Research Article Open Access
Production and Partial Purification of Beta-Mannanase from Aspergillus niger Associated with Ilaje Lake, Ondo State, Nigeria
Oladipo OO* and Olajide AdebowaleDepartment of Microbiology, Federal University of Technology, PMB 704, Akure, Nigeria
- Corresponding Author:
- Oladipo OO
Department of Microbiology
Federal University of Technology
P.M.B 704, Akure, Nigeria
Tel: +2348068054636
E-mail: microladit@gmail.com
Received date: May 17, 2017; Accepted date: June 12, 2017; Published date: June 19, 2017
Citation: Oladipo OO, Adebowale O (2017) Production and Partial Purification of Beta-Mannanase from Aspergillus niger Associated with Ilaje Lake, Ondo State, Nigeria. Biochem Physiol 6:218. doi:10.4172/2168-9652.1000218
Copyright: © 2017 Oladipo OO, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Biochemistry & Physiology: Open Access
Abstract
The present study aimed at the isolation, screening and partial purification of beta-mannanase from fungi isolated from soil and water samples collected from Ilaje Lake, Ondo state, Nigeria. The associated fungi were isolated and counted by standard microbiological methods. Partial purification of crude mannanase was done by standard biochemical methods. Quantitatively, mannanase production was performed in mineral salt medium into which Locust Bean Gum (LBG) had been incorporated as the sole carbon source. Enzyme activity was determined by dinitrosalicylic acid (DNSA) method, while protein content was evaluated by Lowry’s method. The highest fungal counts were recorded for water sample collected from Ilaje Lake with 3.4 × 108 SFU/mL. The organisms encountered include Aspergillus flavus, Rhizopus stolonifer, A. fumigatus, A. niger, R. japonicus, Penicillum italicum, Fusarium solani and Candida albicans. All the fungal isolates encountered from these sources showed varied degrees of mannanase activities. The highest specific mannanase activity was recorded for isolate 4B1, while the lowest value was obtained with isolate 1B2. Purification of crude mannanase from A. niger was carried out by ammonium sulphate precipitation and gel filtration (Sephadex G-200). Fractionation of ammonium sulphate precipitated mannanase from A. niger on Sephadex G-200 produced two activity peaks. In this investigation, fungal isolates evaluated for mannanase production from this source gave appreciable mannanase activity and this could be applied in many industrial processes.
Keywords
Beta-mannanase; Ilaje Lake; Locust bean gum; Fungal counts; Partial purification
Introduction
Beta-mannanase otherwise known as mannan endo-1,4-β- mannosidase or 1,4-β-D-mannanase with an enzyme commission (EC) 3.2.1.78 breakdowns β-1,4-mannosidic bonds in the main chain of glucomannans, galactomannans and β-1,4-mannans [1]. It transforms the abundant mannan-rich heteromannan, glucomannan and galactomannan substrates into manno-oligosaccharides [2] and a small amount of mannose, glucose and galactose [1,3]. A number of plants, bacteria, fungi and various invertebrates had been reported to produce β-mannanase [1,3]. The mannanase of biological origins have found a variety of applications in different industrial sectors [3] including animal feed formulation, pharmaceutical products preparation, pulp biobleaching, [4] as well as pre-treatment of plant biomass for second generation biofuel production [5].
There is a greater demand for biotechnological and industrial stable enzymes with higher catalytic efficiency that can function under harsh environmental conditions. In the recent years, screening and isolation of microorganisms for the production of industrial enzymes of microbial origin with unique and excellent characteristics has become one of the focuses of research in biotechnology. Enzymes required for industrial applications should be stable at high temperature, pH and presence of salts, solvents, etc. Extracellular enzymes from microbes associated with extreme environments such as halophiles are blessed with higher catalytic power to initiate and sustain metabolic activities under high salt concentrations. Gomes and Steiner [6] and Karan and Khare [7] have been reported that enzymes from extreme environments are expected to be active and stable under more than one extreme condition.
The physicochemical analysis carried out on four different coastal waters from Ilaje, Ondo state, Nigeria revealed that they were slightly acidic with a mean pH of 6.69 and also moderately saline with a mean salinity of 16.68% [8]. Hence, mannanase obtained from bacteria isolated from this source are expected to be halo tolerant. Mannanaseproducing microorganisms have been reported from different sources but there has been scanty or no information on mannanase production from bacteria inhabits Ilaje Lake. Hence, enzymes from halo tolerant bacteria from this location should have an edge in terms of stability. In search of high performance mannolytic organisms; we isolated and screened different fungi from soil and water samples in Ilaje, Ondo State, Nigeria. Therefore, the present study dealt with the isolation and screening of fungal isolates for beta-mannanase production and its partial purification from Aspergillus niger.
Materials and Methods
Sources of sample
Water and soil samples were collected from Ilaje Lake, Ondo State, Nigeria in sterile bottles. The samples were transferred to the laboratory and were used as sources for the isolation of mannanase-producing fungi [9,10].
Isolation and identification of mannanase-producing fungi from soil and water samples
The total fungal counts from the samples were determined using the pour plate method on potato dextrose agar. The samples were serially diluted and 1ml of an appropriate dilution was used to inoculate the plate in duplicate. The plates were incubated at 28 ± 2°C for 72 h, after which the total colony count was determined as previously described by Lateef et al. [11]. At the end of incubation, the colonies were subcultured from the mixed cultures and identified on the bases of cultural characters (colour, shape of colony, surface and reverse pigmentation and texture of the colony) as well as microscopic structure (septate or nonseptate hyphae, structure of hyphae and conidia) [12].
Screening of mannanase-producing fungi in submerged state fermentation
Medium composition described by Mandles and Weber [12] modified by Arotupin and Olaniyi [10] was used for submerged fermentation (static condition). The basal medium contained (g/L): LBG 10 g, peptone 2 g, yeast extract 2 g, NaNO3 2 g, KH2PO4 1 g, MgSO4.7H2O 0.5 g, KCl 0.5 g and FeSO4.7H2O traces. The pH of the media was adjusted to 6.8 with pH meter (Denver Instrument, Model 20 pH/Conductivity meter) prior sterilization. Then, 100 mL of the liquid medium was dispensed in 250 mL Erlenmeyer flask and sterilized at 121°C for 15 min. The sterile basal medium was inoculated with 2 discs of 8 mm diameter of the fungal strains from PDA using cork borer. The inoculated media were incubated at 30°C for 5 days at static condition. Crude enzyme preparation was obtained by centrifugation at 6000 rpm for 10 min at 4°C using refrigerated ultracentrifuge (Centurion Scientific Limited). The supernatant was used as the crude extracellular enzyme source. Each treatment was carried out in triplicates and the results obtained throughout the work were the mean of at least 3 experiments.
Enzyme assay and protein determination
Mannanase activity was assayed in the reaction mixture comprising of 0.5 mL of 1% LBG prepared in 50 mM potassium phosphate buffer pH 6.8 and 0.5 mL of supernatant at 45°C for 60 min (modified method of El-Naggar et al. [13]). At the end of the incubation period, tubes were removed from the water bath (Lamfield Medical England Model DK-600) and the reaction was terminated by the addition of 2 ml of 3,5-dinitrosalicylic acid (DNSA) reagent per tube. The tubes were incubated for 5 min in a boiling water bath for colour development and were cooled rapidly. The activity of reaction mixture was measured against a reagent blank at 540 nm. Amount of reducing sugar released was determined by the dinitrosalicylic acid reagent (DNS) [14]. One unit of mannanase activity was defined as amount of enzyme producing 1 μmol of mannose/min under the experimental conditions. The amount of protein produced in the basal medium was evaluated according to the method of Lowry et al. [15,16] using Bovine Serum Albumin (BSA) as the standard.
Mannanase purification
Mannanase purification was performed in three stages. In the first stage, the supernatant obtained after centrifugation was precipitated by the addition of ammonium sulphate to achieve 70% ammonium sulphate concentration [1]. The precipitated enzyme was then diluted in phosphate buffer 50 mM pH 7.0, and dialyzed in the same buffer. Secondly, the enzyme obtained after dialysis was loaded into ion exchange column chromatography with diethylaminoethyl (DEAE) Sephadex G-200 matrix (20 × 2.5 cm, Pharmacia). The fractions obtained were sequentially washed with ion free water and 0.01 M Tris-HCl buffer pH 8.0. The bound protein molecules were eluted with concentration gradient of NaCl. The absorbance of each fraction collected was measured at 280 nm with UV spectrophotometer (Lab- Tech Digital) and the activity of mannanase of each fraction was determined. In the third stage, concentrated enzyme from second stage purification step was loaded onto the column chromatography (2.5 cm in diameter and 30 cm high) which contained Sephadex G-200 (Pharmacia). The concentrated enzyme was eluted with phosphate buffer pH 7.0 at the flow rate of 20 ml/h.A fraction of 10 mL was collected at an interval of 30 minutes and the absorbance was taken at 280 nm. Fractions with mannanase activity were pooled and concentrated in glycerol solution at 30°C.
Results
Populations of mannanase-producing fungi from soil and water samples
Table 1 shows the total fungal count from each of the sources. Water sample had the highest number of fungal population of 3.4 × 108 SFU/mL, while soil sample from Ilaje Lake (2.2 × 108 SFU/g) recorded the least fungal counts.
| Sources | Fungal counts |
|---|---|
| ILS | 2.2 × 108 SFU/g |
| ILW | 3.4 × 108 SFU/ml |
ILS=Soil sample from Ilaje Lake
ILW=Water sample from Ilaje Lake
Table 1: Populations of mannanase-producing fungi from soil and water samples.
Cultural characterization and microscopic observation of fungal isolates
Table 2 shows the cultural characterization and microscopic observation of fungal isolates associated with soil and water samples from Ilaje Lake. Eight fungal isolates were isolated from the samples. The organisms encountered include Aspergillus flavus, A. fumigatus, A. niger, Rhizopus japonicus, R. stolonifer, Penicillum italicum, Fusarium solani and Candida albicans.
| Isolate code | Cultural Characteristics |
Microscopic observation | Suspected organisms |
|---|---|---|---|
| 1B2 and Z3E2 | Greenish yellow mycelial growth | An upright conidiophores that terminates in a davate swelling bearing phialides at the apex or radiating from the entire surface; conidia are 1-celled and globose. | Aspergillus flavus |
| 1C2 | Brown mycelia | Conidiospores upright, simple, terminating in elevates, swelling, bearing phalides at the apex or radiating from the entire surface, L celled globe often variously coloured in mathes in dry basipetal chains. | Aspergillus fumigatus |
| 2E3 and 4E3 | Rapidly growing white colored fungus, swarms over entire plate | Non-septate mycelium gives rise to straight sporangiophores that with sporangium containing a columella; root like hyphae (rhioziod) penetrating the medium. | Rhizopus stolonifer |
| 2K2 | White mycelia with areas of whitish yellow | Fast growing rate with aerial mycelium. They appeared as sickle-shaped. | Fusarium solani |
| 3E2 | Blue mould growth | Septate mycelium bearing Single conidiophores which are branched near the apex ending in phialides that carries the conidia. | Penicillium italicum |
| 4B1 | Black mycelia | Simple, upright conidiophores terminating in ovoid swelling. | A. niger |
| 4B2 | White to cream colour, smooth mycelial growth | Spherical budding cells or Blastoconidia. | Candida albicans |
| 4E2 | Rapidly growing white colored fungus, swarm over entire plate; then showed black spores varying in sizes | Non-septate mycelium gives rise to straight sporangiophores that sporangium; root like hyphae (rhioziod) penetrating the medium. | R. japonicas |
Table 2: Cultural characterization and microscopic observation of fungal isolates.
Percentage occurrence of fungal isolates
Table 3 shows the percentage occurrence of fungal isolates associated with the samples. The highest percentage occurrence of 20% was obtained for Aspergillus flavus and Rhizopus stolonifer, while Aspergillus fumigatus, A. niger, Rhizopus japonicus, Penicillum italicum, Fusarium solani and Candida albicans had the same value of 10%.
| Fungal isolates | % Occurrence of isolates |
|---|---|
| Aspergillus flavus | 20 |
| A. fumigatus | 10 |
| Rhizopus stolonifer | 20 |
| Fusarium solani | 10 |
| Penicillium italicum | 10 |
| A. niger | 10 |
| Candida albicans | 10 |
| R. japonicas | 10 |
Table 3: Percentage occurrence of fungal isolates.
Mannanase production by fungal isolates from different samples
The quantitative determination of mannan-degrading enzyme from different fungal isolates is shown in Table 4. All the fungal isolates encountered from different sources displayed varied degrees of mannanase activities. The highest mannanase activity and specific activity of 9.37 U/mL and 2.59 U/mg respectively was produced by fungal isolate 4B1 sourced from water sample, while the lowest values for mannanase activity and specific activity were obtained from 2E3 and 1B2 respectively. Protein content ranged from 3.06 mg/mL to 5.39 mg/mL was produced by the fungal isolates, with the highest protein content displayed by isolate IB2 obtained from soil sample. Therefore, fungal isolate 4B1 was selected for purification studies because of its highest mannanase-producing potential.
| Source | Isolate code | Mannanase activity (U/mL) | Protein Content (mg/mL) |
Specific mannanase activity (U/mg) |
|---|---|---|---|---|
| ILS | 1B2 | 4.17 | 5.29 | 0.79 |
| 1C2 | 4.47 | 3.27 | 1.37 | |
| 2E3 | 3.58 | 3.28 | 1.09 | |
| 2K2 | 7.13 | 3.30 | 2.16 | |
| ILW | 3E2 | 4.20 | 3.38 | 1.24 |
| Z3E2 | 6.71 | 3.49 | 1.92 | |
| 4B1 | 9.37 | 3.62 | 2.59 | |
| 4B2 | 5.18 | 3.06 | 1.70 | |
| 4C3 | 3.65 | 3.28 | 1.11 | |
| 4E2 | 5.28 | 3.24 | 1.63 |
ILS=Soil sample from Ilaje Lake
ILW=Water sample from Ilaje Lake
Table 4: Mannanase production by fungal isolates from different samples.
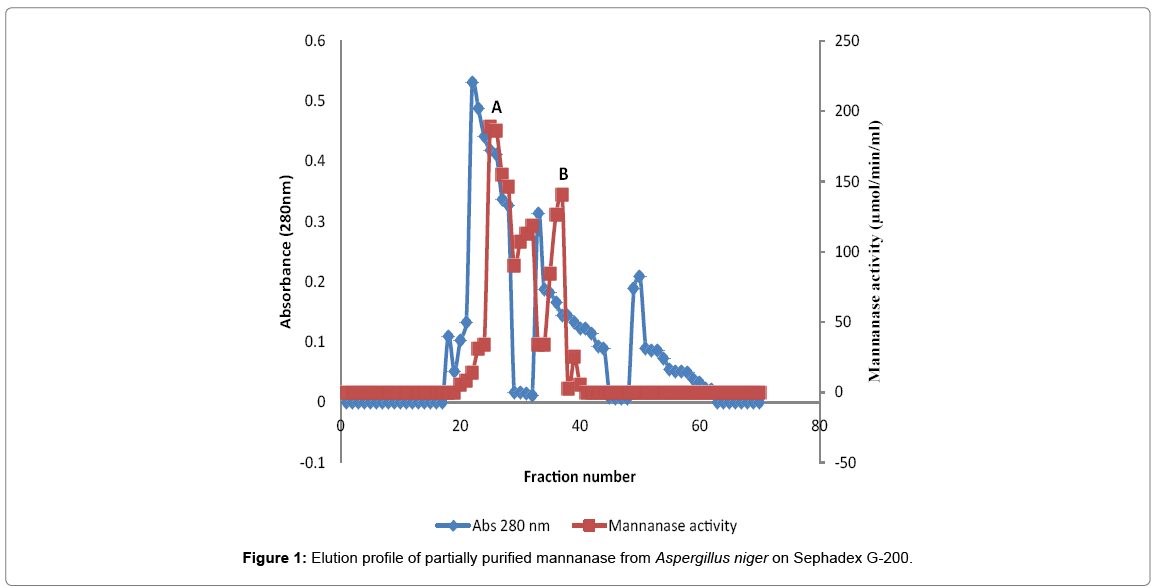
Elution profile of partially purified mannanase on sephadex G-200
After 25% ammonium sulphate saturation of crude enzyme, the precipitates were dialyzed against 50 mM phosphate buffer pH 6.8 for 24 h at 4°C. Dialyzed protein was loaded onto a Sephadex G-200 column which was eluted with the same buffer. The ammonium sulphate-dialysate fraction on Sephadex G-200 tends to produce two activity peaks represented as A and B (Figure 1). Mannanase activity increased appreciably and reached A with an enzyme activity of 98.09 μmol/min/mL, while peak B had enzyme activity of 148.12 μmol/min/ mL.
Summary of purification procedures
Three step purification procedures were adopted and a 7.43-fold purification was achieved with specific activity of 48.06 μmol/min/ mg (Table 5). Enzyme activity increased from 169.13 μmol/min/mL in crude enzyme to 218.46 μmol/min/mL after gel filtration. Crude protein and total protein decreased from 26.14 and 3920.46 mg/mL to 4.55 and 58.18 mg/mL, respectively.
| Step | Vol. | EA | CP | TP | TA | SA | Yield | Fold |
|---|---|---|---|---|---|---|---|---|
| Crude Enzyme | 150 | 169.13 | 26.14 | 3920.46 | 25369.98 | 6.47 | 100 | 1.00 |
| Ammonium sulphate precipitation | 44.1 | 188.87 | 17.05 | 751.70 | 8328.96 | 11.08 | 32.83 | 1.71 |
| Gel filtration | 12.8 | 218.46 | 4.55 | 58.18 | 2796.34 | 48.06 | 11.02 | 7.43 |
EA=Enzyme activity (μmol/min/mL)
CP=Crude protein (mg/mL)
TP=Total protein (mg/mL)
TA=Total activity (μmol/min/mL)
SA=Specific activity (μmol/min/mg)
Table 5: Summary of purification of mannanase from Aspergillus niger.
Discussion
Mannanases from fungi are generally produced into culture medium supplemented with different mannan-rich substrates known as inducers. These include Locust Bean Gum (LBG), guar gum, konjac flour and copra meal. Also, some microorganisms are capable of secreting considerable volume of extracellular enzymes into their basal media and this property has been adopted for industrial enzyme production.
The high fungal counts from water sample may be due to lack of efficient control measures in the discharge of organic wastes into the water bodies. The excessive addition of organic wastes (nutrients) to water bodies is known to stimulate excessive growth of microorganisms [10].
The fungal isolates obtained from Ilaje Lake (water and soil samples) exhibited different mannanase activities in basal media supplemented with LBG as an inducer. The production of mannanase in LBG medium had been reported for Bacillus circulans, Chryseobacterium indologenes, Bacillus sp. MG-33, Bacillus amylolequifaciens10A1 [4], Bacillus sp., Aspergillus niger, Sclerotium rolfsii, Trichoderma sp. and Scopulariopsis candida. The secretion of mannanases by these isolates on LBG media could be attributed to the ability of their genetic makeup to secrete active mannanase coupled with varied diffusion rate [10]. All the tested fungal strains were able to produce extracellular mannanase in submerged state fermentation, although with differences in the rate of enzyme production. These differences might be attributed to the source of isolation and slight variation in their genetic makeup [10].
The variation in protein content generated by each of the strains in submerged state fermentation could be attributed to the production of variety of enzymes (amylases, cellulases, protease and xylanases) in addition to the enzyme been examined in this study. Presumably, the protein from fungal cells and metabolites rich in protein might interfere with mannanase production causing variation in protein contents, since the protein assay could only identify accumulated protein in enzyme production medium [10].
Preliminary investigations in this study revealed that Aspergillus niger gave the highest mannanase activity out of all the fungal isolates encountered from the lake. It was therefore selected for purification studies. Mannanase has been produced by various microorganisms and purified as reported in previous studies. Olaniyi et al. [1] produced and purified mannanase enzyme from Penicillium italicum isolated from yam peel. Certain researchers sourced for β-1,4-mannanase from Scopulariosis candida and purified it while others isolated Geobacillus stearothermophilus L07 from oil palm shell, screened for mannanase production after which it was purified. Some researchers purified β-mannanase from Aspergillus oryzae. The two major activity peaks obtained from the elution profile could be attributed to the source of isolation coupled with environmental influences [1].
Conclusion
Aspergillus niger isolated from water sample in Ilaje, Ondo state, Nigeria showed a potential to convert substrates containing mannan into simple carbohydrates which could be readily used in many applications such as animal foods and a feed stock for production of prebiotics. It is recommended that the characterization of the purified β-mannanase and molecular study should be carried out on the isolate.
References
- Olaniyi OO, Arotupin DJ, Akinyele BJ, Bamidele OS (2014) Kinetic properties of purified β-mannanase from Penicillium italiticum. Br Microbiol Res J 4: 1092-1104.
- Songsiriritthigul C, Buranabanyat B, Haltrich D, Yamabhai M (2010) Efficient recombinant expression and secretion of a thermostable GH26 mannanendo-1,4-β-mannosidase from Bacillus licheniformis in Escherichia coli. Microb Cell Fact 9: 20.
- Dhawan S, Kaur J (2007) Microbial mannanases: An overview of production and applications. Crit Rev Biotechnol 27: 197–216.
- Mabrouk MEM, El Ahwany AMD (2008) Production of β-mannanase by Bacillus amylolequifaciens 10A1 cultured on potato peels. Afr J Biotechnol 7: 1123-1128.
- Wyman CE, Decker SR, Himmel ME, et al (2005) Polysaccharides: Structural diversity and functional versatility. CRC Press Boca Raton, FL. pp: 953-1033.
- Gomes J, Steiner W (2004) The biocatalytic potential of extremophiles and extrozymes. Food Technol Biotechnol 42: 223-225.
- Karan R, Khare SK (2010) Purification and characterization of a solvent stable protease from Geomicrobium sp. EMB2. Environ Technol 31: 1061-1072.
- Ajibare AO (2014) Assessment of physico-chemical parameters of waters in Ilaje local government area of Ondo state, Nigeria. International Journal of Fishery and Aquatic Studies 1: 84-92.
- Abe J, Hossain ZM, Hizukuri S (1994) Isolation of β-mannanase producing microorganisms. Journal of Fermentation and Bioengineering 3: 259-261.
- Arotupin DJ, Olaniyi OO (2013) Screening and identification of mannanase-producing fungi isolated from selected agricultural wastes. Br Microbiol Res J 3: 635-644.
- Lateef A, Oloke JK, Gueguim-Kana EB (2004) Antimicrobial resistance of bacterial strains isolated from orange juice products. Afr J Biotechnol 3: 334-338.
- Mandles M, Weber J (1969) Exoglucanase activity by microorganisms. Adv Chem 95: 391-414.
- El-Naggar MY, El-Aassar SA, Youssef AS, El-Sercy NA, Sercy A et al (2006) Extracellular β-mannanase production by the immobilization of the locally isolated Aspergillus niger. Int J Agric Biol 8: 57-62.
- Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugars. Anal Chem 31: 426-428.
- Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with folin phenol reagent. J Biol Chem 193: 265-275.
- Dhawan S, Jagdeep K (2007) Microbial mannanases: An overview of production and applications. Crit Rev Biotechnol 27: 197-216.
Relevant Topics
- Analytical Biochemistry
- Applied Biochemistry
- Carbohydrate Biochemistry
- Cellular Biochemistry
- Clinical_Biochemistry
- Comparative Biochemistry
- Environmental Biochemistry
- Forensic Biochemistry
- Lipid Biochemistry
- Medical_Biochemistry
- Metabolomics
- Nutritional Biochemistry
- Pesticide Biochemistry
- Process Biochemistry
- Protein_Biochemistry
- Single-Cell Biochemistry
- Soil_Biochemistry
Recommended Journals
- Biosensor Journals
- Cellular Biology Journal
- Journal of Biochemistry and Microbial Toxicology
- Journal of Biochemistry and Cell Biology
- Journal of Biological and Medical Sciences
- Journal of Cell Biology & Immunology
- Journal of Cellular and Molecular Pharmacology
- Journal of Chemical Biology & Therapeutics
- Journal of Phytochemicistry And Biochemistry
Article Tools
Article Usage
- Total views: 4239
- [From(publication date):
June-2017 - Aug 30, 2025] - Breakdown by view type
- HTML page views : 3269
- PDF downloads : 970