gmail.com
Received date: March 07, 2016; Accepted date: March 24, 2016; Published date: March 26, 2016
Citation: Ghosh B, Ali Md N, Saikat G (2016) Response of Rice under Salinity Stress: A Review Update. J Res Rice 4:167. doi: 10.4172/2375-4338.1000167
Copyright: © 2016 Ghosh B, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Rice Research: Open Access
Abstract
Salinity has been a key abiotic constraint devastating crop production worldwide. Attempts in understanding salt tolerance mechanisms has revealed several key enzymes and altered biochemical pathways inferring resistance to crop plants against salt stress. The past decades have witnessed extensive research in development of salt tolerant cultivars via conventional means, improvised by modern era molecular tools and techniques. Rice (Oryza sativa L) is the staple food crop across several countries worldwide. Being a glycophyte by nature, its growth is severely imparted in presence of excess salt. Rice is susceptible to salinity specifically at the early vegetative and later reproductive stages and the response of the crop to excessive salt toxicity at biochemical and molecular level as well as physiological level is well studied and documented. An understanding of the specific response of rice to ion accumulation at the toxic level can aid in identifying the key factors responsible for retarded growth and limited production of rice with the future scope of mitigating the same. The present review summarizes the differential responses of rice, in particular, to salt toxicity enumerating the detailed morphological, physiological, biochemical and molecular changes occurring in the plant. An attempt to explain salinity tolerance and its future scope and implications in screening for salt tolerance has also been elucidated in the present study.
Keywords
Salinity; Biochemical marker; Chlorophyll; Oryza sativa L; morphological descriptor
Abbreviations
CHL: Chlorophyll, EC: Electric Conductivity, PCA: Principal Component Analysis, POD: Peroxide Dismutase, RWC: Relative Water Content, SOD: Superoxide Dismutase, STI: Salt Tolerance Index, UPGMA: Unweighted Pair Group Method Arithmetic Average
Introduction
Rice, the major food crop
Rice (Oryza sativa L) belongs to the family Poaceae. The basic chromosome number of rice is n=12. The species can be either diploid or tetraploid. In this respect, Oryza sativa L. and Oryza glaberrima L both are diploid species (2n= 24) [1]. The Asian cultivated rice (Oryza sativa L) is the first fully sequenced crop genome and is a model crop species. Rice is considered as a major food crop across major countries worldwide. As a food crop, it forms the staple food of more than three billion people accounting for about 50-80% of their daily calorie intake [2]. It yields about one third of the total carbohydrate source. It provides the considerable amount of recommended Zinc and Niacin [3]. Rice protein is biologically richest as its digestibility is very high (88%). It is the 2nd most important crop in the world after wheat, covering almost 90% of area across Asia alone. The use of the crop varies widely ranging from its use as food in cereals, snacks, brewed beverages, flour, rice bran oil to its use in religious events across India. The medicinal value of the crop adds on more to the list.
Rice cultivation has been predominant in India across ages. Rice covers a global area of 156 million hectares of land producing about 650 million tons of crop [4]. Globally covering an area of 149.15 million ha area yielding about 550.19 million tones, this food crop is being cultivated across an area of 44.6 million ha in India. India ranks first in area and second in production following China, the largest producer of rice. As an economically and industrially important crop of India, rice provides about 23% of total world rice production and 45% of the total Indian food grain production. However with the expanding population, the increase in production of the crop is the urgent need of the hour in order to keep in accord to the national food and livelihood security system. India is one of the leading exporters of rice, in specific, basmati rice.
Salinity: an overview
Salinity as an abiotic stress widely limit the crop production severely [5]. A saline soil is usually the reservoir of a number of soluble salts such as Ca2+, Mg2+, Na+ and anions SO42-, Cl-, HCO3- with exceptional amounts of K+, CO32-, and NO3-. A soil can be termed as saline if its EC is 4 dS/m or more [6], (equivalent to approximately 40 mM NaCl) with an osmotic pressure of approximately 0.2 MPa. Salinity is the condition when the EC is sufficient to cause yield reduction of most crops. The pH of saline soils generally ranges from 7-8.5 [7]. However, the pH in saturated soil can vary provoking severe crop damage (Table 1). The arid and semi-arid zones, characterized by low precipitation and high evaporation are the most affected due to minimum lixiviation of salt from the soil profile resulting in increased salt accumulation. Salinity prone areas found in the arid and semiarid zones are usually accounted to the accumulation of salts over ages. Moreover, weathering of the parental rocks has accelerated the process a lot [8,9]. Salinity is a well off natural phenomenon occurring near sea shores due to sea water flooding.
| Electrical conductivity (dS m-1 at 25°C) | Crop response |
|---|---|
| 0-2 | Salinity effect is practically zero |
| 4-Feb | Reduction in yield of very sensitive crops |
| 8-Apr | Reduction in yield of most crops |
| 16-Aug | Only tolerant crops produce satisfactory yield |
| >16 | Few highly tolerant crops produce satisfactory |
Table 1: Crop response to salinity influenced by electrical conductivity of saturated soil extract Source: Adapted from [7].
However, use of irrigation and land clearing can be listed as one of the many well-known anthropogenic reasons behind salinity [10].
Salinity has been a potential threat affecting almost 900 million ha of land which nearly accounts for 20% of the globally cultivated area and also half of the total irrigated land of the world [11,12]. Globally salt affected area accounts to about 1 billion ha of land [13]. In India the scenario accounts for about 8.4 million ha land affected by salinity [14]. In India, irrigated land accounts for only 15% of total cultivated land, nevertheless, it has at least twice the productivity of rain-fed land and yields about one third of the world’s consumption. In accordance to the above facts and keeping in view the present alarming scenario, development of salt tolerant genotypes is definitely an urgent need of the hour.
Salinity: A key abiotic constraint to production
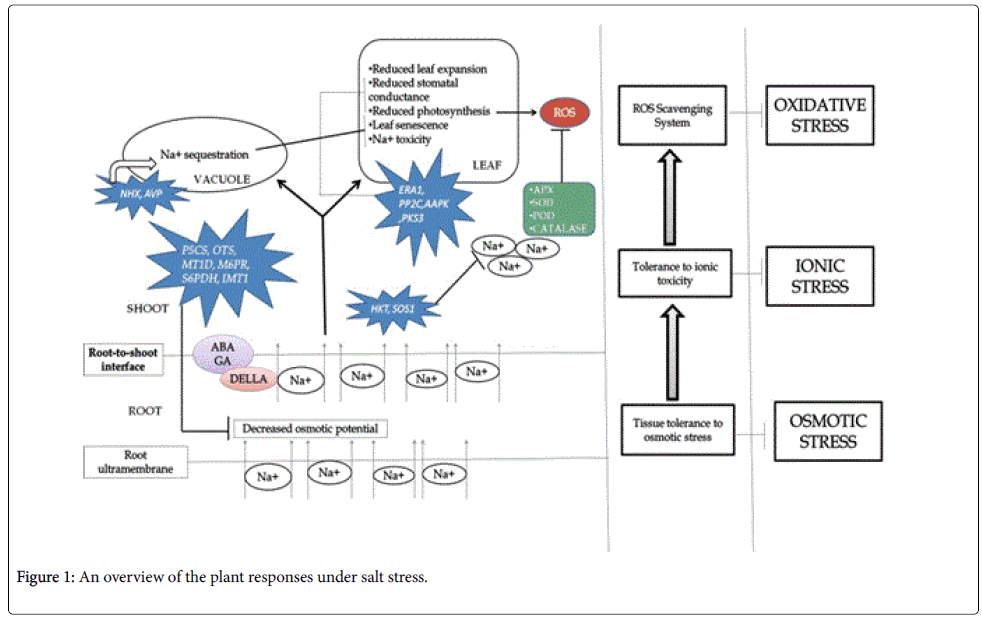
Soil salinity is one of major abiotic stress devastating agricultural production worldwide (Figure 1).
Globally, salt affected area accounts for about 800 million ha of land. Interestingly, this figure covers more than 6% of total global area. Of the 1500 million ha of dry land farming, 32 million ha (2%) are accounted to be salt effected at varying degrees by secondary salinity and to date more than 45 million ha (20%) of the total irrigated land (accounting about 230 million ha) is saline [4].
Soil salinity as an abiotic stress can be traced back in time even when human civilization did never existed or did not even started agriculture as an occupation. However, the devastating effect of it has been aggravated with the practice of irrigation and cultivation.
The effects of salinity in plants are many and interfere greatly with the yield and production of the crops. Salinity affect the plants mainly by imparting two types of stresses: Osmotic Stress (initial stage: caused due to increased osmotic potential of rhizosphere due to high salt concentration) and Ionic Stress (final stage: toxicity resulted by high ionic concentration). The destructive effects of salinity include retarded plant growth due to increased Na+ concentration [15], delay in flowering and impaired fertility, with partial or complete grain loss resulting in poor panicle development in rice [16-19], reduced P3+,K+ and Ca2+ concentrations [13], inhibition of photosynthetic activity [20-23].
Plants vary greatly in their response the salinity. The variation in response is random as salinity is a polygenic trait. NaCl being the most soluble and predominant salt, almost all plants have adapted mechanisms to regulate its concentration. It may be via preferential selection of ions, such as K+ and NO3- commonly present in low concentrations or via effective exclusion of Na+ and Cl- by roots [24]. Halophytes, the natural flora of highly saline soils evolved against glycophytes by this exclusion principle mainly. For example, sea barley grass, Hordeum marinum, excludes both Na+ and Cl- only after it reaches the threshold of 450 mM NaCl [25]. An exclusive salt tolerance mechanism (presence of vesiculated hairs on their surface also known as trichomes) has also been reported in Atriplex halimus, a well known salt tolerant species, for adaptation to salinity stress by accumulating excess salts inside the hairs and thereby protecting the leaf parenchyma and vascular tissues [26].
The differential tolerance to salt toxicity in cereals ranges from barley (Hordium vulgaere ) being the most tolerant followed by Bread Wheat (Triticum aestivum), durum Wheat (Triticum turgidum ssp.drum) with rice being the most sensitive (Oryza sativa) [10]. Strikingly this difference in response is much more in dicotyledonous relatives than in the monocots.
A study of salt tolerance mechanism has been possible with a comparative analysis of the patterns of response between a wild type and the natural halophytic relative. In this regards, light has been thrown into the mechanism of salinity tolerance when Arabidopsis thaliana was being compared with its wild halophytic relative Thellungiella halophilla. The contrasting responses highlighted their selective tolerance to salt: Arabidopsis ceased its growth in presence of the constant exposure to 100 mM salt whereas [27] endured almost little or no effect on the growth rate of Thellungiella [28].
Effect of salinity stress in rice crops
Rice is a crop with great economic importance [29,30] and is cultivated across 114 countries globally [31]. However, the abiotic and biotic stresses can reduce its yield. This problem will be worse in attention to increase of global population and food sources deficiency [32]. Rice is susceptible to salinity, specifically, at the early vegetative and later reproductive stages [5,33]. Rice genotypes show wide variations in salinity tolerance due to additive gene effects [34]. Studies indicated that rice is more resistant at reproductive and grain filling than at germination and vegetative stages [35], as well as low levels of salinity can increase the resistance of rice to higher and lethal salinity levels [36]. At present, salinity is the second type of stress and is the most predominant hindrance to rice production after drought [37]. The effects of salinity on the growth and yield of rice in field have been well studied including the study of genotypic variance for salt tolerance amongst the paddy germplasms [38-40].
Response of rice under salinity
With the aim of generating salt-tolerant rice cultivars, several morpho-physiological studies have been accomplished till date. The main focus of such an approach was to maximize the genetic diversity between parental genotypes. Response of plants to salinity is a random and natural process. Rice as a crop has been reported as sensitive in the seedling and reproductive stages [23,41] and has resulted in reduction of crop yield and productivity [16]. In case of rice, salinity is found to induce both biochemical and physiological changes causing growth inhibition and yield loss [19,42-45]. Several physiological parameters have been studied in the process of evaluation of true salt tolerant lines to understand the drastic effects of salt accumulation at physiological level: plant height, plant dry weight, leaf injury and Na+-K+ ratio [37,46-49].
The effect of salinity on rice is many fold, leading to inhibition of germination, difficulties in crop area establishment, leaf area development, decrease in dry matter production, delay in seed set and also even sterility can occur [40,50]. It has been well documented that the effect of salinity on seedling growth, seedling establishment, grain yield components such as spikelet number, tiller number has successively lead to a reduction in grain yield [51-54]. A wide range of variation has been reported between and within different rice varieties in response to salinity [50,55] checked for varying floodwater salinity tolerance among a number of paddy varieties and reported that floodwater electrical conductivity (EC) reduced germination rate by almost 50% and yield by 80% for the most susceptible cultivar in accordance to the highest salinity level imposed. Further, salinity also resulted in a decrease of the spikelet number per panicle, 1000 grain weight and increased sterility, regardless of the season and development stage.
Morpho-physiological response
Resistance to salinity stress does not rely on a single trait and thereby understanding of the tolerance mechanisms should employ the study of the response of rice under stress. A study of the response of rice to salinity began with the understanding of the underlying physiological mechanisms associated with plant defense mechanisms being activated during stress. The effect of salinity on plants is initiated by the osmotic effect characterized by lowered osmotic potential followed by later ionic effect causing ion toxicity. Studies conducted to interpret the response of rice at physiological level indicated chloroplast and mitochondria to be the most vulnerably affected organs among others [56]. Hence, chlorophyll content, changes in chlorophyll fluorescence (Fv/Fm) and membrane permeability are efficient and potential indicators for understanding the inhibitory effect of salt on the photosynthetic efficiency [20,57]. Salinity has been reported to decrease leaf area tremendously and also showed profound changes in leaf anatomy in rice grown in-vitro [58] or in greenhouse [59] as verified by [60]. The ultra-structural observations briefed by [56] also ensured the inhibitory effect of salt on leaf thus hampering the photosynthetic efficiency: swelling of thylakoids followed by disruption of chloroplastids. Salinity was observed to exert severe detrimental effect on the mesophyll tissue even stretching its harmful effects to the vascular bundles. Evaluation of the response of the crop plant at later stages was to comprehend the toxic ionic effect of salt on the plants. The toxic effect of accumulation of sodium salt are mainly mitigated by the plant via (a) salt exclusion [61,62], (b) selective ion uptake [63,64] and (c) regulation of K+/Na+ ratio [66-69]. A study of the root ultra-structure elucidated the penetrating effect of salinity on rice and the response of the crop plant with an increased rate of vacuolation and vesiculation decreasing mucilage production in treated plants compared to control as observed by [69]. A strong correlation was suggested between sodium content, ratio of K+/ Na+ with seedling growth and grain yield under salt stress [51,52,70]. Evaluation of different cultivars was precised at morpho-physiological level by estimating parameters like tiller number, leaf area, panicle length, root length, biomass, dry weight, RGR (Relative Growth Rate), RWC (Relative Water Content) [16,52,70-72]. Yancey et al. [73] observed an increased level of leaf RWC in paddy under salinity and suggested the role of osmo-protectants in preventing cell injury from salt stress-induced dehydration. An understanding of the STI (Salt tolerance index) evaluating indigenous landraces at the morphobiochemical level has been reported by [49] implicating the importance of landraces as probable sources for de-novo genes inferring tolerance to them as being observed in the study. Thus, the evaluation of different cultivars at morpho-physiological level enabled to build up a comprehensive idea about the several physiological mechanisms operating in the crop plant as a response to salt stress but failed to provide an insight about the exact pathways and defense system and the components being directly or indirectly involved in the process.
Response at the biochemical level
The effect of salinity on crop plants may be categorized as a two-fold process: an initial osmotic effect followed by ionic stress when salt accumulation reaches its toxic level [10]. The initial osmotic effect in plants refers to the lowered water potential to the increased concentration of salt with an enhanced osmotic potential. One of the important plant defenses is the osmotic adjustment in plants accomplished via accumulations of high concentrations of inorganic ions or low molecular weight organic solutes. These compatible osmolytes found in higher plants are actually certain low molecular weight sugars, organic acids, polyols. They also include nitrogen containing compounds such as amino acids, amides, imino acids, proteins and quaternary ammonium compounds. Studies indicated that proline, which occurs widely in higher plants, assimilates in larger amounts in salt stressed plants [39,74,75]. Proline accumulation in rice has also been reported by [76] suggesting its active roles in osmotic adjustment, shielding the enzymes and membranes, also providing energy and nitrogen for utilization during exposure to salinity [77,78].
Soluble sugars and starch has been observed to serve as an osmoticum in a number of plants as a response to salinity [79,80]. An increase in sugar content in shoots has been reported by [81] and also an increased starch content in roots been showed in the roots of rice which contributes to the osmotic adjustment to the crop exposed to salinity by maximizing sufficient storage reserves to prop up the primary metabolism [82,83]. The toxic effects of salinity are masked by accumulation of glycine betaine in rice as being reported by [84]. These compounds are reported to have active role in osmotic adjustment, fortification of the cellular macromolecules, nitrogen storage. They are important to balance the cellular pH, detoxify the cells and scavenge the ROS species.
The other way to respond against salinity is alteration or accumulation of protein level. Exposure to salinity can result in denovo protein synthesis or an up-regulation of the process to increase the concentration of certain proteins already present in the plant [85]. Proteins accumulating in plants grown under saline conditions act as a storage form of nitrogen which is re-utilized in absence of stress [86-88]. Protein synthesis is also destined to play an active role in osmotic adjustment. A significant increase in soluble protein content and positive correlation has been ensured in tolerant rice seedlings compared to the sensitive ones.
Response at molecular level: targeted approach and cellular signaling
Attempts to understand salinity tolerance at the gene or molecular level started with the aim to bred for salinity tolerance and develop salt tolerant lines. Earlier studies on salinity tolerance mechanisms revealed the complexity of salinity as an abiotic stress and that there do exists not only inter-species variation in plant in response to this stress but also inter-varietal differential response as far as rice being concerned as a crop [87-90]. Genetic diversity analysis of several genotypes were thus employed to screen for salinity tolerance by various molecular markers: RFLP and SSLP mapping by [91], RAPD and SSR analysis by [92], morpho-molecular evaluation of landraces by [93]. In order to define the complexity and the nature of the inheritance of salinity in rice, several studies were conducted which included conventional techniques such as positional cloning [94,95] and ‘insertional mutagenesis’ [96,97]. Several genes were identified in rice induced by high salinity levels as elaborated by [98] in the course of monitoring the expression profile of rice under stress, which included genes like salT, catalase and several denovo genes.
The response of plants to combat salinity stress is initiated by their ability to sense the initial osmotic stress or osmo-sensing. Though the physiological mechanisms throw light to the modification taking place in the tissue or cellular level, which is actually measured phenotypically by quantifying the Na+/K+ ratio, however, control at genetic level and the regulatory pathway solely formed the backbone behind understanding of the induced signaling pathway in response to this stress. With an aim to achieve the same, experiments conducted on different mutants of Arabidopsis thaliana, came forth with several regulatory genes and important pathways: Arabidopsis thaliana histidine kinase, ATHK1: candidate osmosensor [99], CDPKs (Ca+ dependent protein kinases: OsCDPK7 being identified in response to salinity in rice by Sajio et al.); MAPKs (Map kinases: OsMAPK5 being identified in rice whose suppression led to hypersensitivity to salinity by [99,100]; SOS pathway (SOS1, SOS2, SOS3:[101]). These multiple genes governing the regulation of the response of the plants to salinity stress ensured the polygenic character of salinity as a trait. Thereby, work initiated to identify QTL (Quantative trait loci) related to salt tolerance. The first breakthrough was achieved by [37] where a major QTL ‘saltol’, has been mapped on chromosome I in an F8 recombinant inbred lines (RIL) population of Pokkali X IR29cross, using amplified fragment length polymorphism (AFLP) markers. In recent years, QTL analysis in rice helped in mapping several QTLs related to several characters correlated with salinity: QTLs mapped on chromosome 1 and 2 for shoot growth [102]; 5 major QTLs: qRL-7 for root length, qDWRO-9a and qDWRO-9b for dry weight root, qBI-1a and qBI-1b for biomass [103]. Separate QTLs being identified each for sodium uptake, potassium uptake, and sodium:potassium selectivity [104]; 8 QTLs accounting each of three for three traits of the shoots, and each of five for four traits of the roots at five chromosomal regions [105] and many more. A total of 35 QTLs were identified by [106] in an F2 mapping population derived from a Sadri/FL478 cross, the major QTL clusters being mapped in chromosomes 2, 4 and 6 for multiple traits under salinity stress. Identification of QTL for salinity tolerance opened a new horizon in the study of salinity and the plant response to cope up with this stress thereafter. Studies are being conducted to formulate and identify different alleles whether associated with the respective QTLs for salinity [107-109].
Defense system of rice against salinity stress
Plant defense mechanisms to combat the toxic effect of salt stress can be categorized into three processes viz. a) Tolerance to osmotic stress via osmotic adjustment, b) Na+ exclusion from leaf blades by selective ion uptake and also regulation of uptake of sodium ions at molecular level, c) Tissue tolerance, i.e. tolerance of tissue to accumulated Na+, or in some species, to Cl− via compartmentalization of absorbed or accumulated salt [10]. In addition to osmotic stress and ionic stress plants are subjected to oxidative stress which is caused mainly due to the inhibitory action of salinity on photosynthesis. In order to cope up with the upcoming photo-inhibitory effects plants undergo modification in their metabolic pathways such as heat debauchery by the xanthophyll pigments and electron transfer to oxygen acceptors (not water) which can result in the formation of ROS (reactive oxygen species). The later response is however mitigated by an initiation of the up regulation of several regulatory enzymes for such as superoxide dismutase, ascorbate peroxidase, catalase, and the various peroxidases [105-109]. The enzymatic antioxidant defense system of plants is inclusive of Superoxide dismutase (SODs), peroxidases, Catalases, and the enzymes of the ascorbate–glutathione cycle: Ascorbate peroxidase (APX), Monodehydro-ascorbate reductase (MDHAR), Dehydro-ascorbate reductase (DHAR), and Glutathionereductase (GR) while non-enzymatic antioxidants include: Ascorbate (AsA) and Glutathione (GSH) [110,111]. The oxidative damage caused is a measure of poise between the formation of ROS and its subsequent removal by the antioxidative scavenging system [109,112]. The active role of the anti oxidative system has also been observed in roots of rice cultivars differing in salt tolerance [113-116]. Sese and Tobita [109] reported a significant decrease of SOD activity in the salt-susceptible varieties whereas an increase in SOD activity in the salt-tolerant variety in response to salt treatment [115]. An increase in peroxidase activity in salt-tolerant cultivars under salt stress has been recorded by various researchers [117-119]. In rice, differential opinions are being established as far as oxidative responses are being concerned. Mishra et al. [120] reported an increase in SOD activity, APX activity and GPX activity, however reporting an decrease in CAT activity with increased exposure to salinity levels. An anticipation of the above results was however elucidated by [121] where an increased CAT activity and decreased SOD and POX activity was observed in salt tolerant lines.
The implication: screening against salinity stress
Differential salinity sensitivity at various growth stages is one of the factors affecting salt tolerance phenotypes. Generally, rice plants is very sensitive to salinity at young seedling stages and less sensitive at reproductive stages [70,52]. To assess the response of germplasms to salinity a proper screening technique is required. In paddy, the screening can be done at its two salt sensitive stages. Screening at seedling stage is a rapid method and based on simple criteria. Screening can be done at field level or in laboratory condition. The former one is difficult because of presence of soil heterogeneity, climatic factors and other environmental factors which may itself interfere with the physiological processes. Environmental influence is a very complicated factor intertangling salt tolerance. Any environmental changes such as humidity and temperature will affect evapo-transpiration and further influence ion transport [5]. This makes the evaluation of the phenotypes at field conditions very difficult. So screening under laboratory condition has many advantages over field screening.
A combined pleotropic effect of different factors alters plant growth and development at three levels of the plants viz. physiological, biochemical and molecular level. Primitive works done ensured screening for salinity tolerance based on agronomic characters. Thus, early experiments conducted on development of salt tolerant genotypes solely relied on conventional selection and breeding techniques [5,74,120-124]. which included screening at the morphological level based on several physiological and agronomical characters.
Morphological screening can be done using the parameters described as effective salinity indices like root length, shoot length, plant biomass, shoot Na+/K+ ratio [38,39,125,126]. The morphological screening can be started at least after 10 days of exposure to saline solution for screening properly the tolerant genotypes from susceptible [37]. However screening for salinity tolerance based on agronomic characters may not prove to be practicable and fruitful technique because they are highly influenced by the environment. The understanding of the salt tolerance mechanisms and furthermore the assessment of salt tolerance is expected to be much more well defined by screening the genotypes at the biochemical level as undertaken by many scientists [75,81,112,119,125,126]. It provided an idea about the potential indicators of salt tolerance at the whole plant, tissue or cellular level. The identification of the metabolic sites affected by salt stress and, conversely, the mechanisms utilized by plants to survive salinity stress has also been well understood. Molecular screening for salt tolerance in rice accounts for the recent approaches for understanding the response of rice under salt stress and thereby mines useful alleles responsible for salt tolerance [91,107,127-129]. The identification of saltol QTL in 2010 by Thomson et al. has widened the horizon for further work and development as far as genetic approach is being concerned [130-135].
Conclusion
Salinity as an abiotic stress is the second most devastating phenomenon after draught and interfering with the production and yield of rice globally. Rice is the major food crop cross across several countries globally. With the increased population worldwide the demand for rice is also increasing in accordance. Rice, a glycophyte, by nature is susceptible to salinity and show wide and vivid response against the detrimental effects of increased salt accumulation. The plant defense system in rice includes arrest and alleviation of the harmful effects of salt toxicity at physiological, biochemical and molecular levels. A comprehensive study of the manifold works done to apprehend the effect of salinity and the response of the crop in turn has been illustrated here. Due to the polygenic nature of the stress, it has been very meticulous to comment in particular about the exact mechanism by which mitigation of the same is achieved. However, a number of metabolic pathways, enzyme complexes, regulatory genes and QTLs have been enumerated till date to throw some light on the various particular responses at various stages of this abiotic stress. With respect to the devastating effect of salinity on this staple food crop, it is very important assimilate the knowledge of the response of the crop towards salinity. This will aid in future work of improvement towards salinity tolerance and breeding for salinity tolerance. A number of works are being conducted across globally to understand salinity and the complex mechanisms of rice to mollify the same. In this regard, it is of prior importance to have an upshot of all the works going on to understand salinity and its effect on rice. This review is a minor attempt to summarize the salient contributions and breakthroughs made in this area in the course of understanding the response and thus the plant defense to fight salinity as a stress.
References
- Brar DS, Khush GS (2003) Utilization of wild species of genus Oryza in rice improvement In: JS Nanda, and SD Sharma (Eds) Monographon Genus Oryza pp: 283-309.
- Khush GS (2005) What will it take to feed 5.0 billion rice consumers in 2030. Plant MolBiol 59: 1-6.
- Gopalan C, Rama Sastri BV, Balasubramanian S (2007) Nutritive Value of Indian Foods, published by National Institute of Nutrition (NIN), ICMR (Indian Council of Medical Research).
- FAO (2008) FAO Land and Plant Nutrition Management Service.
- Shannon MC (1998) Adaptation of plants to salinity. AdvAgron 60: 75-120.
- USDA-ARS (2008) Research Databases Bibliography on Salt Tolerance. George E. Brown, Jr. Salinity Lab. US Dep. Agric., Agric. Res. Serv. Riverside, CA.
- Mengel K, Kirkby EA, Kosegarten H, Appel T (2001) Principles of plant nutrition. Kluwer, Dordrecht.
- Szabolcs I (1989) Salt-Affected Soils Boca Raton, FL: CRC.
- Rengasamy P (2002) Transient salinity and subsoil constraints to dry land farming in Australian sodic soils: an overview. Aust J ExpAgric 42: 351-61.
- Munns R and Tester M (2008) Mechanisms of salinity tolerance. Ann Rev Plant Biol 59: 651-681.
- Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Env 25:239-250.
- FAO (2007) FAO Land and Plant Nutrition Management Service.
- Fageria NK, Stone LF, Santos ABD (2012) Breeding for salinity tolerance. In: R Fritsche-Neto, A Borém (Eds) Plant Breeding for Abiotic Stress Tolerance, Springer-Verlag Berlin, Germany pp: 103-122.
- Tyagi NK, Minta PS (1998) Agricultural salinity management in India.In: NK Tyagi, PS Minhas (Eds.) Agricultural salinity management in India, Central Soil Salinity Research Institute, Karnal, India.
- Saqib M, AKhtar J, Qureshi RH (2008) Sodicity intensifies the effect of salinity on grain yieldand yield components of wheat. J Plant Nutr 31: 689-701.
- Zeng L, Shannon MC (2000) Salinity effects on seedling growth and yield components of rice. Crop Sci 40: 996-1003.
- Abdullah Z, Khan MA, Flowers TJ (2001) Causes of sterility in seed set of rice under salinity stress. J Agron Crop Sci 187: 25-32.
- Kato Y, Kamoshita A, Yamagishi J (2008) Prefloweringabortionreduces spikelet number in upland rice (Oryza sativa L.) under water stress. Crop Sci 48: 2389-2395.
- Rao PS, Mishra B, Gupta SR, Rathore A (2008) Reproductive stage tolerance to salinity and alkalinity stresses in rice genotypes. Plant Breed 127:256-261.
- Netondo GW, Onyango JC, Beck E (2004) Sorghum and salinity: II. Gas exchange and chlorophyll fluorescence of sorghum under salt stress. Crop Sci 44: 806-811.
- Cha-um S, Supaibulwatana K, Kirdmanee C (2006) Water relation, photosynthetic ability and growth of thai jasmine rice (Oryzasativa L. ssp. indica cv. KDML 105) to salt stress by application of exogenous glycine betaine and choline. J Agron Crop Sci 192: 25-36.
- Chaves MM, Flexas J, Pinheiro C (2009) Photosynthesis under drought and salt stress: regulation mechanisms from whole plant to cell. Ann Bot 103: 551-560.
- Moradi F, Ismail AM (2007) Responses of photosynthesis, chlorophyll fluorescence and ROS-scavenging systems to salt stress during seedling and reproductive stages in rice. Ann Bot 99: 1161-1173.
- Munns R (2005) Genes and salt tolerance: bringing them together. New Phytol 167: 645-653.
- Garthwaite AJ, Bothmer RV, Colmer TD (2005) Salt tolerance in wild Hordeum species is associated with restricted entry of Na+ and Cl- into the shoots. J Expt Bot 56: 2365-2378.
- Mozafar A and Goodin JR (1970) Vesiculated Hairs: A Mechanism for Salt Tolerance in Atriplexhelimus L. Plant Physiol 45: 62-65.
- Sickler CM, Edwards GE, Kiirats O, Gao Z, Loescher W (2007) Funct Plant Biol 34: 382-391.
- Kant S, Pragya S, Raveh E, Barak S (2006) Evidence that differential gene expression between the halophyte,Thellungiellahalophila, and Arabidopsis thaliana is responsible for higher levels of the compatible osmolyteproline and tight control of Na+ uptake in T halophila. Plant cell Env 29: 1220-1234.
- Bajaj S, Mohanty A (2005) Recent advances in rice biotechnology towards genetically superior transgenic rice. Plant Biotechnol J 3: 275-307.
- Harkamal W, Clyde W, Linghe Z, Abdelbagi MI, Pascal C, et al. (2007) Genome wide transcriptional analysis of salinity stressed japonica and indica rice genotypes during panicle initiation stage. Plant MolBiol 63: 609-623.
- FAO (2004) FAO Land and Plant Nutrition Management Service.
- Kumar V, Shriram V, Jawali N, Shitole MG (2007) Differential response of indica rice genotypes to NaCl stress in relation to physiological and biochemical parameters. Arch Agron Soil Sci 53:581-592.
- Mass EV, Hoffman GJ (1977) Crop salt tolerance-current assessment. J Irrigation Drainage Div 103: 115-134.
- Sahi C, Singh A, Kumar K, Blumwald E, Grover A (2006) Salt stress response in rice: Genetics, molecular biology and comparative genomics. FuncIntegrative Genomics 6: 263-284.
- Heenan DP, Levin LG, Mcaffery (1988) Salinity tolerance in rice varieties at different growth stages. Aus J ExperAgric 28: 343-349.
- Djanaguiraman M, SheebaJA,Shanker AK, Durga Devi D, Bangarusamy U (2006) Rice can acclimate to lethal level of salinity by pretreatment with subletal level of salinity through osmotic adjustment. Plant Soil 284: 363-373.
- Gregorio GB (1997) Tagging salinity tolerance genes in rice using amplified fragment length polymorphism AFLP. Univ. Philippines, Los Baños, Philippines.
- Akbar M, Gunawardena IE, Ponnamperuma FN (1986) Breeding for soil stresses In: Progress in rain fed lowland rice. IRRI, Manilla, Philippines pp: 263-272.
- Jones RGW (1981) Salt tolerance. In: CB Johnson (Ed), Physiological Processes Limiting Plant Productivity. Butterworths, London pp: 271-292.
- Khatun S, Flowers TJ (1995) Effects of salinity on seed set in rice Plant Cell Environ 18: 61-67.
- Zeng L, Shannon MC, Lesch, SM (2001) Timimg of salinity stress affects rice growth and yield components. Agri Water Manage 48: 191-206.
- Colom MR, Vazzana C (2003) Photosynthesis and PS II functionally of draught resistant and drought sensitive creeping love grass plants. Environ Exp Bot 49: 135-144.
- Chang CS, Sung JM (2004) Nutrient uptake and yield responses of peanuts and rice to lime and fused magnesium phosphate in an acid soil. Field Crops Res 89: 319-325.
- Fukuda T, Saltu A, Wasaki J, Shinana T, Osaki M (2007) Metabolic alterations proposed by proteome in rice roots grown under a low P and high Al concentrations under low pH. Plant Sci 172: 1157-1165.
- Kang DJ, Futakuchi K, Dumnoenng S, Ishii R (2007) High yielding performance of a new rice, IR 53650 in mildly improved acid sulfate soil conditions. Plant Prod Sci 10: 64-67
- Yang F, Liang ZW, Wang Ze, Chen Y (2008) Relationship between diurnal changes of net photosynthetic rate and influencing factors in rice under saline sodic stress. Rice Sc15: 119-124.
- Bhowmik SK, Titov S, Islam MM, Siddika A, Sultana S, et al. (2009) Phenotypic and genotypic screening of rice genotypes at seedling stage for salt tolerance. Afr JBiotechnol 8: 6490-6494.
- Haq TU, Akhtar J, Nawaz S, Ahmad R (2009) Morpho-physiological response of rice (Oryza sativa L) varieties to salinity stress. Pak J Bot 41: 2943-2956.
- Ali MN, Ghosh B, Gantait S, Chakraborty S (2014a) Selection of rice genotypes for salinity tolerance through morpho-biochemical assessment. Rice Sci 21: 288-298.
- Asch F, Wopereis MCS (2001) Responses of field-grown irrigated rice cultivars to varying levels of floodwater salinity in a semi-arid environment. Field Crop Res70: 127-137.
- Khatun S, Rizzo CA, Flowers TJ (1995) Genotypic variation in the effect of salinity on fertility in rice. Plant Soil 173: 239-250.
- Lutts S, Kinet JM, Bouharmont J (1995) Changes in plant response to NaCl during development of rice (Oryza sativa L) varieties differing in salinity resistance. J Exp Bot 46: 1843-1852.
- Shannon MC, Rhoades JD, Draper JH, Scardaci SC, Spyres MD (1998) Assessment of salt tolerance in rice cultivars in response to salinity problems in California. Crop Sci 38: 394-398.
- Zeng L, Lesch SM, Grieve, CM (2003) Rice growth and yield respond to changes in water depth and salinity stress. Agri Water Manage 59: 67-75.
- Yeo AR, Yeo ME, Flowers SA, Flowers TJ (1990) Screening of rice (Oryzasatia L) genotypes for physiological characters contributing to salinity resistance, and their relationship to overall performance. TheorAppl Genet 79: 377-384.
- Rahman MS, Matsumuro T, Miyake H, Takeoka Y (2000) Salinity induced ultrastructural alterations in leaf cells of rice (Oryza sativa L). Plant Prod Sci 3: 422-429.
- Baker NR (2008) Chlorophyll fluorescence: a probe of photosynthesis in vivo. Annu Rev Plant Biol 59: 89-113.
- Bahaji A, Mateu I, Sanz A, Cornejo MJ (2002) Common and distinctive responses of rice seedlings to saline and osmotically generated stress. Plant Growth Regul 38: 83-94.
- Wankhade SD, Bahaji A, MateuAndrésI, Cornejo MJ (2010) phenotypic indicators of NaCl tolerance levels in rice seedlings: variations in development and leaf anatomy. ActaPhysiol Plant 32: 1161-1169.
- Wankhade SD, Cornejo MJ, Mateu-Andrés I, Sanz A (2013) Morpho-physiological variations in response to NaCl stress during vegetative and reproductive development of rice. ActaPhysiol Plant 35: 323-333.
- Garcia A, Rizzo CA, Ud-Din J, Bartos SL, Senadhira D, et al. (1997) Sodium and potassium transport to the xylem are inherited independently in ice, and the mechanism of sodium: potassium selectivity differs between rice and wheat. Plant Cell Env 20: 1167-1174.
- Yeo AR, Flowers SA, Rao G, Welfare K, Senanayake N, et al. (1999) Silicon reduces sodium uptake in rice (Oryza sativa L) in saline conditions and this is accounted for a reduction in the transpirational bypass flow. Plant Cell Environ 22: 559-565.
- Flowers TJ, Yeo AR (1986) Ion relation of plants under drought and salinity. Aust J Plant Physiol 13: 75-91.
- Asch F, Dingkuhn M, Dörffling K (1997) Physiological stresses of irrigated rice caused by soil salinity in the Sahel. Irrigated Rice in the Sahel: Prospects for Sustainable Development, West Africa Rice Development Association, Bouake, Cote d’Ivoire pp: 247-273.
- Qadar A (1988) Potassium status of the rice shoot as Index for salt tolerance. Ind J Plant Physiol 31: 388-393.
- Rajarathinam S, Koodalingam K, Raja VDG (1988) Effect of potassium and sodium in rice for tolerance of soil salinity. J Potassium Res 4: 174-178
- Pandey UK, Srivastava RDL (1991) Leaf potassium as an index of salt tolerance in paddy. Nat AcadSci Let 14: 161-164.
- Rahman MS, Matsumuro T, Miyake H, Takeoka T (2001) Effects of salinity stress on the seminal root tip ultrastructures of rice seedlings (Oryza sativa L). Plant Prod Sci 4: 103-111.
- Flowers TJ, Yeo AR (1981) Variability in the resistance of sodium chloride salinity within rice (Oryza sativa L) varieties. New Phytol 88: 363-373.
- Akita S, Cabuslay G (1990) Physiological basis of differential response to salinity in rice cultivars. Plant and Soil 123: 277-294.
- Yaghoubian Y, Pirdashti H,Mottaghian A, Hosseini S (2012) Effect of fluctuating salinity at different growth stages on physiological and yield related parameters of rice (OryzasativaL). Int J Agricult: Res Rev 2: 266-276.
- Yancey PH, Clark ME, Hand SC, Bowlus RD, Somero GN (1982) Living with water stress, Evolution of osmolyte systems. Science 217: 1212-1222.
- Ashraf M (1994) Breeding for salinity tolerance in plants. Crit Rev Plant Sci 13: 17-42.
- Ali G, Srivastava PS, Iqbal M (1999) Proline accumulation, protein pattern and photosynthesis in regenerants grown under NaCl stress. Biol Plant 42: 89-95.
- Lutts S, Kinet JM, Bouharmont J (1996) NaCl-induced senescence in leaves of rice (Oryza sativa L.) cultivars differing in salinity resistance. Ann Bot 78: 389-398.
- Bandurska H (1993) In vivo and in vitro effect of proline on nitrate reductase activity under osmotic stress in barley. ActaPhysiol Plant 15: 83-88.
- Perez AF, Estan MT, Santa Cruz A, Bolarin MC (1993) Effects of salinity on nitrate, total nitrogen, soluble protein and free amino acid levels in tomato plants. J HortSci 68: 1021-1027.
- Cram WJ (1976) Negative feedback regulation of transport in cells. The maintenance of turgor, volume and nutrient supply. In: U Luttge, MG Pitman (Eds.) Encyclopedia of Plant Physiology New Series Vol. 2, Springer-Verlag, Berlin, Germanypp: 284-316.
- Popp M, Smirnoff N (1995) Polyol accumulation and metabolism during water deficit. In: N Smirnoff (Ed.) Environment and Plant Metabolism: Flexibility and Acclimation. Bios Scientific, Oxford, UK pp: 199-215.
- Amirjani MR (2011) Effect of Salinity stress on Growth, Sugar content, Pigments and Enzyme activity of rice. Int J Bot 7: 73-81.
- Hurry VM, Strand A, Tobiaeson M, Gardarstorm P, Oquist G (1995) Cold hardening of spring and winter wheat and rape results in differential effect of growth carbon metabolism and carbohydrate content. Plant Physiol 109:697-706.
- Dubey RS, Singh AK (1999) Salinity induces accumulation of soluble sugars andalters the activity of sugar metabolizing enzymes in rice plants. Biol Plant 42: 233-239.
- Sakamoto A,Murata A, Murata N (1998) Metabolic engineering of rice leading to biosynthesis of glycine betaine and tolerance to salt and cold. Plant MolBiol 38: 1011-1019.
- Pareek A, Singla SL, Grover A (1997) Salt responsive proteins/genes in crop plants. In: PK Jaiwal,RP Singh, A Gulati (Eds) Strategies for Improving Salt Tolerance in Higher Plants. Oxford and IBH Publication Co., New Delhi, India pp: 365-391.
- Singh NK, Bracken CA, Hasegawa PM, Handa AK, Buckel S, et al. (1987) Characterization of osmotin. A thaumatin-like protein associated with osmotic adjustment in plant cells. Plant Physiol 85: 529-536.
- Jha BN, Singh R (1997) Physiological response of rice varieties to different levels of moisture stress. Ind J Plant Physiol 2: 81-84.
- Akbar M, Yabuno T, Nakao S (1972) Breeding for Saline-resistant Varieties of Rice: I. Variability for Salt Tolerance among Some Rice Varieties. Jpn J Breed 22: 277-284.
- Akbar M, Yabuno T (1975) Breeding for saline-resistant varieties of rice. III. Response of F1 hybrids to salinity in reciprocal crosses between "Jhona 349" and "Magnolia". Jpn J Breed 25: 215-220.
- Ikehashi H, Ponnamperuma FN (1978) Varietal tolerance to rice in adverse soils. In: Soils and rice. IRRI, Los Bafios, Laguna, Philippines pp: 801-823.
- Bonilla PS, Dvorak J, Mackell D, Deal K, Gregorio G (2002) RFLP and SSLP mapping of salinity tolerance genes in chromosome 1 of rice (Oryza sativa L) using recombinant inbred lines. The Phil AgriSci 85: 68-76.
- Kanawapee N, Sanitchon J, Srihaban P, Theerakulpisut P (2011) Genetic diversity analysis of rice cultivars (Oryza sativa L.) differing in salinity tolerance based on RAPD and SSR markers. Electronic J Biotechnol 14: 1-17.
- Ali MN, Yeasmin L, Gantait S, Goswami R, Chakraborty S (2014b) Screening of rice landraces for salinity tolerance at seedling stage through morphological and molecular markers. PhysiolMolBiol Plants.
- Rikke BA, Johnson TE (1998) Towards the cloning of genes underlying murine QTLs. Mamm Genome 9: 963-968.
- Ron M, Weller JI (2007) From QTL to QTN identification in livestock “Winning by points rather than knock-out”. Animal Genetics: a review 38: 429-439.
- Bechtold N, Ellis J, Pelletier G (1993) In planta Agrobacterium mediated gene transfer by infiltration of adult Arabidopsis thaliana plants. C R AcadSci Paris, Life Sciences 316: 1194-1199.
- Salvi S, Tuberosa R (2005) To clone or not to clone plant QTLs: present and future challenges. Trends Plant Sci 10: 297-304.
- Rabbani MA, Maruyama K, Abe H, Khan MA, Katsura K, et al. (2003) Monitoring Expression Profiles of Rice Genes under Cold, Drought, and High Salinity Stresses and Abscisic Acid Application Using cDNA Microarray and RNA Gel-Blot Analyses. Plant Physiology 133: 1755-1767.
- Urao T, Yakubov B, Satoh R, Yamaguchi-Shinozaki K, Seki M, et al. (1999) A transmembrane hybrid-type histidine kinase in Arabidopsis functions as an osmo sensor. The Plant Cell 11: 1743-1754.
- Xiong L, Yang Y (2003) Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid-inducible mitogen-activated protein kinase. Plant Cell 15: 745-759
- Zhu JK (2002) Salt and drought stress signal transduction in plants. Ann Rev Plant Biol 53: 247-273.
- Haq TU, Akhtar J, Gorham J, Steele KA, Khalid M (2008) Genetic mapping of QTLs, controlling shoot fresh and dry weight under salt stress in rice (Oryza sativa L) cross between Co39 × MOROBEREKAN. Pak J Bot 40: 2369-2381.
- Sabouri H,Sabouri A (2008) New evidence of QTLs attributed to salinity tolerance in rice. Afr J Biotechnol 7: 4376-4383.
- Koyama ML, Levesley A, Koebner RMD, Flowers TJ, Yeo AR (2014) Quantitative Trait Loci for Component Physiological Traits Determining Salt Tolerance in Rice. Plant Physiol 125: 406-422
- Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, et al. (2004) QTLs for Na+ and K+ uptake of the shootsand roots controlling rice salt tolerance. TheorAppl Genet 108: 253-260.
- Mohammadi R, Mendioro MS, Diaz GQ, Gregorio GB, Singh RK (2013) Mapping quantitative trait loci associated with yield and yield components under reproductive stage salinity stress in rice (Oryza sativa L). J Genet 92: 433-443.
- Mohammadi NG, Arzani A, Rezail AM, Singh RK, Gregorio GB (2008) Assessment of rice genotypes for salt tolerance using microsatellite markers associated with the saltol QTL. Afr J Biotechnol 7: 730-736.
- Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress and signal transduction. Ann Rev Plant Biol 55: 373-99.
- Logan BA (2005) Reactive oxygen species and photosynthesis. In: N Smirnoff (Eds.) Antioxidants and Reactive Oxygen Species in Plants, Oxford: Blackwell, pp: 250-267.
- Asada K (1999) The water cycle in chloroplast: scavenging of active oxygens and dissipation of excess photons. Annu Rev Plant Biol 50: 601-639.
- Sharma P, Dubey RS (2007) Involvement of oxidative stress and role of antioxidative defense system in growing rice seedlings exposed to toxic concentrations of aluminum. Plant Cell Rep 26: 2027-2038.
- Dionisiosese ML, Tobita J (1998) Antioxidant response of seedlings to salinity stress. Plant Sci 135: 1-9.
- Demiral T, Turkan I (2005) Comparative lipid peroxidation, antioxidant defense systems and proline content in roots of two rice cultivars differing in salt tolerance. EnvExp Bot 53: 247-257.
- Mandhania S, Madan S, Sawhney V (2006) Antioxidant defense mechanism under salt stress in wheat seedlings. Biol Plant 227: 227-231.
- Mittal R, Dubey RS (1991) Behaviour of peroxidases in rice: changes in enzyme activity and isoforms in relation to salt tolerance. Plant PhysiolBiochem 29: 31-40.
- Kalir A, Omri G, Poljakoff-Mayber A (1984) Peroxidase and catalase activity in leaves of Halimioneportulacoides exposed to salinity. Physiol Plant 62: 238-244.
- Sheoran IS, Garg OP (1979) Quantitative and qualitative changes in peroxidase during germination of mung bean under salt stress. Plant Physiol 46: 147-150.
- Garg AK, Kim JK, Owens TG, Ranwala AP, Choi YDO, et al. (2002) Trehalose accumulation in rice plants confershigh tolerance levels to different abiotic stresses. ProcNatlAcadSci USA 99: 15898-15903.
- Mishra P, Bhoomika K, Dubey RS (2013) Differential responses of antioxidative defense system to prolonged salinity stress in salt-tolerant and salt-sensitive Indica rice (Oryza sativa L) seedlings. Protoplasma 250: 3-19.
- Sarkar RK, Mahata KR, Singh DP (2013) Differential responses of antioxidant system and photosynthetic characteristics in four rice cultivars differing in sensitivity to sodium chloride stress. ActaPhysiol Plant 35: 2915-2926.
- Noble CL, Rogers ME (1992) Arguments for the use of physiological criteria for improving the salt tolerance in crops. Plant Soil 146: 99-107.
- Al-Khatib M, McNeilly T, Collins JC (1993) The potential of selection and breeding for improved salt tolerance in lucerne (Medicago sativa L). Euphytica 65:43-51.
- Ashraf M (2002) Salt tolerance of cotton: some new advances. Crit Rev Plant Sci 21: 1-30.
- Mansour MMF (2000) Nitrogen containing compounds and adaptation of plants to salinity stress. Biol Plant 43: 491-500.
- Mittova V, Guy M, Tal M, Volokita M (2002) Response of the cultivated tomato and its wild salt-tolerant relative Lycopersiconpennellii to salt-dependent oxidative stress: increased activities of antioxidant enzymes in root plastids. Free Radic Res 36: 195-202.
- Gregorio GB and Senadhira D (1993) Genetic analysis of salinity tolerance in rice (Oryza sativa L.). TheorAppl Genet 86: 333-338.
- Zeng Y, Zhang H, Li Z, Shen S, Sun J, et al. (2007) Evaluation of Genetic diversity of rice landraces (Oryza sativa L.) in Yunnan, China. Breeding Sci 57: 91-99.
- Roychoudhury A, Basu S, Sarkar SN, Sengupta DN (2008) Comparative physiological and molecular responses of a common aromatic indica rice cultivar to high salinity with non-aromatic indica rice cultivars. Plant Cell Rep 27: 1395-1410.
- Yeo AR, Flowers TJ (1986) Salinity resistance in rice (Oryza sativa L) and a pyramiding approach to breeding varieties for saline soils In NC Turner, JB Passioura (Eds) Effect of drought on plant growth: Salts in soils. CSIRO, Melbourne, Australia pp: 161-173.
- Babu CR, Vijayalakshmi C, Mohandass, S (2005) Evaluation of rice (Oryza sativa L) genotypes for salt tolerance. J Food AgriEnv 3: 190-194.
- Flowers TJ, Yeo AR (1995) Breeding for salinity resistance in crop plants: Where next? Aust J Plant Physiol 22: 875-884.
- Gregorio GB, Dharmawansa S, Mendoza RD (1997) Screening rice for salinity tolerance. IRRl Discussion Paper Series No. 22. IRRI, Manila, Philippines pp: 1-30.
- Maggio A, Raimondi G, Martino A, De Pascale S (2007) Salt stress response in tomato beyond the salinity tolerance threshold. EnvExp Bot 59: 276-282.
- Noble CL, Halloran GM, West DW (1984) Identification and selection for salt tolerance in lucerne (Medicago sativa L). Aust J Agric Res 35: 239-252.
- Soda N, Kushwaha HR, Soni P, Singla-Pareek SL, Pareek A (2013) A suite of new genes defining salinity stress tolerancein seedlings of contrasting rice genotypes. FunctIntegr Genomics 13: 351-365.
- Thompson MJ, Ocampo MD, Egdane J, Rahman MA, Sajise AG, et al. (2010) Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 3: 148-160.
Relevant Topics
- Basmati Rice
- Drought Tolerence
- Golden Rice
- Leaf Diseases
- Long Grain Rice
- Par Boiled Rice
- Raw Rice
- Rice
- Rice and Aquaculture
- Rice and Nutrition
- Rice Blast
- Rice Bran
- Rice Diseases
- Rice Economics
- Rice Genome
- Rice husk
- Rice production
- Rice research
- Rice Yield
- Sticky Rice
- Stress Resistant Rice
- Unpolished Rice
- White Rice
Recommended Journals
Article Tools
Article Usage
- Total views: 24513
- [From(publication date):
May-2016 - Aug 30, 2025] - Breakdown by view type
- HTML page views : 21634
- PDF downloads : 2879